[English] 日本語
 Yorodumi
Yorodumi- EMDB-9382: A high-resolution cryo-electron microscopy structure of a calcito... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9382 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | A high-resolution cryo-electron microscopy structure of a calcitonin receptor-heterotrimeric Gs protein complex | |||||||||||||||
 Map data Map data | Calcitonin receptor-heterotrimeric Gs protein complex, Sharpened map (B -80) | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | GPCR / transmembrane / receptor / calcitonin / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationcalcitonin receptor binding / calcitonin binding / amylin receptor complex 1 / amylin receptor complex 2 / cross-receptor inhibition within G protein-coupled receptor heterodimer / amylin receptor complex 3 / amylin receptor activity / calcitonin receptor activity / calcitonin gene-related peptide receptor activity / amylin receptor signaling pathway ...calcitonin receptor binding / calcitonin binding / amylin receptor complex 1 / amylin receptor complex 2 / cross-receptor inhibition within G protein-coupled receptor heterodimer / amylin receptor complex 3 / amylin receptor activity / calcitonin receptor activity / calcitonin gene-related peptide receptor activity / amylin receptor signaling pathway / positive regulation of adenylate cyclase activity / Calcitonin-like ligand receptors / negative regulation of ossification / positive regulation of protein kinase A signaling / response to amyloid-beta / response to glucocorticoid / regulation of mRNA stability / positive regulation of calcium-mediated signaling / ossification / acrosomal vesicle / osteoclast differentiation / adenylate cyclase-activating G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / hormone activity / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Glucagon signaling in metabolic regulation / cilium / G protein-coupled acetylcholine receptor signaling pathway / G-protein activation / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G beta:gamma signalling through CDC42 / Vasopressin regulates renal water homeostasis via Aquaporins / G beta:gamma signalling through BTK / ADP signalling through P2Y purinoceptor 12 / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / Glucagon-type ligand receptors / Sensory perception of sweet, bitter, and umami (glutamate) taste / photoreceptor disc membrane / G alpha (z) signalling events / Adrenaline,noradrenaline inhibits insulin secretion / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / ADORA2B mediated anti-inflammatory cytokines production / cellular response to catecholamine stimulus / sensory perception of taste / ADP signalling through P2Y purinoceptor 1 / adenylate cyclase-activating dopamine receptor signaling pathway / G beta:gamma signalling through PI3Kgamma / GPER1 signaling / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / cellular response to prostaglandin E stimulus / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / heterotrimeric G-protein complex / G alpha (12/13) signalling events / extracellular vesicle / signaling receptor complex adaptor activity / Thrombin signalling through proteinase activated receptors (PARs) / GTPase binding / retina development in camera-type eye / Ca2+ pathway / amyloid-beta binding / phospholipase C-activating G protein-coupled receptor signaling pathway / positive regulation of cytosolic calcium ion concentration / G alpha (i) signalling events / fibroblast proliferation / G alpha (s) signalling events / G alpha (q) signalling events / Ras protein signal transduction / cell population proliferation / Extra-nuclear estrogen signaling / positive regulation of ERK1 and ERK2 cascade / positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / cell surface receptor signaling pathway / G protein-coupled receptor signaling pathway / lysosomal membrane / axon / GTPase activity / synapse / protein-containing complex binding / positive regulation of gene expression / signal transduction / extracellular space / extracellular exosome / membrane / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   | |||||||||||||||
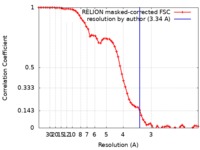
| Method | single particle reconstruction / cryo EM / Resolution: 3.34 Å | |||||||||||||||
 Authors Authors | dal Maso E / Glukhova A | |||||||||||||||
| Funding support |  Australia, 4 items Australia, 4 items
| |||||||||||||||
 Citation Citation |  Journal: ACS Pharmacol Transl Sci / Year: 2019 Journal: ACS Pharmacol Transl Sci / Year: 2019Title: The Molecular Control of Calcitonin Receptor Signaling. Authors: Emma Dal Maso / Alisa Glukhova / Yue Zhu / Javier Garcia-Nafria / Christopher G Tate / Silvia Atanasio / Christopher A Reynolds / Erney Ramírez-Aportela / Jose-Maria Carazo / Caroline A ...Authors: Emma Dal Maso / Alisa Glukhova / Yue Zhu / Javier Garcia-Nafria / Christopher G Tate / Silvia Atanasio / Christopher A Reynolds / Erney Ramírez-Aportela / Jose-Maria Carazo / Caroline A Hick / Sebastian G B Furness / Debbie L Hay / Yi-Lynn Liang / Laurence J Miller / Arthur Christopoulos / Ming-Wei Wang / Denise Wootten / Patrick M Sexton /       Abstract: The calcitonin receptor (CTR) is a class B G protein-coupled receptor (GPCR) that responds to the peptide hormone calcitonin (CT). CTs are clinically approved for the treatment of bone diseases. We ...The calcitonin receptor (CTR) is a class B G protein-coupled receptor (GPCR) that responds to the peptide hormone calcitonin (CT). CTs are clinically approved for the treatment of bone diseases. We previously reported a 4.1 Å structure of the activated CTR bound to salmon CT (sCT) and heterotrimeric Gs protein by cryo-electron microscopy (Liang, Y.