[English] 日本語
 Yorodumi
Yorodumi- EMDB-11930: E. coli NADH quinone oxidoreductase hydrophilic arm (CASP target) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11930 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli NADH quinone oxidoreductase hydrophilic arm (CASP target) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | E. coli / respiratory complex I / ELECTRON TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationoxidoreductase complex / NADH dehydrogenase (quinone) (non-electrogenic) activity / Translocases; Catalysing the translocation of protons; Linked to oxidoreductase reactions / NADH dehydrogenase complex / molybdopterin cofactor binding / NADH dehydrogenase activity / NADH dehydrogenase (ubiquinone) activity / quinone binding / ATP synthesis coupled electron transport / respiratory electron transport chain ...oxidoreductase complex / NADH dehydrogenase (quinone) (non-electrogenic) activity / Translocases; Catalysing the translocation of protons; Linked to oxidoreductase reactions / NADH dehydrogenase complex / molybdopterin cofactor binding / NADH dehydrogenase activity / NADH dehydrogenase (ubiquinone) activity / quinone binding / ATP synthesis coupled electron transport / respiratory electron transport chain / 2 iron, 2 sulfur cluster binding / NAD binding / FMN binding / 4 iron, 4 sulfur cluster binding / oxidoreductase activity / iron ion binding / membrane / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.73 Å | |||||||||
 Authors Authors | Schimpf J / Grishkovskaya I / Haselbach D / Friedrich T | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2022 Journal: Structure / Year: 2022Title: Structure of the peripheral arm of a minimalistic respiratory complex I. Authors: Johannes Schimpf / Sabrina Oppermann / Tatjana Gerasimova / Ana Filipa Santos Seica / Petra Hellwig / Irina Grishkovskaya / Daniel Wohlwend / David Haselbach / Thorsten Friedrich /    Abstract: Respiratory complex I drives proton translocation across energy-transducing membranes by NADH oxidation coupled with (ubi)quinone reduction. In humans, its dysfunction is associated with ...Respiratory complex I drives proton translocation across energy-transducing membranes by NADH oxidation coupled with (ubi)quinone reduction. In humans, its dysfunction is associated with neurodegenerative diseases. The Escherichia coli complex represents the structural minimal form of an energy-converting NADH:ubiquinone oxidoreductase. Here, we report the structure of the peripheral arm of the E. coli complex I consisting of six subunits, the FMN cofactor, and nine iron-sulfur clusters at 2.7 Å resolution obtained by cryo electron microscopy. While the cofactors are in equivalent positions as in the complex from other species, individual subunits are adapted to the absence of supernumerary proteins to guarantee structural stability. The catalytically important subunits NuoC and D are fused resulting in a specific architecture of functional importance. Striking features of the E. coli complex are scrutinized by mutagenesis and biochemical characterization of the variants. Moreover, the arrangement of the subunits sheds light on the unknown assembly of the complex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11930.map.gz emd_11930.map.gz | 78.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11930-v30.xml emd-11930-v30.xml emd-11930.xml emd-11930.xml | 22.8 KB 22.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11930.png emd_11930.png | 69.2 KB | ||
| Filedesc metadata |  emd-11930.cif.gz emd-11930.cif.gz | 10.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11930 http://ftp.pdbj.org/pub/emdb/structures/EMD-11930 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11930 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11930 | HTTPS FTP |
-Related structure data
| Related structure data |  7awtMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11930.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11930.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
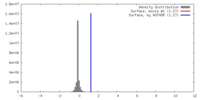
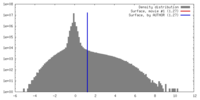
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : NADH quinone oxidoreductase hydrophilic arm
+Supramolecule #1: NADH quinone oxidoreductase hydrophilic arm
+Macromolecule #1: NADH-quinone oxidoreductase subunit B
+Macromolecule #2: NADH-quinone oxidoreductase subunit C/D
+Macromolecule #3: NADH-quinone oxidoreductase subunit E
+Macromolecule #4: NADH-quinone oxidoreductase subunit F
+Macromolecule #5: NADH-quinone oxidoreductase
+Macromolecule #6: NADH-quinone oxidoreductase subunit I
+Macromolecule #7: IRON/SULFUR CLUSTER
+Macromolecule #8: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #9: MAGNESIUM ION
+Macromolecule #10: FLAVIN MONONUCLEOTIDE
+Macromolecule #11: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 6 Component:
| |||||||||||||||
| Grid | Model: Quantifoil / Material: COPPER / Mesh: 200 / Support film - Material: CARBON | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Chamber temperature: 277 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Average exposure time: 1.0 sec. / Average electron dose: 67.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.73 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 2.42) / Number images used: 223751 |
| Initial angle assignment | Type: PROJECTION MATCHING / Software - Name: cryoSPARC (ver. 2.42) |
| Final angle assignment | Type: PROJECTION MATCHING / Software - Name: cryoSPARC (ver. 2.42) |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)