+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8qa4 | ||||||
|---|---|---|---|---|---|---|---|
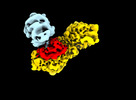
| Title | MTHFR + SAH symmetric dis-inhibited state | ||||||
 Components Components | Methylenetetrahydrofolate reductase (NADPH) | ||||||
 Keywords Keywords | FLAVOPROTEIN / Dis-inhibited / allosteric / folate / S-adenosylhomocysteine | ||||||
| Function / homology |  Function and homology information Function and homology informationmethylenetetrahydrofolate reductase (NADPH) / response to vitamin B2 / methylenetetrahydrofolate reductase (NADPH) activity / methylenetetrahydrofolate reductase (NAD(P)H) activity / methionine metabolic process / modified amino acid binding / homocysteine metabolic process / S-adenosylmethionine metabolic process / Metabolism of folate and pterines / methionine biosynthetic process ...methylenetetrahydrofolate reductase (NADPH) / response to vitamin B2 / methylenetetrahydrofolate reductase (NADPH) activity / methylenetetrahydrofolate reductase (NAD(P)H) activity / methionine metabolic process / modified amino acid binding / homocysteine metabolic process / S-adenosylmethionine metabolic process / Metabolism of folate and pterines / methionine biosynthetic process / response to folic acid / tetrahydrofolate interconversion / response to amino acid / heterochromatin organization / response to interleukin-1 / FAD binding / neural tube closure / NADP binding / flavin adenine dinucleotide binding / response to hypoxia / response to xenobiotic stimulus / protein-containing complex binding / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 2.8 Å | ||||||
 Authors Authors | Blomgren, L.K.M. / Yue, W.W. / Froese, D.S. / McCorvie, T.J. | ||||||
| Funding support |  Switzerland, 1items Switzerland, 1items
| ||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Dynamic inter-domain transformations mediate the allosteric regulation of human 5, 10-methylenetetrahydrofolate reductase. Authors: Linnea K M Blomgren / Melanie Huber / Sabrina R Mackinnon / Céline Bürer / Arnaud Baslé / Wyatt W Yue / D Sean Froese / Thomas J McCorvie /   Abstract: 5,10-methylenetetrahydrofolate reductase (MTHFR) commits folate-derived one-carbon units to generate the methyl-donor S-adenosyl-L-methionine (SAM). Eukaryotic MTHFR appends to the well-conserved ...5,10-methylenetetrahydrofolate reductase (MTHFR) commits folate-derived one-carbon units to generate the methyl-donor S-adenosyl-L-methionine (SAM). Eukaryotic MTHFR appends to the well-conserved catalytic domain (CD) a unique regulatory domain (RD) that confers feedback inhibition by SAM. Here we determine the cryo-electron microscopy structures of human MTHFR bound to SAM and its demethylated product S-adenosyl-L-homocysteine (SAH). In the active state, with the RD bound to a single SAH, the CD is flexible and exposes its active site for catalysis. However, in the inhibited state the RD pocket is remodelled, exposing a second SAM-binding site that was previously occluded. Dual-SAM bound MTHFR demonstrates a substantially rearranged inter-domain linker that reorients the CD, inserts a loop into the active site, positions Tyr404 to bind the cofactor FAD, and blocks substrate access. Our data therefore explain the long-distance regulatory mechanism of MTHFR inhibition, underpinned by the transition between dual-SAM and single-SAH binding in response to cellular methylation status. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8qa4.cif.gz 8qa4.cif.gz | 199.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8qa4.ent.gz pdb8qa4.ent.gz | 153.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8qa4.json.gz 8qa4.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  8qa4_validation.pdf.gz 8qa4_validation.pdf.gz | 1.6 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  8qa4_full_validation.pdf.gz 8qa4_full_validation.pdf.gz | 1.6 MB | Display | |
| Data in XML |  8qa4_validation.xml.gz 8qa4_validation.xml.gz | 36.7 KB | Display | |
| Data in CIF |  8qa4_validation.cif.gz 8qa4_validation.cif.gz | 49.3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qa/8qa4 https://data.pdbj.org/pub/pdb/validation_reports/qa/8qa4 ftp://data.pdbj.org/pub/pdb/validation_reports/qa/8qa4 ftp://data.pdbj.org/pub/pdb/validation_reports/qa/8qa4 | HTTPS FTP |
-Related structure data
| Related structure data | 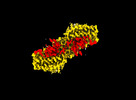 18298MC  8qa5C  8qa6C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 75461.195 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: MTHFR / Production host: Homo sapiens (human) / Gene: MTHFR / Production host:  #2: Chemical | #3: Water | ChemComp-HOH / | Has ligand of interest | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Human 5,10-methylenetetrahydrofolate reductase in complex with S-Adenosyl-L-homocysteine, regulatory domains Type: COMPLEX / Entity ID: #1 / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 0.15 MDa / Experimental value: NO |
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Source (recombinant) | Organism:  |
| Buffer solution | pH: 7.5 Details: 20 mM HEPES, pH 7.5, 150 mM NaCl, 0.0025% Tween20, 1 mM S-Adenosyl-L-homocysteine, filter sterilised |
| Specimen | Conc.: 2 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Specimen support | Grid material: GOLD / Grid mesh size: 300 divisions/in. / Grid type: UltrAuFoil R1.2/1.3 |
| Vitrification | Instrument: FEI VITROBOT MARK III / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 295 K |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: TFS GLACIOS |
|---|---|
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 240000 X / Nominal defocus max: 2100 nm / Nominal defocus min: 900 nm / Cs: 2.7 mm / C2 aperture diameter: 100 µm |
| Specimen holder | Cryogen: NITROGEN |
| Image recording | Average exposure time: 5.18 sec. / Electron dose: 50 e/Å2 / Film or detector model: FEI FALCON IV (4k x 4k) / Num. of grids imaged: 2 / Num. of real images: 5606 |
- Processing
Processing
| EM software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 1900000 | ||||||||||||||||||||||||||||||||||||||||
| Symmetry | Point symmetry: C2 (2 fold cyclic) | ||||||||||||||||||||||||||||||||||||||||
| 3D reconstruction | Resolution: 2.8 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 105879 / Algorithm: FOURIER SPACE / Symmetry type: POINT | ||||||||||||||||||||||||||||||||||||||||
| Atomic model building | B value: 90.1 / Protocol: FLEXIBLE FIT / Space: REAL / Target criteria: Cross-correlation coeficient | ||||||||||||||||||||||||||||||||||||||||
| Atomic model building | PDB-ID: 6fcx Pdb chain-ID: A / Accession code: 6fcx / Source name: PDB / Type: experimental model | ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller




 PDBj
PDBj




