[English] 日本語
 Yorodumi
Yorodumi- PDB-7z14: Cryo-EM structure of Torpedo nicotinic acetylcholine receptor in ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7z14 | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Torpedo nicotinic acetylcholine receptor in complex with a short-chain neurotoxin. | |||||||||||||||||||||
 Components Components |
| |||||||||||||||||||||
 Keywords Keywords | MEMBRANE PROTEIN / Ion channel / toxin | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationacetylcholine-gated monoatomic cation-selective channel activity / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / transmembrane signaling receptor activity / postsynaptic membrane / neuron projection Similarity search - Function | |||||||||||||||||||||
| Biological species | synthetic construct (others) | |||||||||||||||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||||||||||||||
 Authors Authors | Nys, M.A.E.M. / Zarkadas, E. / Ulens, C. / Nury, H. | |||||||||||||||||||||
| Funding support |  Belgium, Belgium,  United Kingdom, United Kingdom,  France, European Union, 6items France, European Union, 6items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: The molecular mechanism of snake short-chain α-neurotoxin binding to muscle-type nicotinic acetylcholine receptors. Authors: Mieke Nys / Eleftherios Zarkadas / Marijke Brams / Aujan Mehregan / Kumiko Kambara / Jeroen Kool / Nicholas R Casewell / Daniel Bertrand / John E Baenziger / Hugues Nury / Chris Ulens /       Abstract: Bites by elapid snakes (e.g. cobras) can result in life-threatening paralysis caused by venom neurotoxins blocking neuromuscular nicotinic acetylcholine receptors. Here, we determine the cryo-EM ...Bites by elapid snakes (e.g. cobras) can result in life-threatening paralysis caused by venom neurotoxins blocking neuromuscular nicotinic acetylcholine receptors. Here, we determine the cryo-EM structure of the muscle-type Torpedo receptor in complex with ScNtx, a recombinant short-chain α-neurotoxin. ScNtx is pinched between loop C on the principal subunit and a unique hairpin in loop F on the complementary subunit, thereby blocking access to the neurotransmitter binding site. ScNtx adopts a binding mode that is tilted toward the complementary subunit, forming a wider network of interactions than those seen in the long-chain α-Bungarotoxin complex. Certain mutations in ScNtx at the toxin-receptor interface eliminate inhibition of neuronal α7 nAChRs, but not of human muscle-type receptors. These observations explain why ScNtx binds more tightly to muscle-type receptors than neuronal receptors. Together, these data offer a framework for understanding subtype-specific actions of short-chain α-neurotoxins and inspire strategies for design of new snake antivenoms. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7z14.cif.gz 7z14.cif.gz | 360.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7z14.ent.gz pdb7z14.ent.gz | 291.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7z14.json.gz 7z14.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  7z14_validation.pdf.gz 7z14_validation.pdf.gz | 1.3 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  7z14_full_validation.pdf.gz 7z14_full_validation.pdf.gz | 1.3 MB | Display | |
| Data in XML |  7z14_validation.xml.gz 7z14_validation.xml.gz | 50.2 KB | Display | |
| Data in CIF |  7z14_validation.cif.gz 7z14_validation.cif.gz | 72.1 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/z1/7z14 https://data.pdbj.org/pub/pdb/validation_reports/z1/7z14 ftp://data.pdbj.org/pub/pdb/validation_reports/z1/7z14 ftp://data.pdbj.org/pub/pdb/validation_reports/z1/7z14 | HTTPS FTP |
-Related structure data
| Related structure data | 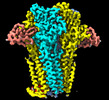 14440MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Acetylcholine receptor subunit ... , 4 types, 5 molecules ADBCE
| #1: Protein | Mass: 50168.164 Da / Num. of mol.: 2 / Source method: isolated from a natural source Source: (natural)  References: UniProt: P02710 #2: Protein | | Mass: 53731.773 Da / Num. of mol.: 1 / Source method: isolated from a natural source Source: (natural)  References: UniProt: P02712 #3: Protein | | Mass: 57625.711 Da / Num. of mol.: 1 / Source method: isolated from a natural source Source: (natural)  References: UniProt: P02718 #4: Protein | | Mass: 56335.684 Da / Num. of mol.: 1 / Source method: isolated from a natural source Source: (natural)  References: UniProt: P02714 |
|---|
-Protein , 1 types, 2 molecules FG
| #5: Protein | Mass: 6848.960 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Details: Consensus sequence from Elapidae family (8602) / Source: (gene. exp.) synthetic construct (others) / Production host:  Strain (production host): E. coli K-12 derived Origami 2 cells |
|---|
-Sugars , 4 types, 6 molecules 
| #6: Polysaccharide | Source method: isolated from a genetically manipulated source #7: Polysaccharide | Source method: isolated from a genetically manipulated source #8: Polysaccharide | alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1- ...alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | Type: oligosaccharide / Mass: 951.875 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source #9: Sugar | ChemComp-NAG / | |
|---|
-Details
| Has ligand of interest | N |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment |
 Movie
Movie Controller
Controller


 PDBj
PDBj

