[English] 日本語
 Yorodumi
Yorodumi- EMDB-1241: Visualizing the ATPase cycle in a protein disaggregating machine:... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1241 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Visualizing the ATPase cycle in a protein disaggregating machine: structural basis for substrate binding by ClpB. | |||||||||
 Map data Map data | volume file of TClpB apo state | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Thermus thermophilus (bacteria) Thermus thermophilus (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 17.7 Å | |||||||||
 Authors Authors | Tsai F | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2007 Journal: Mol Cell / Year: 2007Title: Visualizing the ATPase cycle in a protein disaggregating machine: structural basis for substrate binding by ClpB. Authors: Sukyeong Lee / Jae-Mun Choi / Francis T F Tsai /  Abstract: ClpB is a ring-shaped molecular chaperone that has the remarkable ability to disaggregate stress-damaged proteins. Here we present the electron cryomicroscopy reconstruction of an ATP-activated ClpB ...ClpB is a ring-shaped molecular chaperone that has the remarkable ability to disaggregate stress-damaged proteins. Here we present the electron cryomicroscopy reconstruction of an ATP-activated ClpB trap mutant, along with reconstructions of ClpB in the AMPPNP, ADP, and in the nucleotide-free state. We show that motif 2 of the ClpB M domain is positioned between the D1-large domains of neighboring subunits and could facilitate a concerted, ATP-driven conformational change in the AAA-1 ring. We further demonstrate biochemically that ATP is essential for high-affinity substrate binding to ClpB and cannot be substituted with AMPPNP. Our structures show that in the ATP-activated state, the D1 loops are stabilized at the central pore, providing the structural basis for high-affinity substrate binding. Taken together, our results support a mechanism by which ClpB captures substrates on the upper surface of the AAA-1 ring before threading them through the ClpB hexamer in an ATP hydrolysis-driven step. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1241.map.gz emd_1241.map.gz | 879.5 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1241-v30.xml emd-1241-v30.xml emd-1241.xml emd-1241.xml | 8.4 KB 8.4 KB | Display Display |  EMDB header EMDB header |
| Images |  1241.gif 1241.gif | 23.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1241 http://ftp.pdbj.org/pub/emdb/structures/EMD-1241 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1241 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1241 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1241.map.gz / Format: CCP4 / Size: 5.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1241.map.gz / Format: CCP4 / Size: 5.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | volume file of TClpB apo state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.17 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
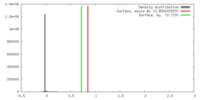
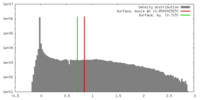
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ClpB
| Entire | Name: ClpB |
|---|---|
| Components |
|
-Supramolecule #1000: ClpB
| Supramolecule | Name: ClpB / type: sample / ID: 1000 / Oligomeric state: homohexamer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 600 KDa / Theoretical: 600 KDa |
-Macromolecule #1: ClpB
| Macromolecule | Name: ClpB / type: protein_or_peptide / ID: 1 / Oligomeric state: hexamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:   Thermus thermophilus (bacteria) / Cell: E.Coli / Location in cell: cytoplasm Thermus thermophilus (bacteria) / Cell: E.Coli / Location in cell: cytoplasm |
| Molecular weight | Experimental: 600 KDa / Theoretical: 600 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.03 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 50mM MOPS pH7.5, 150mM KCl, 5mM MgCl2 |
| Grid | Details: 400 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 90 K / Instrument: REICHERT-JUNG PLUNGER / Details: Vitrification instrument: Reichert plunger |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2010F |
|---|---|
| Temperature | Average: 90 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 400,000x magnification |
| Date | Oct 31, 2003 |
| Image recording | Category: CCD / Film or detector model: GENERIC GATAN (4k x 4k) / Number real images: 34 / Average electron dose: 15 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 1.0 mm / Nominal defocus max: 3.9 µm / Nominal defocus min: 1.9 µm / Nominal magnification: 69000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: each CCD frame |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C6 (6 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 17.7 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: EMAN / Number images used: 10453 |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)