[English] 日本語
 Yorodumi
Yorodumi- EMDB-22820: BG505 SOSIP.664 in complex with the V3-targeting rhesus macaque a... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22820 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | BG505 SOSIP.664 in complex with the V3-targeting rhesus macaque antibody 1485 and human gp120-gp41 interface antibody 8ANC195 | |||||||||
 Map data Map data | unsharpened main map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HIV-1 / broadly neutralizing antibody / Env / envelope / BG505 / bNAb / VIRAL PROTEIN / VIRAL PROTEIN-Immune System complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell ...symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   Human immunodeficiency virus 1 Human immunodeficiency virus 1 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.55 Å | |||||||||
 Authors Authors | Barnes CO / Bjorkman PJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: A broadly neutralizing macaque monoclonal antibody against the HIV-1 V3-Glycan patch. Authors: Zijun Wang / Christopher O Barnes / Rajeev Gautam / Julio C Cetrulo Lorenzi / Christian T Mayer / Thiago Y Oliveira / Victor Ramos / Melissa Cipolla / Kristie M Gordon / Harry B Gristick / ...Authors: Zijun Wang / Christopher O Barnes / Rajeev Gautam / Julio C Cetrulo Lorenzi / Christian T Mayer / Thiago Y Oliveira / Victor Ramos / Melissa Cipolla / Kristie M Gordon / Harry B Gristick / Anthony P West / Yoshiaki Nishimura / Henna Raina / Michael S Seaman / Anna Gazumyan / Malcolm Martin / Pamela J Bjorkman / Michel C Nussenzweig / Amelia Escolano /  Abstract: A small fraction of HIV-1- infected humans develop broadly neutralizing antibodies (bNAbs) against HIV-1 that protect macaques from simian immunodeficiency HIV chimeric virus (SHIV). Similarly, a ...A small fraction of HIV-1- infected humans develop broadly neutralizing antibodies (bNAbs) against HIV-1 that protect macaques from simian immunodeficiency HIV chimeric virus (SHIV). Similarly, a small number of macaques infected with SHIVs develop broadly neutralizing serologic activity, but less is known about the nature of simian antibodies. Here, we report on a monoclonal antibody, Ab1485, isolated from a macaque infected with SHIVAD8 that developed broadly neutralizing serologic activity targeting the V3-glycan region of HIV-1 Env. Ab1485 neutralizes 38.1% of HIV-1 isolates in a 42-pseudovirus panel with a geometric mean IC50 of 0.055 µg/mLl and SHIVAD8 with an IC50 of 0.028 µg/mLl. Ab1485 binds the V3-glycan epitope in a glycan-dependent manner. A 3.5 Å cryo-electron microscopy structure of Ab1485 in complex with a native-like SOSIP Env trimer showed conserved contacts with the N332gp120 glycan and gp120 GDIR peptide motif, but in a distinct Env-binding orientation relative to human V3/N332gp120 glycan-targeting bNAbs. Intravenous infusion of Ab1485 protected macaques from a high dose challenge with SHIVAD8. We conclude that macaques can develop bNAbs against the V3-glycan patch that resemble human V3-glycan bNAbs. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22820.map.gz emd_22820.map.gz | 108.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22820-v30.xml emd-22820-v30.xml emd-22820.xml emd-22820.xml | 23.3 KB 23.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22820_fsc.xml emd_22820_fsc.xml | 11.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_22820.png emd_22820.png | 39.1 KB | ||
| Masks |  emd_22820_msk_1.map emd_22820_msk_1.map | 115.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-22820.cif.gz emd-22820.cif.gz | 7.8 KB | ||
| Others |  emd_22820_additional_1.map.gz emd_22820_additional_1.map.gz | 89.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22820 http://ftp.pdbj.org/pub/emdb/structures/EMD-22820 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22820 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22820 | HTTPS FTP |
-Related structure data
| Related structure data |  7kdeMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22820.map.gz / Format: CCP4 / Size: 115.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22820.map.gz / Format: CCP4 / Size: 115.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened main map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
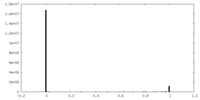
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.897 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_22820_msk_1.map emd_22820_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
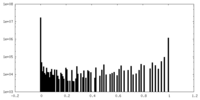
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Additional map
| File | emd_22820_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Additional map | ||||||||||||
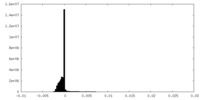
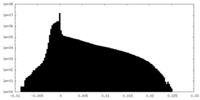
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of HIV-1 Env glycoprotein, BG505. SOSIP.664, with...
| Entire | Name: Ternary complex of HIV-1 Env glycoprotein, BG505. SOSIP.664, with broadly neutralizing antibodies 1485 and 8ANC195 |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of HIV-1 Env glycoprotein, BG505. SOSIP.664, with...
