+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 9ilp | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the bacteriophage T5 portal complex | |||||||||||||||||||||
 Components Components |
| |||||||||||||||||||||
 Keywords Keywords | VIRAL PROTEIN / Complex | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcytoplasmic icosahedral capsid assembly / symbiont genome ejection through host cell envelope, long flexible tail mechanism / viral capsid assembly / virion component / viral capsid / host cell cytoplasm Similarity search - Function | |||||||||||||||||||||
| Biological species |  Escherichia phage T5 (virus) Escherichia phage T5 (virus) | |||||||||||||||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||||||||||||||
 Authors Authors | Peng, Y.N. / Liu, H.R. | |||||||||||||||||||||
| Funding support |  China, 4items China, 4items
| |||||||||||||||||||||
 Citation Citation |  Journal: Int J Mol Sci / Year: 2024 Journal: Int J Mol Sci / Year: 2024Title: Structures of Mature and Urea-Treated Empty Bacteriophage T5: Insights into Siphophage Infection and DNA Ejection. Authors: Yuning Peng / Huanrong Tang / Hao Xiao / Wenyuan Chen / Jingdong Song / Jing Zheng / Hongrong Liu /  Abstract: T5 is a siphophage that has been extensively studied by structural and biochemical methods. However, the complete in situ structures of T5 before and after DNA ejection remain unknown. In this study, ...T5 is a siphophage that has been extensively studied by structural and biochemical methods. However, the complete in situ structures of T5 before and after DNA ejection remain unknown. In this study, we used cryo-electron microscopy (cryo-EM) to determine the structures of mature T5 (a laboratory-adapted, fiberless T5 mutant) and urea-treated empty T5 (lacking the tip complex) at near-atomic resolutions. Atomic models of the head, connector complex, tail tube, and tail tip were built for mature T5, and atomic models of the connector complex, comprising the portal protein pb7, adaptor protein p144, and tail terminator protein p142, were built for urea-treated empty T5. Our findings revealed that the aforementioned proteins did not undergo global conformational changes before and after DNA ejection, indicating that these structural features were conserved among most myophages and siphophages. The present study elucidates the underlying mechanisms of siphophage infection and DNA ejection. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  9ilp.cif.gz 9ilp.cif.gz | 1 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb9ilp.ent.gz pdb9ilp.ent.gz | 907.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  9ilp.json.gz 9ilp.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/il/9ilp https://data.pdbj.org/pub/pdb/validation_reports/il/9ilp ftp://data.pdbj.org/pub/pdb/validation_reports/il/9ilp ftp://data.pdbj.org/pub/pdb/validation_reports/il/9ilp | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  60672MC  8zviC  9ilvC 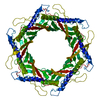 9imhC  9imvC  9inyC  9iozC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 19231.912 Da / Num. of mol.: 12 / Source method: isolated from a natural source / Source: (natural)  Escherichia phage T5 (virus) / References: UniProt: Q6QGD9 Escherichia phage T5 (virus) / References: UniProt: Q6QGD9#2: Protein | Mass: 45137.234 Da / Num. of mol.: 12 / Source method: isolated from a natural source / Source: (natural)  Escherichia phage T5 (virus) / References: UniProt: Q6QGD5 Escherichia phage T5 (virus) / References: UniProt: Q6QGD5Has protein modification | N | |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Escherichia phage T5 / Type: VIRUS / Entity ID: all / Source: NATURAL |
|---|---|
| Source (natural) | Organism:  Escherichia phage T5 (virus) Escherichia phage T5 (virus) |
| Details of virus | Empty: NO / Enveloped: NO / Isolate: SPECIES / Type: VIRION |
| Buffer solution | pH: 7.4 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal defocus max: 2200 nm / Nominal defocus min: 1600 nm |
| Image recording | Electron dose: 32 e/Å2 / Film or detector model: GATAN K3 (6k x 4k) |
- Processing
Processing
| EM software | Name: PHENIX / Category: model refinement | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Type: NONE | ||||||||||||||||||||||||
| 3D reconstruction | Resolution: 3.4 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 18398 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller









 PDBj
PDBj