+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12341 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Akirin2 bound human proteasome | |||||||||
 Map data Map data | denoised map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | proteasome / nuclear import / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationproteasome localization / regulation of muscle cell differentiation / positive regulation of B cell activation / nuclear protein quality control by the ubiquitin-proteasome system / positive regulation of adaptive immune response / embryo development ending in birth or egg hatching / purine ribonucleoside triphosphate binding / positive regulation of innate immune response / Antigen processing: Ub, ATP-independent proteasomal degradation / sperm glycocalyx ...proteasome localization / regulation of muscle cell differentiation / positive regulation of B cell activation / nuclear protein quality control by the ubiquitin-proteasome system / positive regulation of adaptive immune response / embryo development ending in birth or egg hatching / purine ribonucleoside triphosphate binding / positive regulation of innate immune response / Antigen processing: Ub, ATP-independent proteasomal degradation / sperm glycocalyx / Regulation of ornithine decarboxylase (ODC) / Proteasome assembly / proteasome core complex / perinuclear theca / Cross-presentation of soluble exogenous antigens (endosomes) / Somitogenesis / myofibril / proteasomal ubiquitin-independent protein catabolic process / sperm head-tail coupling apparatus / immune system process / proteasome endopeptidase complex / NF-kappaB binding / proteasome core complex, beta-subunit complex / threonine-type endopeptidase activity / proteasome assembly / proteasome core complex, alpha-subunit complex / transcription repressor complex / : / ciliary tip / proteasome complex / Regulation of activated PAK-2p34 by proteasome mediated degradation / sarcomere / transcription coregulator activity / Autodegradation of Cdh1 by Cdh1:APC/C / APC/C:Cdc20 mediated degradation of Securin / Asymmetric localization of PCP proteins / Ubiquitin-dependent degradation of Cyclin D / SCF-beta-TrCP mediated degradation of Emi1 / NIK-->noncanonical NF-kB signaling / AUF1 (hnRNP D0) binds and destabilizes mRNA / TNFR2 non-canonical NF-kB pathway / centriole / sperm end piece / negative regulation of inflammatory response to antigenic stimulus / P-body / Assembly of the pre-replicative complex / Vpu mediated degradation of CD4 / lipopolysaccharide binding / Cdc20:Phospho-APC/C mediated degradation of Cyclin A / Dectin-1 mediated noncanonical NF-kB signaling / Degradation of DVL / Degradation of AXIN / Degradation of CRY and PER proteins / Hh mutants are degraded by ERAD / Activation of NF-kappaB in B cells / G2/M Checkpoints / Degradation of GLI1 by the proteasome / Hedgehog ligand biogenesis / Regulation of RUNX3 expression and activity / Autodegradation of the E3 ubiquitin ligase COP1 / Defective CFTR causes cystic fibrosis / GSK3B and BTRC:CUL1-mediated-degradation of NFE2L2 / Negative regulation of NOTCH4 signaling / cerebral cortex development / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / Hedgehog 'on' state / Vif-mediated degradation of APOBEC3G / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / Degradation of GLI2 by the proteasome / GLI3 is processed to GLI3R by the proteasome / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / MAPK6/MAPK4 signaling / Degradation of CDH1 / positive regulation of interleukin-6 production / Degradation of beta-catenin by the destruction complex / Oxygen-dependent proline hydroxylation of Hypoxia-inducible Factor Alpha / CDK-mediated phosphorylation and removal of Cdc6 / ABC-family protein mediated transport / CLEC7A (Dectin-1) signaling / SCF(Skp2)-mediated degradation of p27/p21 / FCERI mediated NF-kB activation / response to virus / Regulation of expression of SLITs and ROBOs / nuclear matrix / Regulation of PTEN stability and activity / Interleukin-1 signaling / Orc1 removal from chromatin / protein import into nucleus / Regulation of RAS by GAPs / Regulation of RUNX2 expression and activity / The role of GTSE1 in G2/M progression after G2 checkpoint / Separation of Sister Chromatids / UCH proteinases / KEAP1-NFE2L2 pathway / Downstream TCR signaling / peptidase activity / Antigen processing: Ubiquitination & Proteasome degradation / RUNX1 regulates transcription of genes involved in differentiation of HSCs / sperm principal piece / ER-Phagosome pathway Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Singh K / Brunner H / Grishkovskaya I / de Almeida M / Hinterndorfer M / Zuber J / Haselbach D | |||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: AKIRIN2 controls the nuclear import of proteasomes in vertebrates. Authors: Melanie de Almeida / Matthias Hinterndorfer / Hanna Brunner / Irina Grishkovskaya / Kashish Singh / Alexander Schleiffer / Julian Jude / Sumit Deswal / Robert Kalis / Milica Vunjak / Thomas ...Authors: Melanie de Almeida / Matthias Hinterndorfer / Hanna Brunner / Irina Grishkovskaya / Kashish Singh / Alexander Schleiffer / Julian Jude / Sumit Deswal / Robert Kalis / Milica Vunjak / Thomas Lendl / Richard Imre / Elisabeth Roitinger / Tobias Neumann / Susanne Kandolf / Michael Schutzbier / Karl Mechtler / Gijs A Versteeg / David Haselbach / Johannes Zuber /     Abstract: Protein expression and turnover are controlled through a complex interplay of transcriptional, post-transcriptional and post-translational mechanisms to enable spatial and temporal regulation of ...Protein expression and turnover are controlled through a complex interplay of transcriptional, post-transcriptional and post-translational mechanisms to enable spatial and temporal regulation of cellular processes. To systematically elucidate such gene regulatory networks, we developed a CRISPR screening assay based on time-controlled Cas9 mutagenesis, intracellular immunostaining and fluorescence-activated cell sorting that enables the identification of regulatory factors independent of their effects on cellular fitness. We pioneered this approach by systematically probing the regulation of the transcription factor MYC, a master regulator of cell growth. Our screens uncover a highly conserved protein, AKIRIN2, that is essentially required for nuclear protein degradation. We found that AKIRIN2 forms homodimers that directly bind to fully assembled 20S proteasomes to mediate their nuclear import. During mitosis, proteasomes are excluded from condensing chromatin and re-imported into newly formed daughter nuclei in a highly dynamic, AKIRIN2-dependent process. Cells undergoing mitosis in the absence of AKIRIN2 become devoid of nuclear proteasomes, rapidly causing accumulation of MYC and other nuclear proteins. Collectively, our study reveals a dedicated pathway controlling the nuclear import of proteasomes in vertebrates and establishes a scalable approach to decipher regulators in essential cellular processes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12341.map.gz emd_12341.map.gz | 150.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12341-v30.xml emd-12341-v30.xml emd-12341.xml emd-12341.xml | 35 KB 35 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_12341.png emd_12341.png | 72.5 KB | ||
| Filedesc metadata |  emd-12341.cif.gz emd-12341.cif.gz | 9.1 KB | ||
| Others |  emd_12341_additional_1.map.gz emd_12341_additional_1.map.gz | 141.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12341 http://ftp.pdbj.org/pub/emdb/structures/EMD-12341 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12341 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12341 | HTTPS FTP |
-Related structure data
| Related structure data |  7nhtMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10752 (Title: Single particle cryo EM Data of a human Akirin2 proteasome complex EMPIAR-10752 (Title: Single particle cryo EM Data of a human Akirin2 proteasome complexData size: 2.0 TB Data #1: Unaligned Multiframe micrographs of Akirin2 bound to the human proteasome [micrographs - multiframe] Data #2: Motion corrected files dose weighted and summed as well as without dose weigthening [micrographs - single frame] Data #3: Extracted particles of an Akirin2 bound proteasome [picked particles - single frame - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12341.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12341.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | denoised map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
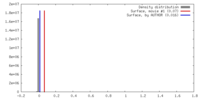
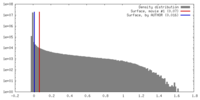
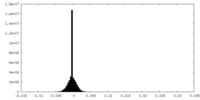
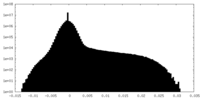
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: unsharpend map
| File | emd_12341_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpend map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Akirin2 bound to 20S proteasome
+Supramolecule #1: Akirin2 bound to 20S proteasome
+Supramolecule #2: 20S proteasome
+Supramolecule #3: Akirin2
+Macromolecule #1: Proteasome subunit alpha type-2
+Macromolecule #2: Proteasome subunit alpha type-4
+Macromolecule #3: Proteasome subunit alpha type-7
+Macromolecule #4: Proteasome subunit alpha type-5
+Macromolecule #5: Proteasome subunit alpha type-1
+Macromolecule #6: Proteasome subunit alpha type-3
+Macromolecule #7: Proteasome subunit alpha type-6
+Macromolecule #8: Proteasome subunit beta type-7
+Macromolecule #9: Proteasome subunit beta type-3
+Macromolecule #10: Proteasome subunit beta type-2
+Macromolecule #11: Proteasome subunit beta type-5
+Macromolecule #12: Proteasome subunit beta type-1
+Macromolecule #13: Proteasome subunit beta type-4
+Macromolecule #14: Proteasome subunit beta type-6
+Macromolecule #15: Akirin-2
+Macromolecule #16: POTASSIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.5 Component:
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R3.5/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 4595 / Average exposure time: 1.0 sec. / Average electron dose: 33.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)