[English] 日本語
 Yorodumi
Yorodumi- PDB-8jr1: Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Fo i... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8jr1 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Mycobacterium tuberculosis ATP synthase Fo in complex with TBAJ-587 | ||||||||||||
 Components Components |
| ||||||||||||
 Keywords Keywords | MEMBRANE PROTEIN / ATP synthase / Mycobacterium tuberculosis / cryo-EM | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationproton motive force-driven plasma membrane ATP synthesis / proton-transporting two-sector ATPase complex, proton-transporting domain / proton-transporting ATP synthase complex / proton-transporting ATP synthase activity, rotational mechanism / hydrolase activity / lipid binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3.17 Å | ||||||||||||
 Authors Authors | Zhang, Y. / Lai, Y. / Liu, F. / Rao, Z. / Gong, H. | ||||||||||||
| Funding support |  China, 3items China, 3items
| ||||||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Inhibition of M. tuberculosis and human ATP synthase by BDQ and TBAJ-587. Authors: Yuying Zhang / Yuezheng Lai / Shan Zhou / Ting Ran / Yue Zhang / Ziqing Zhao / Ziyan Feng / Long Yu / Jinxu Xu / Kun Shi / Jianyun Wang / Yu Pang / Liang Li / Hongming Chen / Luke W Guddat / ...Authors: Yuying Zhang / Yuezheng Lai / Shan Zhou / Ting Ran / Yue Zhang / Ziqing Zhao / Ziyan Feng / Long Yu / Jinxu Xu / Kun Shi / Jianyun Wang / Yu Pang / Liang Li / Hongming Chen / Luke W Guddat / Yan Gao / Fengjiang Liu / Zihe Rao / Hongri Gong /   Abstract: Bedaquiline (BDQ), a first-in-class diarylquinoline anti-tuberculosis drug, and its analogue, TBAJ-587, prevent the growth and proliferation of Mycobacterium tuberculosis by inhibiting ATP synthase. ...Bedaquiline (BDQ), a first-in-class diarylquinoline anti-tuberculosis drug, and its analogue, TBAJ-587, prevent the growth and proliferation of Mycobacterium tuberculosis by inhibiting ATP synthase. However, BDQ also inhibits human ATP synthase. At present, how these compounds interact with either M. tuberculosis ATP synthase or human ATP synthase is unclear. Here we present cryogenic electron microscopy structures of M. tuberculosis ATP synthase with and without BDQ and TBAJ-587 bound, and human ATP synthase bound to BDQ. The two inhibitors interact with subunit a and the c-ring at the leading site, c-only sites and lagging site in M. tuberculosis ATP synthase, showing that BDQ and TBAJ-587 have similar modes of action. The quinolinyl and dimethylamino units of the compounds make extensive contacts with the protein. The structure of human ATP synthase in complex with BDQ reveals that the BDQ-binding site is similar to that observed for the leading site in M. tuberculosis ATP synthase, and that the quinolinyl unit also interacts extensively with the human enzyme. This study will improve researchers' understanding of the similarities and differences between human ATP synthase and M. tuberculosis ATP synthase in terms of the mode of BDQ binding, and will allow the rational design of novel diarylquinolines as anti-tuberculosis drugs. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8jr1.cif.gz 8jr1.cif.gz | 161.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8jr1.ent.gz pdb8jr1.ent.gz | 132.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8jr1.json.gz 8jr1.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/jr/8jr1 https://data.pdbj.org/pub/pdb/validation_reports/jr/8jr1 ftp://data.pdbj.org/pub/pdb/validation_reports/jr/8jr1 ftp://data.pdbj.org/pub/pdb/validation_reports/jr/8jr1 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  36590MC  8j0sC  8j0tC 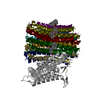 8j57C  8j58C  8jr0C  8khfC  8ki3C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 8058.423 Da / Num. of mol.: 9 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Mycolicibacterium smegmatis (bacteria) / References: UniProt: A0A045H4W8 Mycolicibacterium smegmatis (bacteria) / References: UniProt: A0A045H4W8#2: Protein | | Mass: 27488.436 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: atpB, ERS007657_00358, ERS007661_00092, ERS007663_00105, ERS007665_00910, ERS007670_00031, ERS007679_03316, ERS007681_03471, ERS007688_02939, ERS007703_00159, ERS007720_03212, ERS007722_00190, ...Gene: atpB, ERS007657_00358, ERS007661_00092, ERS007663_00105, ERS007665_00910, ERS007670_00031, ERS007679_03316, ERS007681_03471, ERS007688_02939, ERS007703_00159, ERS007720_03212, ERS007722_00190, ERS007739_02359, ERS007741_03568, ERS024276_02583, ERS027646_02991, ERS027659_00329, ERS027661_00021, SAMEA2683035_01568 Production host:  Mycolicibacterium smegmatis (bacteria) / References: UniProt: A0A045J1C5 Mycolicibacterium smegmatis (bacteria) / References: UniProt: A0A045J1C5#3: Chemical | ChemComp-UTI / ( Mass: 614.503 Da / Num. of mol.: 7 / Source method: obtained synthetically / Formula: C30H33BrFN3O5 Has ligand of interest | N | |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Mycobacterium tuberculosis ATP synthase / Type: COMPLEX / Entity ID: #1-#2 / Source: RECOMBINANT |
|---|---|
| Source (natural) | Organism:  |
| Source (recombinant) | Organism:  Mycolicibacterium smegmatis (bacteria) Mycolicibacterium smegmatis (bacteria) |
| Buffer solution | pH: 7.4 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal defocus max: 2400 nm / Nominal defocus min: 1200 nm |
| Image recording | Electron dose: 50 e/Å2 / Film or detector model: FEI FALCON IV (4k x 4k) |
- Processing
Processing
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION |
|---|---|
| 3D reconstruction | Resolution: 3.17 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 81528 / Symmetry type: POINT |
 Movie
Movie Controller
Controller

















 PDBj
PDBj