-L., . Phase-plate cryo- EM structure of a class B GPCR-G protein complex. , , 118-123). In the current study, we have reprocessed the electron micrographs to yield a 3.3 Å map of the complex. This has allowed us to model extracellular loops (ECLs) 2 and 3, and the peptide N-terminus that previously could not be resolved. We have also performed alanine scanning mutagenesis of ECL1 and the upper segment of transmembrane helix 1 (TM1) and its extension into the receptor extracellular domain (TM1 stalk), with effects on peptide binding and function assessed by cAMP accumulation and ERK1/2 phosphorylation. These data were combined with previously published alanine scanning mutagenesis of ECL2 and ECL3 and the new structural information to provide a comprehensive 3D map of the molecular surface of the CTR that controls binding and signaling of distinct CT and related peptides. The work highlights distinctions in how different, related, class B receptors may be activated. The new mutational data on the TM1 stalk and ECL1 have also provided critical insights into the divergent control of cAMP versus pERK signaling and, collectively with previous mutagenesis data, offer evidence that the conformations linked to these different signaling pathways are, in many ways, mutually exclusive. This study furthers our understanding of the complex nature of signaling elicited by GPCRs and, in particular, that of the therapeutically important class B subfamily. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9382.map.gz emd_9382.map.gz | 2.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9382-v30.xml emd-9382-v30.xml emd-9382.xml emd-9382.xml | 28.3 KB 28.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9382_fsc.xml emd_9382_fsc.xml | 6.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_9382.png emd_9382.png | 106.7 KB | ||
| Masks |  emd_9382_msk_1.map emd_9382_msk_1.map | 22.2 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-9382.cif.gz emd-9382.cif.gz | 7.6 KB | ||
| Others |  emd_9382_additional_1.map.gz emd_9382_additional_1.map.gz emd_9382_additional_2.map.gz emd_9382_additional_2.map.gz emd_9382_half_map_1.map.gz emd_9382_half_map_1.map.gz emd_9382_half_map_2.map.gz emd_9382_half_map_2.map.gz | 20.9 MB 16.9 MB 17 MB 17 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9382 http://ftp.pdbj.org/pub/emdb/structures/EMD-9382 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9382 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9382 | HTTPS FTP |
-Validation report
| Summary document |  emd_9382_validation.pdf.gz emd_9382_validation.pdf.gz | 713 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9382_full_validation.pdf.gz emd_9382_full_validation.pdf.gz | 712.6 KB | Display | |
| Data in XML |  emd_9382_validation.xml.gz emd_9382_validation.xml.gz | 12 KB | Display | |
| Data in CIF |  emd_9382_validation.cif.gz emd_9382_validation.cif.gz | 16.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9382 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9382 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9382 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9382 | HTTPS FTP |
-Related structure data
| Related structure data |  6niyMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9382.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9382.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Calcitonin receptor-heterotrimeric Gs protein complex, Sharpened map (B -80) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_9382_msk_1.map emd_9382_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Calcitonin receptor-heterotrimeric Gs protein complex, LocalDeblur sharpened map...
| File | emd_9382_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Calcitonin receptor-heterotrimeric Gs protein complex, LocalDeblur sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Calcitonin receptor-heterotrimeric Gs protein complex, unsharpened map...
| File | emd_9382_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Calcitonin receptor-heterotrimeric Gs protein complex, unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Calcitonin receptor-heterotrimeric Gs protein complex, final half map...
| File | emd_9382_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Calcitonin receptor-heterotrimeric Gs protein complex, final half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Calcitonin receptor-heterotrimeric Gs protein complex, final half map...
| File | emd_9382_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Calcitonin receptor-heterotrimeric Gs protein complex, final half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of a full-length, active-state calcitonin receptor with p...
| Entire | Name: Complex of a full-length, active-state calcitonin receptor with peptide ligand, heterotrimeric Gs protein and nanobody 35 |
|---|---|
| Components |
|
-Supramolecule #1: Complex of a full-length, active-state calcitonin receptor with p...
