+ Open data
Open data
- Basic information
Basic information
| Entry | 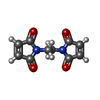 Database: PDB chemical components / ID: ME7 Database: PDB chemical components / ID: ME7 |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||
|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: ME7 / Ideal coordinates details: Corina / Model coordinates PDB-ID: 3QVY | ||||
| History |
| ||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  ChemicalBook / ChemicalBook /  CompTox / CompTox /  Nikkaji / Nikkaji /  PubChem / PubChem /  PubChem_TPharma / PubChem_TPharma /  ZINC / ZINC /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 12.01 | | CACTVS 3.370 | OpenEye OEToolkits 1.7.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.370 | | OpenEye OEToolkits 1.7.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 12.01 | | OpenEye OEToolkits 1.7.0 | |
|---|
-PDB entries
Showing all 5 items

PDB-3qvy: 
Crystal structure of the Zn-RIDC1 complex stabilized by BMOE crosslinks

PDB-3qvz: 
Crystal structure of the Zn-RIDC1 complex stabilized by BMOE crosslinks cocrystallized in the presence of Cu(II)

PDB-7ojx: 
E2 UBE2K covalently linked to donor Ub, acceptor di-Ub, and RING E3 primed for K48-linked Ub chain synthesis

PDB-7qy3: 
Crystal structure of the halohydrin dehalogenase HheG D114C mutant cross-linked with BMOE

PDB-9ff7: 
Structure of the BMOE-crosslinked transcription termination factor Rho in the presence of ppGpp; S84C/M405C double mutant
 Movie
Movie Controller
Controller



