[English] 日本語
 Yorodumi
Yorodumi- EMDB-14440: Cryo-EM structure of Torpedo nicotinic acetylcholine receptor in ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
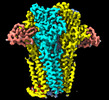
| Title | Cryo-EM structure of Torpedo nicotinic acetylcholine receptor in complex with a short-chain neurotoxin. | |||||||||||||||||||||
 Map data Map data | ||||||||||||||||||||||
 Sample Sample |
| |||||||||||||||||||||
 Keywords Keywords | Ion channel / toxin / membrane protein | |||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationacetylcholine-gated monoatomic cation-selective channel activity / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / transmembrane signaling receptor activity / postsynaptic membrane Similarity search - Function | |||||||||||||||||||||
| Biological species |  | |||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | |||||||||||||||||||||
 Authors Authors | Nys MAEM / Zarkadas E / Ulens C / Nury H | |||||||||||||||||||||
| Funding support |  Belgium, Belgium,  United Kingdom, United Kingdom,  France, European Union, 6 items France, European Union, 6 items
| |||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: The molecular mechanism of snake short-chain α-neurotoxin binding to muscle-type nicotinic acetylcholine receptors. Authors: Mieke Nys / Eleftherios Zarkadas / Marijke Brams / Aujan Mehregan / Kumiko Kambara / Jeroen Kool / Nicholas R Casewell / Daniel Bertrand / John E Baenziger / Hugues Nury / Chris Ulens /       Abstract: Bites by elapid snakes (e.g. cobras) can result in life-threatening paralysis caused by venom neurotoxins blocking neuromuscular nicotinic acetylcholine receptors. Here, we determine the cryo-EM ...Bites by elapid snakes (e.g. cobras) can result in life-threatening paralysis caused by venom neurotoxins blocking neuromuscular nicotinic acetylcholine receptors. Here, we determine the cryo-EM structure of the muscle-type Torpedo receptor in complex with ScNtx, a recombinant short-chain α-neurotoxin. ScNtx is pinched between loop C on the principal subunit and a unique hairpin in loop F on the complementary subunit, thereby blocking access to the neurotransmitter binding site. ScNtx adopts a binding mode that is tilted toward the complementary subunit, forming a wider network of interactions than those seen in the long-chain α-Bungarotoxin complex. Certain mutations in ScNtx at the toxin-receptor interface eliminate inhibition of neuronal α7 nAChRs, but not of human muscle-type receptors. These observations explain why ScNtx binds more tightly to muscle-type receptors than neuronal receptors. Together, these data offer a framework for understanding subtype-specific actions of short-chain α-neurotoxins and inspire strategies for design of new snake antivenoms. | |||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14440.map.gz emd_14440.map.gz | 59.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14440-v30.xml emd-14440-v30.xml emd-14440.xml emd-14440.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14440.png emd_14440.png | 130.2 KB | ||
| Filedesc metadata |  emd-14440.cif.gz emd-14440.cif.gz | 7.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14440 http://ftp.pdbj.org/pub/emdb/structures/EMD-14440 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14440 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14440 | HTTPS FTP |
-Related structure data
| Related structure data |  7z14MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14440.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14440.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.145 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Cryo-EM structure of Torpedo nicotinic acetylcholine receptor in ...
| Entire | Name: Cryo-EM structure of Torpedo nicotinic acetylcholine receptor in complex with a short-chain neurotoxin. |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of Torpedo nicotinic acetylcholine receptor in ...
| Supramolecule | Name: Cryo-EM structure of Torpedo nicotinic acetylcholine receptor in complex with a short-chain neurotoxin. type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1, #3, #2, #4-#5 |
|---|
-Macromolecule #1: Acetylcholine receptor subunit alpha
| Macromolecule | Name: Acetylcholine receptor subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 50.168164 KDa |
