[English] 日本語
 Yorodumi
Yorodumi- EMDB-0344: CryoEM structure of Leviviridae PP7 WT coat protein dimer capsid ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0344 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
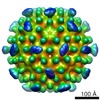
| Title | CryoEM structure of Leviviridae PP7 WT coat protein dimer capsid (PP7PP7-WT) | |||||||||
 Map data Map data | Final map autosharp | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | T4 / icosahedral / PP7 / biotechnology / vaccine / drug delivery / VIRUS LIKE PARTICLE | |||||||||
| Function / homology | Bacteriophage PP7, coat / Phage PP7 coat protein / Bacteriophage RNA-type, capsid / T=3 icosahedral viral capsid / RNA binding / identical protein binding / Capsid protein Function and homology information Function and homology information | |||||||||
| Biological species |  Pseudomonas phage PP7 (virus) Pseudomonas phage PP7 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Liangjun Z / Kopylov M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: ACS Nano / Year: 2019 Journal: ACS Nano / Year: 2019Title: Engineering the PP7 Virus Capsid as a Peptide Display Platform. Authors: Liangjun Zhao / Mykhailo Kopylov / Clinton S Potter / Bridget Carragher / M G Finn /  Abstract: As self-assembling polyvalent nanoscale structures that can tolerate substantial genetic and chemical modification, virus-like particles are useful in a variety of fields. Here we describe the ...As self-assembling polyvalent nanoscale structures that can tolerate substantial genetic and chemical modification, virus-like particles are useful in a variety of fields. Here we describe the genetic modification and structural characterization of the Leviviridae PP7 capsid protein as a platform for the presentation of functional polypeptides. This particle was shown to tolerate the display of sequences from 1 kDa (a cell penetrating peptide) to 14 kDa (the Fc-binding double Z-domain) on its exterior surface as C-terminal genetic fusions to the coat protein. In addition, a dimeric construct allowed the presentation of exogenous loops between capsid monomers and the simultaneous presentation of two different peptides at different positions on the icosahedral structure. The PP7 particle is thereby significantly more tolerant of these types of polypeptide additions than Qβ and MS2, the other Leviviridae-derived VLPs in common use. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0344.map.gz emd_0344.map.gz | 480.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0344-v30.xml emd-0344-v30.xml emd-0344.xml emd-0344.xml | 11.3 KB 11.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0344.png emd_0344.png | 197.8 KB | ||
| Masks |  emd_0344_msk_1.map emd_0344_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-0344.cif.gz emd-0344.cif.gz | 5.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0344 http://ftp.pdbj.org/pub/emdb/structures/EMD-0344 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0344 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0344 | HTTPS FTP |
-Related structure data
| Related structure data |  6n4vMC  0351C  0352C  0353C  0354C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0344.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0344.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Final map autosharp | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0733 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_0344_msk_1.map emd_0344_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Pseudomonas phage PP7
| Entire | Name:  Pseudomonas phage PP7 (virus) Pseudomonas phage PP7 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Pseudomonas phage PP7
| Supramolecule | Name: Pseudomonas phage PP7 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 12023 / Sci species name: Pseudomonas phage PP7 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: Yes |
|---|
-Macromolecule #1: Coat protein
| Macromolecule | Name: Coat protein / type: protein_or_peptide / ID: 1 / Number of copies: 120 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage PP7 (virus) Pseudomonas phage PP7 (virus) |
| Molecular weight | Theoretical: 28.144008 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SKTIVLSVGE ATRTLTEIQS TADRQIFEEK VGPLVGRLRL TASLRQNGAK TAYRVNLKLD QADVVDCSTS VCGELPKVRY TQVWSHDVT IVANSTEASR KSLYDLTKSL VATSQVEDLV VNLVPLGRAY GGSKTIVLSV GEATRTLTEI QSTADRQIFE E KVGPLVGR ...String: SKTIVLSVGE ATRTLTEIQS TADRQIFEEK VGPLVGRLRL TASLRQNGAK TAYRVNLKLD QADVVDCSTS VCGELPKVRY TQVWSHDVT IVANSTEASR KSLYDLTKSL VATSQVEDLV VNLVPLGRAY GGSKTIVLSV GEATRTLTEI QSTADRQIFE E KVGPLVGR LRLTASLRQN GAKTAYRVNL KLDQADVVDC STSVCGELPK VRYTQVWSHD VTIVANSTEA SRKSLYDLTK SL VATSQVE DLVVNLVPLG R UniProtKB: Capsid protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 35.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus min: 0.002 µm / Nominal magnification: 22500 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)