[English] 日本語
 Yorodumi
Yorodumi- EMDB-12715: IMC-Arches C6 at 8.33A - Refinement with C6 symmetry of the Inner... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | IMC-Arches C6 at 8.33A - Refinement with C6 symmetry of the Inner Membrane Complex (IMC) with the Arches from the fully-assembled R388 type IV secretion system. | |||||||||||||||
 Map data Map data | Inner Membrane Complex (IMC) with the Arches, 6-fold symmetry and sharpened map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | type IV secretion system / type 4 secretion system / T4SS / IMC / inner membrane complex / inner membrane / arches / periplasmic / R388 plasmid / conjugation / bacterial secretion / secretion / secretion system / protein complex / VirB3 / VirB4 / VirB8 / TrwM / TrwK / TrwG / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationprotein secretion by the type IV secretion system / ATP hydrolysis activity / ATP binding / membrane Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.6 Å | |||||||||||||||
 Authors Authors | Mace K / Vadakkepat AK | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Cryo-EM structure of a type IV secretion system. Authors: Kévin Macé / Abhinav K Vadakkepat / Adam Redzej / Natalya Lukoyanova / Clasien Oomen / Nathalie Braun / Marta Ukleja / Fang Lu / Tiago R D Costa / Elena V Orlova / David Baker / Qian Cong ...Authors: Kévin Macé / Abhinav K Vadakkepat / Adam Redzej / Natalya Lukoyanova / Clasien Oomen / Nathalie Braun / Marta Ukleja / Fang Lu / Tiago R D Costa / Elena V Orlova / David Baker / Qian Cong / Gabriel Waksman /     Abstract: Bacterial conjugation is the fundamental process of unidirectional transfer of DNAs, often plasmid DNAs, from a donor cell to a recipient cell. It is the primary means by which antibiotic resistance ...Bacterial conjugation is the fundamental process of unidirectional transfer of DNAs, often plasmid DNAs, from a donor cell to a recipient cell. It is the primary means by which antibiotic resistance genes spread among bacterial populations. In Gram-negative bacteria, conjugation is mediated by a large transport apparatus-the conjugative type IV secretion system (T4SS)-produced by the donor cell and embedded in both its outer and inner membranes. The T4SS also elaborates a long extracellular filament-the conjugative pilus-that is essential for DNA transfer. Here we present a high-resolution cryo-electron microscopy (cryo-EM) structure of a 2.8 megadalton T4SS complex composed of 92 polypeptides representing 8 of the 10 essential T4SS components involved in pilus biogenesis. We added the two remaining components to the structural model using co-evolution analysis of protein interfaces, to enable the reconstitution of the entire system including the pilus. This structure describes the exceptionally large protein-protein interaction network required to assemble the many components that constitute a T4SS and provides insights on the unique mechanism by which they elaborate pili. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12715.map.gz emd_12715.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12715-v30.xml emd-12715-v30.xml emd-12715.xml emd-12715.xml | 20.1 KB 20.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_12715.png emd_12715.png | 61.6 KB | ||
| Masks |  emd_12715_msk_1.map emd_12715_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-12715.cif.gz emd-12715.cif.gz | 6.2 KB | ||
| Others |  emd_12715_additional_1.map.gz emd_12715_additional_1.map.gz emd_12715_half_map_1.map.gz emd_12715_half_map_1.map.gz emd_12715_half_map_2.map.gz emd_12715_half_map_2.map.gz | 31.1 MB 59.3 MB 59.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12715 http://ftp.pdbj.org/pub/emdb/structures/EMD-12715 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12715 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12715 | HTTPS FTP |
-Validation report
| Summary document |  emd_12715_validation.pdf.gz emd_12715_validation.pdf.gz | 1009.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12715_full_validation.pdf.gz emd_12715_full_validation.pdf.gz | 1008.9 KB | Display | |
| Data in XML |  emd_12715_validation.xml.gz emd_12715_validation.xml.gz | 12.6 KB | Display | |
| Data in CIF |  emd_12715_validation.cif.gz emd_12715_validation.cif.gz | 14.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12715 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12715 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12715 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12715 | HTTPS FTP |
-Related structure data
| Related structure data |  7o41MC  7o3jC  7o3tC  7o3vC  7o42C  7o43C  7oiuC  7q1vC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12715.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12715.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Inner Membrane Complex (IMC) with the Arches, 6-fold symmetry and sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.134 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_12715_msk_1.map emd_12715_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
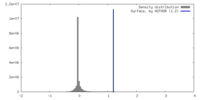
| Density Histograms |
-Additional map: Inner Membrane Complex (IMC) with the Arches, 6-fold...
| File | emd_12715_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Inner Membrane Complex (IMC) with the Arches, 6-fold symmetry and unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
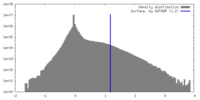
| Density Histograms |
-Half map: Inner Membrane Complex (IMC) with the Arches, 6-fold...