| Sequence | String: SEHETRLVAN LLENYNKVIR PVEHHTHFVD ITVGLQLIQL ISVDEVNQIV ETNVRLRQQW IDVRLRWNPA DYGGIKKIRL PSDDVWLPD LVLYNNADGD FAIVHMTKLL LDYTGKIMWT PPAIFKSYCE IIVTHFPFDQ QNCTMKLGIW TYDGTKVSIS P ESDRPDLS ...String: SEHETRLVAN LLENYNKVIR PVEHHTHFVD ITVGLQLIQL ISVDEVNQIV ETNVRLRQQW IDVRLRWNPA DYGGIKKIRL PSDDVWLPD LVLYNNADGD FAIVHMTKLL LDYTGKIMWT PPAIFKSYCE IIVTHFPFDQ QNCTMKLGIW TYDGTKVSIS P ESDRPDLS TFMESGEWVM KDYRGWKHWV YYTCCPDTPY LDITYHFIMQ RIPLYFVVNV IIPCLLFSFL TGLVFYLPTD SG EKMTLSI SVLLSLTVFL LVIVELIPST SSAVPLIGKY MLFTMIFVIS SIIITVVVIN THHRSPSTHT MPQWVRKIFI DTI PNVMFF STMKRASKEK QENKIFADDI DISDISGKQV TGEVIFQTPL IKNPDVKSAI EGVKYIAEHM KSDEESSNAA EEWK YVAMV IDHILLCVFM LICIIGTVSV FAGRLIELSQ EG UniProtKB: Acetylcholine receptor subunit alpha |
-Macromolecule #2: Acetylcholine receptor subunit beta
| Macromolecule | Name: Acetylcholine receptor subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 53.731773 KDa |
| Sequence | String: SVMEDTLLSV LFETYNPKVR PAQTVGDKVT VRVGLTLTNL LILNEKIEEM TTNVFLNLAW TDYRLQWDPA AYEGIKDLRI PSSDVWQPD IVLMNNNDGS FEITLHVNVL VQHTGAVSWQ PSAIYRSSCT IKVMYFPFDW QNCTMVFKSY TYDTSEVTLQ H ALDAKGER ...String: SVMEDTLLSV LFETYNPKVR PAQTVGDKVT VRVGLTLTNL LILNEKIEEM TTNVFLNLAW TDYRLQWDPA AYEGIKDLRI PSSDVWQPD IVLMNNNDGS FEITLHVNVL VQHTGAVSWQ PSAIYRSSCT IKVMYFPFDW QNCTMVFKSY TYDTSEVTLQ H ALDAKGER EVKEIVINKD AFTENGQWSI EHKPSRKNWR SDDPSYEDVT FYLIIQRKPL FYIVYTIIPC ILISILAILV FY LPPDAGE KMSLSISALL AVTVFLLLLA DKVPETSLSV PIIIRYLMFI MILVAFSVIL SVVVLNLHHR SPNTHTMPNW IRQ IFIETL PPFLWIQRPV TTPSPDSKPT IISRANDEYF IRKPAGDFVC PVDNARVAVQ PERLFSEMKW HLNGLTQPVT LPQD LKEAV EAIKYIAEQL ESASEFDDLK KDWQYVAMVA DRLFLYVFFV ICSIGTFSIF LDASHNVPPD NPFA UniProtKB: Acetylcholine receptor subunit beta |
-Macromolecule #3: Acetylcholine receptor subunit delta
| Macromolecule | Name: Acetylcholine receptor subunit delta / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 57.625711 KDa |
| Sequence | String: VNEEERLIND LLIVNKYNKH VRPVKHNNEV VNIALSLTLS NLISLKETDE TLTSNVWMDH AWYDHRLTWN ASEYSDISIL RLPPELVWI PDIVLQNNND GQYHVAYFCN VLVRPNGYVT WLPPAIFRSS CPINVLYFPF DWQNCSLKFT ALNYDANEIT M DLMTDTID ...String: VNEEERLIND LLIVNKYNKH VRPVKHNNEV VNIALSLTLS NLISLKETDE TLTSNVWMDH AWYDHRLTWN ASEYSDISIL RLPPELVWI PDIVLQNNND GQYHVAYFCN VLVRPNGYVT WLPPAIFRSS CPINVLYFPF DWQNCSLKFT ALNYDANEIT M DLMTDTID GKDYPIEWII IDPEAFTENG EWEIIHKPAK KNIYPDKFPN GTNYQDVTFY LIIRRKPLFY VINFITPCVL IS FLASLAF YLPAESGEKM STAISVLLAQ AVFLLLTSQR LPETALAVPL IGKYLMFIMS LVTGVIVNCG IVLNFHFRTP STH VLSTRV KQIFLEKLPR ILHMSRADES EQPDWQNDLK LRRSSSVGYI SKAQEYFNIK SRSELMFEKQ SERHGLVPRV TPRI GFGNN NENIAASDQL HDEIKSGIDS TNYIVKQIKE KNAYDEEVGN WNLVGQTIDR LSMFIITPVM VLGTIFIFVM GNFNH PPAK PFEGDPFDYS SDHPRCA UniProtKB: Acetylcholine receptor subunit delta |
-Macromolecule #4: Acetylcholine receptor subunit gamma
| Macromolecule | Name: Acetylcholine receptor subunit gamma / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 56.335684 KDa |
| Sequence | String: ENEEGRLIEK LLGDYDKRII PAKTLDHIID VTLKLTLTNL ISLNEKEEAL TTNVWIEIQW NDYRLSWNTS EYEGIDLVRI PSELLWLPD VVLENNVDGQ FEVAYYANVL VYNDGSMYWL PPAIYRSTCP IAVTYFPFDW QNCSLVFRSQ TYNAHEVNLQ L SAEEGEAV ...String: ENEEGRLIEK LLGDYDKRII PAKTLDHIID VTLKLTLTNL ISLNEKEEAL TTNVWIEIQW NDYRLSWNTS EYEGIDLVRI PSELLWLPD VVLENNVDGQ FEVAYYANVL VYNDGSMYWL PPAIYRSTCP IAVTYFPFDW QNCSLVFRSQ TYNAHEVNLQ L SAEEGEAV EWIHIDPEDF TENGEWTIRH RPAKKNYNWQ LTKDDTDFQE IIFFLIIQRK PLFYIINIIA PCVLISSLVV LV YFLPAQA GGQKCTLSIS VLLAQTIFLF LIAQKVPETS LNVPLIGKYL IFVMFVSMLI VMNCVIVLNV SLRTPNTHSL SEK IKHLFL GFLPKYLGMQ LEPSEETPEK PQPRRRSSFG IMIKAEEYIL KKPRSELMFE EQKDRHGLKR VNKMTSDIDI GTTV DLYKD LANFAPEIKS CVEACNFIAK STKEQNDSGS ENENWVLIGK VIDKACFWIA LLLFSIGTLA IFLTGHFNQV PEFPF PGDP RKYVP UniProtKB: Acetylcholine receptor subunit gamma |
-Macromolecule #5: Consensus short-chain short-chain alpha-neurotoxin ScNtx
| Macromolecule | Name: Consensus short-chain short-chain alpha-neurotoxin ScNtx type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 6.84896 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MICYNQQSSQ PPTTKTCSET SCYKKTWRDH RGTIIERGCG CPKVKPGIKL HCCRTDKCNN |
-Macromolecule #9: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 9 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.7000000000000001 µm |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.15 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 26581 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















