+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11193 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | COPII on membranes, outer coat right-handed rod, class1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PROTEIN TRANSPORT / SECRETION / TRAFFICKING | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of ER to Golgi vesicle-mediated transport / Seh1-associated complex / COPII-coated vesicle budding / nuclear pore localization / protein exit from endoplasmic reticulum / COPII-mediated vesicle transport / regulation of TORC1 signaling / nuclear pore outer ring / COPII-coated vesicle cargo loading / Regulation of Glucokinase by Glucokinase Regulatory Protein ...positive regulation of ER to Golgi vesicle-mediated transport / Seh1-associated complex / COPII-coated vesicle budding / nuclear pore localization / protein exit from endoplasmic reticulum / COPII-mediated vesicle transport / regulation of TORC1 signaling / nuclear pore outer ring / COPII-coated vesicle cargo loading / Regulation of Glucokinase by Glucokinase Regulatory Protein / positive regulation of protein exit from endoplasmic reticulum / : / Regulation of HSF1-mediated heat shock response / COPII vesicle coat / SUMOylation of SUMOylation proteins / mating projection tip / endoplasmic reticulum organization / SUMOylation of RNA binding proteins / SUMOylation of chromatin organization proteins / vacuolar membrane / nucleocytoplasmic transport / endoplasmic reticulum exit site / positive regulation of TOR signaling / nuclear pore / mRNA transport / ERAD pathway / positive regulation of TORC1 signaling / cell periphery / protein import into nucleus / nuclear envelope / protein transport / endoplasmic reticulum membrane / positive regulation of DNA-templated transcription / structural molecule activity / endoplasmic reticulum Similarity search - Function | |||||||||
| Biological species |  | |||||||||
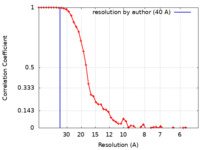
| Method | subtomogram averaging / cryo EM / Resolution: 40.0 Å | |||||||||
 Authors Authors | Zanetti G / Hutchings J | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Structure of the complete, membrane-assembled COPII coat reveals a complex interaction network. Authors: Joshua Hutchings / Viktoriya G Stancheva / Nick R Brown / Alan C M Cheung / Elizabeth A Miller / Giulia Zanetti /   Abstract: COPII mediates Endoplasmic Reticulum to Golgi trafficking of thousands of cargoes. Five essential proteins assemble into a two-layer architecture, with the inner layer thought to regulate coat ...COPII mediates Endoplasmic Reticulum to Golgi trafficking of thousands of cargoes. Five essential proteins assemble into a two-layer architecture, with the inner layer thought to regulate coat assembly and cargo recruitment, and the outer coat forming cages assumed to scaffold membrane curvature. Here we visualise the complete, membrane-assembled COPII coat by cryo-electron tomography and subtomogram averaging, revealing the full network of interactions within and between coat layers. We demonstrate the physiological importance of these interactions using genetic and biochemical approaches. Mutagenesis reveals that the inner coat alone can provide membrane remodelling function, with organisational input from the outer coat. These functional roles for the inner and outer coats significantly move away from the current paradigm, which posits membrane curvature derives primarily from the outer coat. We suggest these interactions collectively contribute to coat organisation and membrane curvature, providing a structural framework to understand regulatory mechanisms of COPII trafficking and secretion. | |||||||||
| History |
|
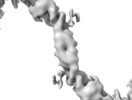
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11193.map.gz emd_11193.map.gz | 7.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11193-v30.xml emd-11193-v30.xml emd-11193.xml emd-11193.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11193_fsc.xml emd_11193_fsc.xml | 6 KB | Display |  FSC data file FSC data file |
| Images |  emd_11193.png emd_11193.png | 47.1 KB | ||
| Masks |  emd_11193_msk_1.map emd_11193_msk_1.map | 8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-11193.cif.gz emd-11193.cif.gz | 6.7 KB | ||
| Others |  emd_11193_half_map_1.map.gz emd_11193_half_map_1.map.gz emd_11193_half_map_2.map.gz emd_11193_half_map_2.map.gz | 4.2 MB 4.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11193 http://ftp.pdbj.org/pub/emdb/structures/EMD-11193 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11193 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11193 | HTTPS FTP |