| Supramolecule | Name: Ternary complex of HIV-1 Env glycoprotein, BG505. SOSIP.664, with broadly neutralizing antibodies 1485 and 8ANC195 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 600 KDa |
-Macromolecule #1: HIV-1 Envelope Glycoprotein BG505 SOSIP.664 gp41
| Macromolecule | Name: HIV-1 Envelope Glycoprotein BG505 SOSIP.664 gp41 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.146482 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RNLLSGIVQQ QSNLLRAPEA QQHLLKLTVW GIKQLQARVL AVERYLRDQQ LLGIWGCSG KLICCTNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNEQDL LALD UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #2: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 53.864086 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: NLWVTVYYGV PVWKDAETTL FCASDAKAYE TEKHNVWATH ACVPTDPNPQ EIHLENVTEE FNMWKNNMVE QMHTDIISLW DQSLKPCVK LTPLCVTLQC TNVTNNITDD MRGELKNCSF NMTTELRDKK QKVYSLFYRL DVVQINENQG NRSNNSNKEY R LINCNTSA ...String: NLWVTVYYGV PVWKDAETTL FCASDAKAYE TEKHNVWATH ACVPTDPNPQ EIHLENVTEE FNMWKNNMVE QMHTDIISLW DQSLKPCVK LTPLCVTLQC TNVTNNITDD MRGELKNCSF NMTTELRDKK QKVYSLFYRL DVVQINENQG NRSNNSNKEY R LINCNTSA ITQACPKVSF EPIPIHYCAP AGFAILKCKD KKFNGTGPCP SVSTVQCTHG IKPVVSTQLL LNGSLAEEEV MI RSENITN NAKNILVQFN TPVQINCTRP NNNTRKSIRI GPGQAFYATG DIIGDIRQAH CNVSKATWNE TLGKVVKQLR KHF GNNTII RFANSSGGDL EVTTHSFNCG GEFFYCNTSG LFNSTWISNT SVQGSNSTGS NDSITLPCRI KQIINMWQRI GQAM YAPPI QGVIRCVSNI TGLILTRDGG STNSTTETFR PGGGDMRDNW RSELYKYKVV KIEPLGVAPT RCKRRVVGRR RRRR UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #3: Ab 1485 Heavy Chain
| Macromolecule | Name: Ab 1485 Heavy Chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.236078 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLVESGPG LVKASETLSL TCTVSGYNIR SNNWWSWVRQ PPGKGLEWIG GVYANSEITN YNSSLKSRVS ISQDAWRNKF SLKLKSVTD ADTAVYYCVR GPNHWEYFDS GNNEYFEFWG QGALVTVSSA STKGPSVFPL APSSKSTSGG TAALGCLVKD Y FPEPVTVS ...String: EVQLVESGPG LVKASETLSL TCTVSGYNIR SNNWWSWVRQ PPGKGLEWIG GVYANSEITN YNSSLKSRVS ISQDAWRNKF SLKLKSVTD ADTAVYYCVR GPNHWEYFDS GNNEYFEFWG QGALVTVSSA STKGPSVFPL APSSKSTSGG TAALGCLVKD Y FPEPVTVS WNSGALTSGV HTFPAVLQSS GLYSLSSVVT VPSSSLGTQT YICNVNHKPS NTKVDKRVEP KSCDK |
-Macromolecule #4: 8ANC195 Fab Heavy Chain
| Macromolecule | Name: 8ANC195 Fab Heavy Chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 26.153283 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QIHLVQSGTE VKKPGSSVTV SCKAYGVNTF GLYAVNWVRQ APGQSLEYIG QIWRWKSSAS HHFRGRVLIS AVDLTGSSPP ISSLEIKNL TSDDTAVYFC TTTSTYDRWS GLHHDGVMAF SSWGQGTLIS VSAASTKGPS VFPLAPSSKS TSGGTAALGC L VKDYFPEP ...String: QIHLVQSGTE VKKPGSSVTV SCKAYGVNTF GLYAVNWVRQ APGQSLEYIG QIWRWKSSAS HHFRGRVLIS AVDLTGSSPP ISSLEIKNL TSDDTAVYFC TTTSTYDRWS GLHHDGVMAF SSWGQGTLIS VSAASTKGPS VFPLAPSSKS TSGGTAALGC L VKDYFPEP VTVSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYICNVN HKPSNTKVDK RVEPKSCDKT HH HHHH |
-Macromolecule #5: 8ANC195 Fab Light Chain
| Macromolecule | Name: 8ANC195 Fab Light Chain / type: protein_or_peptide / ID: 5 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.460047 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQMTQSPST LSASTGDTVR ISCRASQSIT GNWVAWYQQR PGKAPRLLIY RGAALLGGVP SRFRGSAAGT DFTLTIGNLQ AEDFGTFYC QQYDTYPGTF GQGTKVEVKR TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD ...String: DIQMTQSPST LSASTGDTVR ISCRASQSIT GNWVAWYQQR PGKAPRLLIY RGAALLGGVP SRFRGSAAGT DFTLTIGNLQ AEDFGTFYC QQYDTYPGTF GQGTKVEVKR TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGEC |
-Macromolecule #6: Ab 1485 Light Chain
| Macromolecule | Name: Ab 1485 Light Chain / type: protein_or_peptide / ID: 6 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.167529 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSALTQPPSV SGAPGQRVTI SCTGTSSNIE NYHAFWYQQF SGMAPKLLIR DNDKRPSGVS DRFSGSKSGA SASLTITGLQ TDDEADYYC QSYDSGLRSY IFASGTRLTV LGQPKASPTV TLFPPSSEEL QANKATLVCL ISDFYPGAVT VAWKADSSPV K AGVETTTP ...String: QSALTQPPSV SGAPGQRVTI SCTGTSSNIE NYHAFWYQQF SGMAPKLLIR DNDKRPSGVS DRFSGSKSGA SASLTITGLQ TDDEADYYC QSYDSGLRSY IFASGTRLTV LGQPKASPTV TLFPPSSEEL QANKATLVCL ISDFYPGAVT VAWKADSSPV K AGVETTTP SKQSNNKYAA SSYLSLTPEQ WKSHRSYSCQ VTHEGSTVEK TVAPTECS |
-Macromolecule #17: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 17 / Number of copies: 34 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Details: 0.15 mA current | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV / Details: 3s blot, blot force 0. | |||||||||
| Details | monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 1 / Number real images: 1926 / Average exposure time: 3.6 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 45000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||
|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 118 / Target criteria: Correlation coefficient | ||||||
| Output model |  PDB-7kde: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)