+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11356 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of detergent-extracted full-length E.coli BcsB | ||||||||||||||||||
 Map data Map data | Main map non-uniformly refined. It represents two BcsB octamers (6YG8) linked by a central miscellaneous structure. | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Function / homology | Cellulose synthase, subunit B / Biofilm formation / Cellulose synthase BcsB, bacterial / Bacterial cellulose synthase subunit / cellulose biosynthetic process / UDP-alpha-D-glucose metabolic process / plasma membrane / Cyclic di-GMP-binding protein / Cyclic di-GMP-binding protein Function and homology information Function and homology information | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.9 Å | ||||||||||||||||||
 Authors Authors | Abidi W / Zouhir S / Krasteva P | ||||||||||||||||||
| Funding support | 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2021 Journal: Sci Adv / Year: 2021Title: Architecture and regulation of an enterobacterial cellulose secretion system. Authors: Wiem Abidi / Samira Zouhir / Meryem Caleechurn / Stéphane Roche / Petya Violinova Krasteva /  Abstract: Many free-living and pathogenic enterobacteria secrete biofilm-promoting cellulose using a multicomponent, envelope-embedded Bcs secretion system under the control of intracellular second messenger c- ...Many free-living and pathogenic enterobacteria secrete biofilm-promoting cellulose using a multicomponent, envelope-embedded Bcs secretion system under the control of intracellular second messenger c-di-GMP. The molecular understanding of system assembly and cellulose secretion has been largely limited to the crystallographic studies of a distantly homologous BcsAB synthase tandem and a low-resolution reconstruction of an assembled macrocomplex that encompasses most of the inner membrane and cytosolic subunits and features an atypical layered architecture. Here, we present cryo-EM structures of the assembled Bcs macrocomplex, as well as multiple crystallographic snapshots of regulatory Bcs subcomplexes. The structural and functional data uncover the mechanism of asymmetric secretion system assembly and periplasmic crown polymerization and reveal unexpected subunit stoichiometry, multisite c-di-GMP recognition, and ATP-dependent regulation. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11356.map.gz emd_11356.map.gz | 129.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11356-v30.xml emd-11356-v30.xml emd-11356.xml emd-11356.xml | 20.1 KB 20.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11356.png emd_11356.png | 90 KB | ||
| Others |  emd_11356_additional_1.map.gz emd_11356_additional_1.map.gz emd_11356_additional_2.map.gz emd_11356_additional_2.map.gz emd_11356_additional_3.map.gz emd_11356_additional_3.map.gz | 248 MB 162.6 MB 241.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11356 http://ftp.pdbj.org/pub/emdb/structures/EMD-11356 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11356 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11356 | HTTPS FTP |
-Related structure data
| Related structure data |  6yarC  6yayC  6yb3C  6yb5C  6ybbC  6ybuC  6yg8C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11356.map.gz / Format: CCP4 / Size: 262.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11356.map.gz / Format: CCP4 / Size: 262.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main map non-uniformly refined. It represents two BcsB octamers (6YG8) linked by a central miscellaneous structure. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.13 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Main map non-uniformly refined and sharpened.
| File | emd_11356_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main map non-uniformly refined and sharpened. | ||||||||||||
| Projections & Slices |
| ||||||||||||
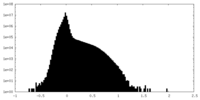
| Density Histograms |
-Additional map: The volumes corresponding to the octamers were extracted...
| File | emd_11356_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The volumes corresponding to the octamers were extracted and locally refined to be finally combined resulting in a map without the central miscellaneous structure. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: The volumes corresponding to the octamers were extracted...
| File | emd_11356_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The volumes corresponding to the octamers were extracted and locally refined to be finally combined and sharpened resulting in a map without the central miscellaneous structure. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Self-assembly of the BcsB protein
| Entire | Name: Self-assembly of the BcsB protein |
|---|---|
| Components |
|
-Supramolecule #1: Self-assembly of the BcsB protein
| Supramolecule | Name: Self-assembly of the BcsB protein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 1.378 MDa |
-Macromolecule #1: Bacterial cellulose synthase regulator protein BcsB
| Macromolecule | Name: Bacterial cellulose synthase regulator protein BcsB / type: protein_or_peptide / ID: 1 Details: BcsB harbors a signal sequence to be addressed to the periplasm where according the ServerP4.1 Server prediction would result in mature form starting with the residue #26 TPATQ... Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: TPATQPLINA EPAVAAQTEQ NPQVGQVMPG VQGADAPVVA QNGPSRDVKL TFAQIAPPPG SMVLRGINPN GSIEFGMRSD EVVTKAMLNL EYTPS PSLL PVQSQLKVYL NDELMGVLPV TKEQLGKKTL AQMPINPLFI TDFNRVRLEF VGHYQD VCE NPASTTLWLD ...String: TPATQPLINA EPAVAAQTEQ NPQVGQVMPG VQGADAPVVA QNGPSRDVKL TFAQIAPPPG SMVLRGINPN GSIEFGMRSD EVVTKAMLNL EYTPS PSLL PVQSQLKVYL NDELMGVLPV TKEQLGKKTL AQMPINPLFI TDFNRVRLEF VGHYQD VCE NPASTTLWLD VGRSSGLDLT YQTLNVKNDL SHFPVPFFDP RDNRTNTLPM VFAGAPD VG LQQASAIVAS WFGSRSGWRG QNFPVLYNQL PDRNAIVFAT NDKRPDFLRD HPAVKAPV I EMINHPQNPY VKLLVVFGRD DKDLLQAAKG IAQGNILFRG ESVVVNEVKP LLPRKPYDA PNWVRTDRPV TFGELKTYEE QLQSSGLEPA AINVSLNLPP DLYLMRSTGI DMDINYRYTM PPVKDSSRM DISLNNQFLQ SFNLSSKQEA NRLLLRIPVL QGLLDGKTDV SIPALKLGAT N QLRFDFEY MNPMPGGSVD NCITFQPVQN HVVIGDDSTI DFSKYYHFIP MPDLRAFANA GF PFSRMAD LSQTITVMPK APNEAQMETL LNTVGFIGAQ TGFPAINLTV TDDGSTIQGK DAD IMIIGG IPDKLKDDKQ IDLLVQATES WVKTPMRQTP FPGIVPDESD RAAETRSTLT SSGA MAAVI GFQSPYNDQR SVIALLADSP RGYEMLNDAV NDSGKRATMF GSVAVIRESG INSLR VGDV YYVGHLPWFE RLWYALANHP ILLAVLAAIS VILLAWVLWR LLRIISRRRL NPDNEAAALE HHHHHH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
Details: BcsB was incubated with a mix of detergents: 0.4% DDM, 0.4 % digitonin, 0.4% DM-NPG, 0.2 % GDN-101, and 0.2% LM-NPG and then purified on IMAC using free-detergents buffers. | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS TALOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 0.74 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.75 µm / Nominal defocus min: 0.75 µm / Nominal magnification: 36000 |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)