-Related structure data
| Related structure data |  6zg5MC  6zg6C  6zgaC  6zl0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11193.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11193.map.gz / Format: CCP4 / Size: 8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.66 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11193_msk_1.map emd_11193_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_11193_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_11193_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : COPII coat assembled on lipid bilayer
| Entire | Name: COPII coat assembled on lipid bilayer |
|---|---|
| Components |
|
-Supramolecule #1: COPII coat assembled on lipid bilayer
| Supramolecule | Name: COPII coat assembled on lipid bilayer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Protein transport protein SEC31
| Macromolecule | Name: Protein transport protein SEC31 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 138.833422 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVKLAEFSRT ATFAWSHDKI PLLVSGTVSG TVDANFSTDS SLELWSLLAA DSEKPIASLQ VDSKFNDLDW SHNNKIIAGA LDNGSLELY STNEANNAIN SMARFSNHSS SVKTVKFNAK QDNVLASGGN NGEIFIWDMN KCTESPSNYT PLTPGQSMSS V DEVISLAW ...String: MVKLAEFSRT ATFAWSHDKI PLLVSGTVSG TVDANFSTDS SLELWSLLAA DSEKPIASLQ VDSKFNDLDW SHNNKIIAGA LDNGSLELY STNEANNAIN SMARFSNHSS SVKTVKFNAK QDNVLASGGN NGEIFIWDMN KCTESPSNYT PLTPGQSMSS V DEVISLAW NQSLAHVFAS AGSSNFASIW DLKAKKEVIH LSYTSPNSGI KQQLSVVEWH PKNSTRVATA TGSDNDPSIL IW DLRNANT PLQTLNQGHQ KGILSLDWCH QDEHLLLSSG RDNTVLLWNP ESAEQLSQFP ARGNWCFKTK FAPEAPDLFA CAS FDNKIE VQTLQNLTNT LDEQETETKQ QESETDFWNN VSREESKEKP SVFHLQAPTW YGEPSPAAHW AFGGKLVQIT PDGK GVSIT NPKISGLESN TTLSEALKTK DFKPLINQRL VKVIDDVNEE DWNLLEKLSM DGTEEFLKEA LAFDNDESDA QDDAN NEKE DDGEEFFQQI ETNFQPEGDF SLSGNIEQTI SKNLVSGNIK SAVKNSLEND LLMEAMVIAL DSNNERLKES VKNAYF AKY GSKSSLSRIL YSISKREVDD LVENLDVSQW KFISKAIQNL YPNDIAQRNE MLIKLGDRLK ENGHRQDSLT LYLAAGS LD KVASIWLSEF PDLEDKLKKD NKTIYEAHSE CLTEFIERFT VFSNFINGSS TINNEQLIAK FLEFINLTTS TGNFELAT E FLNSLPSDNE EVKTEKARVL IASGKSLPAQ NPATATTSKA KYTNAKTNKN VPVLPTPGMP STTSIPSMQA PFYGMTPGA SANALPPKPY VPATTTSAPV HTEGKYAPPS QPSMASPFVN KTNSSTRLNS FAPPPNPYAT ATVPATNVST TSIPQNTFAP IQPGMPIMG DYNAQSSSIP SQPPINAVSG QTPHLNRKAN DGWNDLPLKV KEKPSRAKAV SVAPPNILST PTPLNGIPAN A ASTMPPPP LSRAPSSVSM VSPPPLHKNS RVPSLVATSE SPRASISNPY APPQSSQQFP IGTISTANQT SNTAQVASSN PY APPPQQR VATPLSGGVP PAPLPKASNP YAPTATTQPN GSSYPPTGPY TNNHTMTSPP PVFNKPPTGP PPISMKKRSN KLA SIEQNP SQGATYPPTL SSSASPLQPS QPPTLASQVN TSAENVSHEI PADQQPIVDF LKEELARVTP LTPKEYSKQL KDCD KRLKI LFYHLEKQDL LTQPTIDCLH DLVALMKEKK YKEAMVIHAN IATNHAQEGG NWLTGVKRLI GIAEATLN UniProtKB: Protein transport protein SEC31 |
-Macromolecule #2: Protein transport protein SEC13
| Macromolecule | Name: Protein transport protein SEC13 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 33.082965 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVVIANAHNE LIHDAVLDYY GKRLATCSSD KTIKIFEVEG ETHKLIDTLT GHEGPVWRVD WAHPKFGTIL ASCSYDGKVL IWKEENGRW SQIAVHAVHS ASVNSVQWAP HEYGPLLLVA SSDGKVSVVE FKENGTTSPI IIDAHAIGVN SASWAPATIE E DGEHNGTK ...String: MVVIANAHNE LIHDAVLDYY GKRLATCSSD KTIKIFEVEG ETHKLIDTLT GHEGPVWRVD WAHPKFGTIL ASCSYDGKVL IWKEENGRW SQIAVHAVHS ASVNSVQWAP HEYGPLLLVA SSDGKVSVVE FKENGTTSPI IIDAHAIGVN SASWAPATIE E DGEHNGTK ESRKFVTGGA DNLVKIWKYN SDAQTYVLES TLEGHSDWVR DVAWSPTVLL RSYLASVSQD RTCIIWTQDN EQ GPWKKTL LKEEKFPDVL WRASWSLSGN VLALSGGDNK VTLWKENLEG KWEPAGEVHQ UniProtKB: Protein transport protein SEC13 |
-Macromolecule #3: Protein transport protein SEC13
| Macromolecule | Name: Protein transport protein SEC13 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 33.191188 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVVIANAHNE MIHDAVMDYY GKRMATCSSD KTIKIFEVEG ETHKLIDTLT GHEGPVWRVD WAHPKFGTIL ASCSYDGKVM IWKEENGRW SQIAVHAVHS ASVNSVQWAP HEYGPMLLVA SSDGKVSVVE FKENGTTSPI IIDAHAIGVN SASWAPATIE E DGEHNGTK ...String: MVVIANAHNE MIHDAVMDYY GKRMATCSSD KTIKIFEVEG ETHKLIDTLT GHEGPVWRVD WAHPKFGTIL ASCSYDGKVM IWKEENGRW SQIAVHAVHS ASVNSVQWAP HEYGPMLLVA SSDGKVSVVE FKENGTTSPI IIDAHAIGVN SASWAPATIE E DGEHNGTK ESRKFVTGGA DNLVKIWKYN SDAQTYVLES TLEGHSDWVR DVAWSPTVLL RSYMASVSQD RTCIIWTQDN EQ GPWKKTL LKEEKFPDVL WRASWSLSGN VLALSGGDNK VTLWKENLEG KWEPAGEVHQ UniProtKB: Protein transport protein SEC13 |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 185 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Buffer | pH: 6.8 |
|---|---|
| Grid | Details: C-FLAT GRIDS |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 3.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)