[English] 日本語
 Yorodumi
Yorodumi- SASDCB6: Nmul_A1745 protein from Nitrosospira multiformis, Northeast Struc... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDCB6 |
|---|---|
 Sample Sample | Nmul_A1745 protein from Nitrosospira multiformis, Northeast Structural Genomics Consortium Target NmR72
|
| Function / homology | Uncharacterized cysteine-rich protein YhjQ-like / Protein of unknown function DUF326 / Copper storage protein / Ferredoxin Function and homology information Function and homology information |
| Biological species |  Nitrosospira multiformis (strain ATCC 25196 / NCIMB 11849 / C 71) (bacteria) Nitrosospira multiformis (strain ATCC 25196 / NCIMB 11849 / C 71) (bacteria) |
 Citation Citation |  Journal: Biopolymers / Year: 2011 Journal: Biopolymers / Year: 2011Title: Small angle X-ray scattering as a complementary tool for high-throughput structural studies. Authors: Thomas D Grant / Joseph R Luft / Jennifer R Wolfley / Hiro Tsuruta / Anne Martel / Gaetano T Montelione / Edward H Snell /  Abstract: Structural crystallography and nuclear magnetic resonance (NMR) spectroscopy are the predominant techniques for understanding the biological world on a molecular level. Crystallography is constrained ...Structural crystallography and nuclear magnetic resonance (NMR) spectroscopy are the predominant techniques for understanding the biological world on a molecular level. Crystallography is constrained by the ability to form a crystal that diffracts well and NMR is constrained to smaller proteins. Although powerful techniques, they leave many soluble, purified structurally uncharacterized protein samples. Small angle X-ray scattering (SAXS) is a solution technique that provides data on the size and multiple conformations of a sample, and can be used to reconstruct a low-resolution molecular envelope of a macromolecule. In this study, SAXS has been used in a high-throughput manner on a subset of 28 proteins, where structural information is available from crystallographic and/or NMR techniques. These crystallographic and NMR structures were used to validate the accuracy of molecular envelopes reconstructed from SAXS data on a statistical level, to compare and highlight complementary structural information that SAXS provides, and to leverage biological information derived by crystallographers and spectroscopists from their structures. All the ab initio molecular envelopes calculated from the SAXS data agree well with the available structural information. SAXS is a powerful albeit low-resolution technique that can provide additional structural information in a high-throughput and complementary manner to improve the functional interpretation of high-resolution structures. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Models
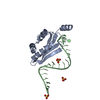
| Model #1409 |  Type: dummy / Radius of dummy atoms: 2.00 A / Chi-square value: 2.119936  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|---|
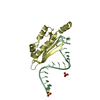
| Model #1416 |  Type: atomic / Radius of dummy atoms: 1.90 A / Chi-square value: 2.6569  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
- Sample
Sample
 Sample Sample | Name: Nmul_A1745 protein from Nitrosospira multiformis, Northeast Structural Genomics Consortium Target NmR72 Specimen concentration: 0.73-2.23 |
|---|---|
| Buffer | Name: 5 mM DTT 100 mM NaCl 10 mM Tris-HCl 0.02 % NaN3 / pH: 7.5 |
| Entity #745 | Type: protein / Description: Uncharacterized protein / Formula weight: 13.694 / Num. of mol.: 4 Source: Nitrosospira multiformis (strain ATCC 25196 / NCIMB 11849 / C 71) References: UniProt: Q2Y879 Sequence: MFLYTETDQN LQACIDACNH CYRTCLRMAM NHCLEAGGKH VEADHLRLMM NCAEICQTSL NFMLSGSRFS PKVCGVCAEI CDACAKSCEQ LDGMEECVQT CRQCAEHCRK MAALEHHHHH H |
-Experimental information
| Beam | Instrument name: Stanford Synchrotron Radiation Lightsource (SSRL) BL4-2 City: Stanford, CA / 国: USA  / Type of source: X-ray synchrotron / Wavelength: 0.13 Å / Dist. spec. to detc.: 1.5 mm / Type of source: X-ray synchrotron / Wavelength: 0.13 Å / Dist. spec. to detc.: 1.5 mm | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Rayonix MX225-HE | ||||||||||||||||||
| Scan | Measurement date: Feb 12, 2010 / Storage temperature: -80 °C / Cell temperature: 20 °C / Exposure time: 1 sec. / Number of frames: 20 / Unit: 1/A /
| ||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller

 SASDCB6
SASDCB6