| File | emd_12715_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Inner Membrane Complex (IMC) with the Arches, 6-fold symmetry - Half_B | ||||||||||||
| Projections & Slices |
| ||||||||||||
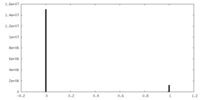
| Density Histograms |
-Half map: Inner Membrane Complex (IMC) with the Arches, 6-fold...
| File | emd_12715_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Inner Membrane Complex (IMC) with the Arches, 6-fold symmetry - Half_A | ||||||||||||
| Projections & Slices |
| ||||||||||||
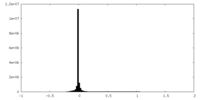
| Density Histograms |
- Sample components
Sample components
-Entire : Type IV secretion system complex
| Entire | Name: Type IV secretion system complex |
|---|---|
| Components |
|
-Supramolecule #1: Type IV secretion system complex
| Supramolecule | Name: Type IV secretion system complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 2.808 MDa |
-Macromolecule #1: TrwM protein
| Macromolecule | Name: TrwM protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 12.292585 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKPPQQQHEA FPLFKGATRL PTIWGVPMIP LMAMVMGVAV IALTVSIWWW ALVPPLWFIM AQITKNDDKA FRIWWLWIDT KFRNRNKGF WGASSYSPAN YRKRR UniProtKB: TrwM protein |
-Macromolecule #2: TrwK protein
| Macromolecule | Name: TrwK protein / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 93.76993 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGAIESRKLL ASETPVGQFI PYSHHVTDTI ISTKNAEYLS VWKIDGRSHQ SASEADVFQW IRELNNTLRG ISSANLSLWT HIVRRRVYE YPDAEFDNVF CRQLDEKYRE SFTGYNLMVN DLYLTVVYRP VSDKVLSFFA KRERETPDQK KHRQESCIKA L EDINRTLG ...String: MGAIESRKLL ASETPVGQFI PYSHHVTDTI ISTKNAEYLS VWKIDGRSHQ SASEADVFQW IRELNNTLRG ISSANLSLWT HIVRRRVYE YPDAEFDNVF CRQLDEKYRE SFTGYNLMVN DLYLTVVYRP VSDKVLSFFA KRERETPDQK KHRQESCIKA L EDINRTLG QSFKRYGAEL LSVYEKGGHA FSAPLEFLAR LVNGEHIPMP ICRDRFSDYM AVNRPMFSKW GEVGELRSLT GL RRFGMLE IREYDDATEP GQLNVLLESD YEFVLTHSFS VLSRPAAKEY LQRHQKNLID ARDVATDQIE EIDEALNQLI SGH FVMGEH HCTLTVYGET VQQVRDNLAH ASAAMLDVAV LPKPVDLALE AGYWAQLPAN WQWRPRPAPI TSLNFLSFSP FHNF MSGKP TGNPWGPAVT ILKTVSGTPL YFNFHASKEE EDATDKRLLG NTMLIGQSSS GKTVLLGFLL AQAQKFKPTI VAFDK DRGM EISIRAMGGR YLPLKTGEPS GFNPFQLPPT HANLIFLKQF VKKLAAAGGE VTHRDEEEID QAITAMMSDS IDKSLR RLS LLLQFLPNPR SDDMDARPTV HARLVKWCEG GDYGWLFDNP TDALDLSTHQ IYGFDITEFL DNPEARTPVM MYLLYRT ES MIDGRRFMYV FDEFWKPLQD EYFEDLAKNK QKTIRKQNGI FVFATQEPSD ALESNIAKTL IQQCATYIFL ANPKADYE D YTQGFKLTDS EFELVRGLGE FSRRFLIKQG DQSALAEMNL GKFRTIVDGE TVERDFDDEL LVLSGTPDNA EIAESIIAE VGDDPAVWLP IFLDRVKAER SDV UniProtKB: Type IV secretion system protein virB4 |
-Macromolecule #3: TrwG protein
| Macromolecule | Name: TrwG protein / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25.799994 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSKKQPKPVK AEQLKSYYEE SRGLERDLIG EFVKSRKTAW RVATASGLFG LLGMVCGIVG FSQPAPAPLV LRVDNATGAV DVVTTLREH ESSYGEVVDT YWLNQYVLNR EAYDYNTIQM NYDTTALLSA PAVQQDYYKL FDGSNARDRV LGNKARITVR V RSIQPNGR ...String: MSKKQPKPVK AEQLKSYYEE SRGLERDLIG EFVKSRKTAW RVATASGLFG LLGMVCGIVG FSQPAPAPLV LRVDNATGAV DVVTTLREH ESSYGEVVDT YWLNQYVLNR EAYDYNTIQM NYDTTALLSA PAVQQDYYKL FDGSNARDRV LGNKARITVR V RSIQPNGR GQATVRFTTQ QHNSNGTVEA PQHQIATIGY TYIGAPMRSS DRLLNPLGFQ VTSYRADPEI LNN UniProtKB: TrwG protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 57.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: CryoSPARC ab-initio |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C6 (6 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 7.6 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 126975 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: OTHER / Details: Stochastic gradient descent (SGD) |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)