[English] 日本語
 Yorodumi
Yorodumi- EMDB-8936: Bundibugyo virus GP (mucin-deleted) in complex with pan-ebolaviru... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8936 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Bundibugyo virus GP (mucin-deleted) in complex with pan-ebolavirus human antibody ADI-15878 Fab | |||||||||
 Map data Map data | Bundibugyo virus glycoprotein (mucin-deleted) ectodomain bound to human survivor antibody ADI-15878 Fab | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | antibody / pan-ebolavirus / internal fusion loop / HR1 / glycoprotein / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated-mediated suppression of host tetherin activity / clathrin-dependent endocytosis of virus by host cell / entry receptor-mediated virion attachment to host cell / symbiont-mediated suppression of host innate immune response / fusion of virus membrane with host endosome membrane / viral envelope / lipid binding / host cell plasma membrane / virion membrane / extracellular region Similarity search - Function | |||||||||
| Biological species |   Bundibugyo virus / Bundibugyo virus /  Homo sapiens (human) / Homo sapiens (human) /  Bundibugyo ebolavirus Bundibugyo ebolavirus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.29 Å | |||||||||
 Authors Authors | Murin CD / Ward AB | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2018 Journal: Cell Rep / Year: 2018Title: Structural Basis of Pan-Ebolavirus Neutralization by an Antibody Targeting the Glycoprotein Fusion Loop. Authors: Charles D Murin / Jessica F Bruhn / Zachary A Bornholdt / Jeffrey Copps / Robyn Stanfield / Andrew B Ward /  Abstract: Monoclonal antibodies (mAbs) with pan-ebolavirus cross-reactivity are highly desirable, but development of such mAbs is limited by a lack of a molecular understanding of cross-reactive epitopes. The ...Monoclonal antibodies (mAbs) with pan-ebolavirus cross-reactivity are highly desirable, but development of such mAbs is limited by a lack of a molecular understanding of cross-reactive epitopes. The antibody ADI-15878 was previously identified from a human survivor of Ebola virus Makona variant (EBOV/Mak) infection. This mAb demonstrated potent neutralizing activity against all known ebolaviruses and provided protection in rodent and ferret models against three ebolavirus species. Here, we describe the unliganded crystal structure of ADI-15878 as well as the cryo-EM structures of ADI-15878 in complex with the EBOV/Mak and Bundibugyo virus (BDBV) glycoproteins (GPs). ADI-15878 binds through an induced-fit mechanism by targeting highly conserved residues in the internal fusion loop (IFL), bridging across GP protomers via the heptad repeat 1 (HR1) region. Our structures provide a more complete description of the ebolavirus immunogenic landscape, as well as a molecular basis for how rare but potent antibodies target conserved filoviral fusion machinery. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8936.map.gz emd_8936.map.gz | 7.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8936-v30.xml emd-8936-v30.xml emd-8936.xml emd-8936.xml | 25.8 KB 25.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8936_fsc.xml emd_8936_fsc.xml | 11.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_8936.png emd_8936.png | 66.4 KB | ||
| Filedesc metadata |  emd-8936.cif.gz emd-8936.cif.gz | 8.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8936 http://ftp.pdbj.org/pub/emdb/structures/EMD-8936 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8936 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8936 | HTTPS FTP |
-Related structure data
| Related structure data |  6dzmMC  8935C  6dzlC  6dznC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8936.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8936.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Bundibugyo virus glycoprotein (mucin-deleted) ectodomain bound to human survivor antibody ADI-15878 Fab | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
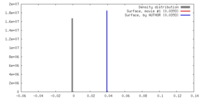
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bundibugyo virus glycoprotein (mucin-deleted) ectodomain bound to...
| Entire | Name: Bundibugyo virus glycoprotein (mucin-deleted) ectodomain bound to human survivor antibody ADI-15878 Fab |
|---|---|
| Components |
|
-Supramolecule #1: Bundibugyo virus glycoprotein (mucin-deleted) ectodomain bound to...
