+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0800 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Abo1 structure with AAA+ and bromodomains in apo state | ||||||||||||||||||
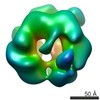
 Map data Map data | apo-Abo1 with bromodomain | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.9 Å | ||||||||||||||||||
 Authors Authors | Cho C / Jang J | ||||||||||||||||||
| Funding support |  Korea, Republic Of, 5 items Korea, Republic Of, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Structural basis of nucleosome assembly by the Abo1 AAA+ ATPase histone chaperone. Authors: Carol Cho / Juwon Jang / Yujin Kang / Hiroki Watanabe / Takayuki Uchihashi / Seung Joong Kim / Koichi Kato / Ja Yil Lee / Ji-Joon Song /   Abstract: The fundamental unit of chromatin, the nucleosome, is an intricate structure that requires histone chaperones for assembly. ATAD2 AAA+ ATPases are a family of histone chaperones that regulate ...The fundamental unit of chromatin, the nucleosome, is an intricate structure that requires histone chaperones for assembly. ATAD2 AAA+ ATPases are a family of histone chaperones that regulate nucleosome density and chromatin dynamics. Here, we demonstrate that the fission yeast ATAD2 homolog, Abo1, deposits histone H3-H4 onto DNA in an ATP-hydrolysis-dependent manner by in vitro reconstitution and single-tethered DNA curtain assays. We present cryo-EM structures of an ATAD2 family ATPase to atomic resolution in three different nucleotide states, revealing unique structural features required for histone loading on DNA, and directly visualize the transitions of Abo1 from an asymmetric spiral (ATP-state) to a symmetric ring (ADP- and apo-states) using high-speed atomic force microscopy (HS-AFM). Furthermore, we find that the acidic pore of ATP-Abo1 binds a peptide substrate which is suggestive of a histone tail. Based on these results, we propose a model whereby Abo1 facilitates H3-H4 loading by utilizing ATP. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0800.map.gz emd_0800.map.gz | 95.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0800-v30.xml emd-0800-v30.xml emd-0800.xml emd-0800.xml | 11.5 KB 11.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0800.png emd_0800.png | 67.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0800 http://ftp.pdbj.org/pub/emdb/structures/EMD-0800 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0800 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0800 | HTTPS FTP |
-Related structure data
| Related structure data |  9870C  9871C  9872C  6jpqC  6jpuC  6jq0C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0800.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0800.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | apo-Abo1 with bromodomain | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : apo-Abo1
| Entire | Name: apo-Abo1 |
|---|---|
| Components |
|
-Supramolecule #1: apo-Abo1
| Supramolecule | Name: apo-Abo1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Macromolecule #1: Abo1
| Macromolecule | Name: Abo1 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MKEEASEHGG SADETQELSP VSDSSDEMPN NAKRRRRSQS MIANKRIHQA FQEDEGDEDW EEEEHKPKA KRRYNTRSNE SFSEGDDEPF EVSESSALED ELSDSEDSFI RSVRSKPKYK P GTRRSTRL RNRRSQDEEE SEEEHRPILR ERTSRINYSV PLAFPPVDEM ...String: MKEEASEHGG SADETQELSP VSDSSDEMPN NAKRRRRSQS MIANKRIHQA FQEDEGDEDW EEEEHKPKA KRRYNTRSNE SFSEGDDEPF EVSESSALED ELSDSEDSFI RSVRSKPKYK P GTRRSTRL RNRRSQDEEE SEEEHRPILR ERTSRINYSV PLAFPPVDEM DGDPSSQVNQ SR SRKTHSE LAITKLLRQQ VSSFMPYIDS SGSESESDNT RIKKSSAKTI KALTDPANSG GPP DFGRIR EKSDLADSDP LGVDSSLSFE SVGGLDNYIN QLKEMVMLPL LYPEIFQRFN MQPP RGVLF HGPPGTGKTL MARALAAACS SENKKVSFYM RKGADCLSKW VGEAERQLRL LFEEA KSTQ PSIIFFDEID GLAPVRSSKQ EQIHASIVST LLALMDGMES RGQVIIIGAT NRPDAV DPA LRRPGRFDRE FYFPLPDRDA RKKIIEIHTR NWDPPVPEWL CSMLAEKSKG YGGADLR AL CTEAALNSIK RTYPQLYRST KRLQIDPKTI KVKVKDFVMS MKRMIPSSER SSISPSKP L SPELKPLLNE AFQDIEKTLQ KLMPVASKLN PLEEVMYDDP KENDFEYQQR LETFETLRI YKPRFLICGR KGLGQTALGP AILQQYEGVH VQSFDMSTLL QDSTQSIETS IIHLFLEVRR HTPSIIYIP DIDNWLNVLP LTAITTFSSM LERLDFSDQI LFLALSSSPL SELHPQLREW F SSKQSVYS LQYPTRDSII AFFQPILELI KASPTELPGG IPRKRRVLPE LPLAPDPPPF TS QKITLKQ TKQADMRLLN KLKIKLNALL GSLRARYRKF KKPLIDFNDI YCVDPETGHS YRS REECHY EFVDDVVKQI GSDQKFSMMS LEEIEKRTWD NCYCTPKQFV HDIKLILRDA LQLE DSETI KRAQEMYANV LLGVEDMEDD QFSQRCERMA LREAERRKLR HGKLQKHLDE TKADM QFTS EKPSVDESIT EVDDAIKDGP PVLAETLTNS LMEDVGPENV DMDIEDNEIF TNQSTM SVP SMLVENEESP KPDEYIDQKD KVQSPLLNGK SPVGVPSEAA LRVSTDVSTN ISSNGRA DI PVDTLITSPA DVPNNAPTDA HNITSADGHI ENIEQEVVFP DLVFDEDRLT PLKQLLID S TTGFTVDQLL HLHSFLYQII WNTKSEWNRN SVVDECERAV KEFMINALQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.015 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 6.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 23581 |
|---|---|
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