| Supramolecule | Name: Complex of a full-length, active-state calcitonin receptor with peptide ligand, heterotrimeric Gs protein and nanobody 35 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 150 KDa |
-Macromolecule #1: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 45.72552 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGAGE SGKSTIVKQM RILHVNGFNG EGGEEDPQAA RSNSDGEKA TKVQDIKNNL KEAIETIVAA MSNLVPPVEL ANPENQFRVD YILSVMNVPD FDFPPEFYEH AKALWEDEGV R ACYERSNE ...String: MGCLGNSKTE DQRNEEKAQR EANKKIEKQL QKDKQVYRAT HRLLLLGAGE SGKSTIVKQM RILHVNGFNG EGGEEDPQAA RSNSDGEKA TKVQDIKNNL KEAIETIVAA MSNLVPPVEL ANPENQFRVD YILSVMNVPD FDFPPEFYEH AKALWEDEGV R ACYERSNE YQLIDCAQYF LDKIDVIKQA DYVPSDQDLL RCRVLTSGIF ETKFQVDKVN FHMFDVGGQR DERRKWIQCF ND VTAIIFV VASSSYNMVI REDNQTNRLQ EALNLFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK IEDYFPEFAR YTT PEDATP EPGEDPRVTR AKYFIRDEFL RISTASGDGR HYCYPHFTCA VDTENIRRVF NDCRDIIQRM HLRQYELL UniProtKB: UNIPROTKB: P63092 |
-Macromolecule #2: Nanobody35
| Macromolecule | Name: Nanobody35 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.140742 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QVQLQESGGG LVQPGGSLRL SCAASGFTFS NYKMNWVRQA PGKGLEWVSD ISQSGASISY TGSVKGRFTI SRDNAKNTLY LQMNSLKPE DTAVYYCARC PAPFTRDCFD VTSTTYAYRG QGTQVTVSSH HHHHHEPEA |
-Macromolecule #3: Calcitonin receptor
| Macromolecule | Name: Calcitonin receptor / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.387316 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MRFTFTSRCL ALFLLLNHPT PILPAFSNQT YPTIEPKPFL YVVGRKKMMD AQYKCYDRMQ QLPAYQGEGP YCNRTWDGWL CWDDTPAGV LSYQFCPDYF PDFDPSEKVT KYCDEKGVWF KHPENNRTWS NYTMCNAFTP EKLKNAYVLY YLAIVGHSLS I FTLVISLG ...String: MRFTFTSRCL ALFLLLNHPT PILPAFSNQT YPTIEPKPFL YVVGRKKMMD AQYKCYDRMQ QLPAYQGEGP YCNRTWDGWL CWDDTPAGV LSYQFCPDYF PDFDPSEKVT KYCDEKGVWF KHPENNRTWS NYTMCNAFTP EKLKNAYVLY YLAIVGHSLS I FTLVISLG IFVFFRSLGC QRVTLHKNMF LTYILNSMII IIHLVEVVPN GELVRRDPVS CKILHFFHQY MMACNYFWML CE GIYLHTL IVVAVFTEKQ RLRWYYLLGW GFPLVPTTIH AITRAVYFND NCWLSVETHL LYIIHGPVMA ALVVNFFFLL NIV RVLVTK MRETHEAESH MYLKAVKATM ILVPLLGIQF VVFPWRPSNK MLGKIYDYVM HSLIHFQGFF VATIYCFCNN EVQT TVKRQ WAQFKIQWNQ RWGRRPSNRS ARAAAAAAEA GDIPIYICHQ EPRNEPANNQ GEESAEIIPL NIIEQESSA UniProtKB: Calcitonin receptor |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.41693 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL ...String: MSELDQLRQE AEQLKNQIRD ARKACADATL SQITNNIDPV GRIQMRTRRT LRGHLAKIYA MHWGTDSRLL VSASQDGKLI IWDSYTTNK VHAIPLRSSW VMTCAYAPSG NYVACGGLDN ICSIYNLKTR EGNVRVSREL AGHTGYLSCC RFLDDNQIVT S SGDTTCAL WDIETGQQTT TFTGHTGDVM SLSLAPDTRL FVSGACDASA KLWDVREGMC RQTFTGHESD INAICFFPNG NA FATGSDD ATCRLFDLRA DQELMTYSHD NIICGITSVS FSKSGRLLLA GYDDFNCNVW DALKADRAGV LAGHDNRVSC LGV TDDGMA VATGSWDSFL KIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #5: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #6: Calcitonin
| Macromolecule | Name: Calcitonin / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 3.438886 KDa |
| Sequence | String: CSNLSTCVLG KLSQELHKLQ TYPRTNTGSG TP UniProtKB: Calcitonin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: OTHER |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 47170 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)