| Supramolecule | Name: Bundibugyo virus glycoprotein (mucin-deleted) ectodomain bound to human survivor antibody ADI-15878 Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: #3-#4 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 45 KDa |
-Supramolecule #2: Bundibugyo virus glycoprotein (mucin-deleted) ectodomain
| Supramolecule | Name: Bundibugyo virus glycoprotein (mucin-deleted) ectodomain type: complex / ID: 2 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Bundibugyo virus Bundibugyo virus |
-Supramolecule #3: Human survivor antibody ADI-15878 Fab
| Supramolecule | Name: Human survivor antibody ADI-15878 Fab / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3-#4 / Details: Fab was generated by papain digestion of IgG |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Bundibugyo virus GP1
| Macromolecule | Name: Bundibugyo virus GP1 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Bundibugyo ebolavirus Bundibugyo ebolavirus |
| Molecular weight | Theoretical: 34.697938 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: IPLGVVHNNT LQVSDIDKLV CRDKLSSTSQ LKSVGLNLEG NGVATDVPTA TKRWGFRAGV PPKVVNYEAG EWAENCYNLD IKKADGSEC LPEAPEGVRG FPRCRYVHKV SGTGPCPEGY AFHKEGAFFL YDRLASTIIY RSTTFSEGVV AFLILPETKK D FFQSPPLH ...String: IPLGVVHNNT LQVSDIDKLV CRDKLSSTSQ LKSVGLNLEG NGVATDVPTA TKRWGFRAGV PPKVVNYEAG EWAENCYNLD IKKADGSEC LPEAPEGVRG FPRCRYVHKV SGTGPCPEGY AFHKEGAFFL YDRLASTIIY RSTTFSEGVV AFLILPETKK D FFQSPPLH EPANMTTDPS SYYHTVTLNY VADNFGTNMT NFLFQVDHLT YVQLEPRFTP QFLVQLNETI YTNGRRSNTT GT LIWKVNP TVDTGVGEWA FWENKKNFTK TLSSEELSVI FVPIDISEST EPGPLTNTTR GAANLLTGSR RTRR UniProtKB: Envelope glycoprotein |
-Macromolecule #2: Bundibugyo virus GP2
| Macromolecule | Name: Bundibugyo virus GP2 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Bundibugyo ebolavirus Bundibugyo ebolavirus |
| Molecular weight | Theoretical: 19.683902 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EITLRTQAKC NPNLHYWTTQ DEGAAIGLAW IPYFGPAAEG IYTEGIMHNQ NGLICGLRQL ANETTQALQL FLRATTELRT FSILNRKAI DFLLQRWGGT CHILGPDCCI EPHDWTKNIT DKIDQIIHDF IDKPLPDQTD VEVDDDDKAG WSHPQFEKGG G SGGGSGGG SWSHPQFEK UniProtKB: Envelope glycoprotein |
-Macromolecule #3: Antibody ADI-15878 Fab, light chain
| Macromolecule | Name: Antibody ADI-15878 Fab, light chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.333459 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DIVLTQSPST LSASVGDRVT ITCRASQSIS SWLAWYQQKP GEAPKLLISD ASSLESGVPS RFSGSGSGTE FTLTISSLQP DDFATYYCQ QYYSSPTFGG GTKVEIK |
-Macromolecule #4: Antibody ADI-15878 Fab, heavy chain
| Macromolecule | Name: Antibody ADI-15878 Fab, heavy chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.043574 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: VQLVQSGVTL VQPGGSLRVS CAASGFTFSS YAMSWVRQAP GKGLEWVSAI SGLGGSTYYA DSVKGRFTIS RDNSKNTLYL QMNSLRAED TAVYYCAKDH RVWAAGYHFD YWGQGALVTV S |
-Macromolecule #7: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 7 / Number of copies: 6 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Solutions were made fresh from 10X concentrate and filtered with a 0.2 micrometer filter into a sterile glass container. | |||||||||
| Grid | Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 5 sec. / Pretreatment - Atmosphere: OTHER / Details: unspecified | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: 3.5s blot on both sides of the grid using Whattman No. 1 filter paper. | |||||||||
| Details | Antibody ADI-15878 Fab was added in 10 molar excess to glycoprotein and allowed to incubate overnight at 4 degrees Celsius followed by size exclusion chromatography to purify the complex and rid of excess Fab. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Details | Preliminary grid screening was performed manually |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated magnification: 38167 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)