+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12054 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | TRAPP C11 region of the whole TRAPPIII complex | |||||||||
 Map data Map data | Cryo-EM map of the TRAPPC11 region of the Drosophila melanogaster TRAPPIII complex. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Golgi / GEFS / Rab1 / TRAPP / EXOCYTOSIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationRAB GEFs exchange GTP for GDP on RABs / TRAPPIII protein complex / Golgi vesicle transport / protein secretion / long-term memory / intracellular transport / Golgi apparatus / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
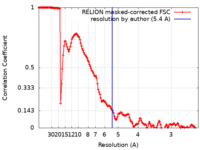
| Method | single particle reconstruction / cryo EM / Resolution: 5.4 Å | |||||||||
 Authors Authors | Galindo A / Munro S | |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2021 Journal: EMBO J / Year: 2021Title: Cryo-EM structure of metazoan TRAPPIII, the multi-subunit complex that activates the GTPase Rab1. Authors: Antonio Galindo / Vicente J Planelles-Herrero / Gianluca Degliesposti / Sean Munro /  Abstract: The TRAPP complexes are nucleotide exchange factors that play essential roles in membrane traffic and autophagy. TRAPPII activates Rab11, and TRAPPIII activates Rab1, with the two complexes sharing a ...The TRAPP complexes are nucleotide exchange factors that play essential roles in membrane traffic and autophagy. TRAPPII activates Rab11, and TRAPPIII activates Rab1, with the two complexes sharing a core of small subunits that affect nucleotide exchange but being distinguished by specific large subunits that are essential for activity in vivo. Crystal structures of core subunits have revealed the mechanism of Rab activation, but how the core and the large subunits assemble to form the complexes is unknown. We report a cryo-EM structure of the entire Drosophila TRAPPIII complex. The TRAPPIII-specific subunits TRAPPC8 and TRAPPC11 hold the catalytic core like a pair of tongs, with TRAPPC12 and TRAPPC13 positioned at the joint between them. TRAPPC2 and TRAPPC2L link the core to the two large arms, with the interfaces containing residues affected by disease-causing mutations. The TRAPPC8 arm is positioned such that it would contact Rab1 that is bound to the core, indicating how the arm could determine the specificity of the complex. A lower resolution structure of TRAPPII shows a similar architecture and suggests that the TRAPP complexes evolved from a single ur-TRAPP. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12054.map.gz emd_12054.map.gz | 226.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12054-v30.xml emd-12054-v30.xml emd-12054.xml emd-12054.xml | 13.9 KB 13.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12054_fsc.xml emd_12054_fsc.xml | 14.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_12054.png emd_12054.png | 144.7 KB | ||
| Masks |  emd_12054_msk_1.map emd_12054_msk_1.map | 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-12054.cif.gz emd-12054.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12054 http://ftp.pdbj.org/pub/emdb/structures/EMD-12054 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12054 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12054 | HTTPS FTP |
-Related structure data
| Related structure data |  7b6hMC  7b6dC  7b6eC  7b6rC  7b6xC  7b6yC  7b6zC  7b70C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12054.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12054.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of the TRAPPC11 region of the Drosophila melanogaster TRAPPIII complex. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.248 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
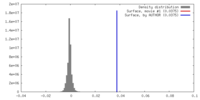
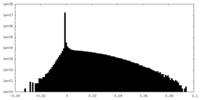
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12054_msk_1.map emd_12054_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : TRAPPIII complex - TRAPPC11 subunit
| Entire | Name: TRAPPIII complex - TRAPPC11 subunit |
|---|---|
| Components |
|
-Supramolecule #1: TRAPPIII complex - TRAPPC11 subunit
| Supramolecule | Name: TRAPPIII complex - TRAPPC11 subunit / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Partial model of the TRAPP C11 subunits from the C11 map. Part of the TRAPPIII complex |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Trafficking protein particle complex subunit 11
| Macromolecule | Name: Trafficking protein particle complex subunit 11 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 81.847164 KDa |
| Recombinant expression | Organism:  Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) |
| Sequence | String: MTMDATALPS ELLVTPQPLV GFCGLDTARV SVHKAVWEAF SGSLQRKAAD RAAVQYKLLP PNYEFPVAKP KRASYEWYHP KGILKRNWM LKHLHVLPSV VVLFQDMEWN DLQWTEKQVQ CAAIVQALKN TLQERNTRLC LVLLQRAAPL PPGEDLLAAE R AASLTNAC ...String: MTMDATALPS ELLVTPQPLV GFCGLDTARV SVHKAVWEAF SGSLQRKAAD RAAVQYKLLP PNYEFPVAKP KRASYEWYHP KGILKRNWM LKHLHVLPSV VVLFQDMEWN DLQWTEKQVQ CAAIVQALKN TLQERNTRLC LVLLQRAAPL PPGEDLLAAE R AASLTNAC GITSKMLFIL PHTEHLTGYA LRLESAFLDM AQSYYALMSK RIRNHRDQLT AAHTSLKIRH QFKLGFVAEM RQ DFSTGQK HYFQAYANLD EIRINDGNCL EIKTLAGFLN YKICRLMFKL KTPRDAINQF IIHVEKHKSR VGFKDLAFEH HAW LSTQHS VFAELFCEAI KNGLPALQTQ HPGIYYHKAA EFVMKRRDAA MEAYAAMQAS SEATPTPIQN PLSLYTEFFG IRAV KTGDL VAEQQANMQL CDQERSYNHS AAIIALLSQA MAQFKIYKCL RFRKKLAIDM AEEYLKSGDH AKALTLYSLM LPDYR QEKW TTIFTDVLLK TLRCALLSGS VADYIACSVE ALSLRHQSDQ SERILILENL WQVFQGVPPM PKTQLTPEAQ ALWTSA LAN VKSPIQIDLD KVNDVVEMCA TFERVQLSND DLLQLQLIVR VLTDIPLRIR SFHVILADAG NPQNSYKLEA LKYFCFP TL TQLRGQKQPD DEQLENPSQE PKNFEKNMRL EPGSYYQLFC STEAQQFHEN TQLRIVRLEA HMGTDQVAAL LTCSSN UniProtKB: Trafficking protein particle complex subunit 11 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.05 mg/mL | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.001 kPa | ||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 285.15 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Details | 32% of the images were acquiring tilted the stage 19 degrees. |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 4 / Number real images: 3671 / Average exposure time: 0.8 sec. / Average electron dose: 30.0 e/Å2 Details: 1190 from the whole dataset were collected with the stage tilted at 19 degrees |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated magnification: 75000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.2 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 302.4 |
|---|---|
| Output model |  PDB-7b6h: |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)