[English] 日本語
 Yorodumi
Yorodumi- EMDB-8780: An envelope of a filamentous hyperthermophilic virus carries lipi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8780 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | An envelope of a filamentous hyperthermophilic virus carries lipids in a horseshoe conformation | ||||||||||||
 Map data Map data | envelope of a filamentous hyperthermophilic virus | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | AFV1 / hyperthermophile / filamentous virus / A-form DNA / VIRUS | ||||||||||||
| Function / homology | : / Filamentous archaeal viruses coat protein / helical viral capsid / DNA binding / Major capsid protein 1 / Major capsid protein 2 Function and homology information Function and homology information | ||||||||||||
| Biological species |  Acidianus filamentous virus 1 Acidianus filamentous virus 1 | ||||||||||||
| Method | helical reconstruction / cryo EM / Resolution: 4.5 Å | ||||||||||||
 Authors Authors | Kasson P / DiMaio F | ||||||||||||
| Funding support |  United States, United States,  France, 3 items France, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2017 Journal: Elife / Year: 2017Title: Model for a novel membrane envelope in a filamentous hyperthermophilic virus. Authors: Peter Kasson / Frank DiMaio / Xiong Yu / Soizick Lucas-Staat / Mart Krupovic / Stefan Schouten / David Prangishvili / Edward H Egelman /    Abstract: Biological membranes create compartments, and are usually formed by lipid bilayers. However, in hyperthermophilic archaea that live optimally at temperatures above 80°C the membranes are monolayers ...Biological membranes create compartments, and are usually formed by lipid bilayers. However, in hyperthermophilic archaea that live optimally at temperatures above 80°C the membranes are monolayers which resemble fused bilayers. Many double-stranded DNA viruses which parasitize such hosts, including the filamentous virus AFV1 of , are enveloped with a lipid-containing membrane. Using cryo-EM, we show that the membrane in AFV1 is a ~2 nm-thick monolayer, approximately half the expected membrane thickness, formed by host membrane-derived lipids which adopt a U-shaped 'horseshoe' conformation. We hypothesize that this unusual viral envelope structure results from the extreme curvature of the viral capsid, as 'horseshoe' lipid conformations favor such curvature and host membrane lipids that permit horseshoe conformations are selectively recruited into the viral envelope. The unusual envelope found in AFV1 also has many implications for biotechnology, since this membrane can survive the most aggressive conditions involving extremes of temperature and pH. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8780.map.gz emd_8780.map.gz | 10.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8780-v30.xml emd-8780-v30.xml emd-8780.xml emd-8780.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8780.png emd_8780.png | 63.4 KB | ||
| Filedesc metadata |  emd-8780.cif.gz emd-8780.cif.gz | 5.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8780 http://ftp.pdbj.org/pub/emdb/structures/EMD-8780 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8780 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8780 | HTTPS FTP |
-Related structure data
| Related structure data |  5w7gMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_8780.map.gz / Format: CCP4 / Size: 28.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8780.map.gz / Format: CCP4 / Size: 28.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | envelope of a filamentous hyperthermophilic virus | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
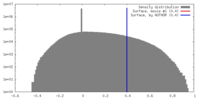
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Acidianus filamentous virus 1
| Entire | Name:  Acidianus filamentous virus 1 Acidianus filamentous virus 1 |
|---|---|
| Components |
|
-Supramolecule #1: Acidianus filamentous virus 1
| Supramolecule | Name: Acidianus filamentous virus 1 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 235266 / Sci species name: Acidianus filamentous virus 1 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|
-Macromolecule #1: ORF140
| Macromolecule | Name: ORF140 / type: protein_or_peptide / ID: 1 / Number of copies: 21 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Acidianus filamentous virus 1 Acidianus filamentous virus 1 |
| Molecular weight | Theoretical: 15.832856 KDa |
| Sequence | String: MAKLNRKLRQ DSTDRYKTKL YLWRNLGGLI PEDMAISVTE SITADWKQYN DMMSKVRNET LDILKTNKVA TEDYIGYIAF AEELAHQVW KNKNSSPDPN TANEASKTDL ESKYSDVYGL DVTVLDAIYN AVIPIIMGGG S UniProtKB: Major capsid protein 1 |
-Macromolecule #2: ORF132
| Macromolecule | Name: ORF132 / type: protein_or_peptide / ID: 2 / Number of copies: 21 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Acidianus filamentous virus 1 Acidianus filamentous virus 1 |
| Molecular weight | Theoretical: 15.017159 KDa |
| Sequence | String: MVNKKYRQDS KDRYQYKQYI YRSIGGIVPP EMAETVTANQ TAQWEAGFTP YHKLRLAIKE ICKTDGIPNI KWGMYIAFGE KLLKSYLKM KAGSASSDMI AEYINNAISA FSSRTGISQE TAQKIADFIT SNY UniProtKB: Major capsid protein 2 |
-Macromolecule #3: DNA (253-MER)
| Macromolecule | Name: DNA (253-MER) / type: dna / ID: 3 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Acidianus filamentous virus 1 Acidianus filamentous virus 1 |
| Molecular weight | Theoretical: 77.747367 KDa |
| Sequence | String: (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT) (DA)(DT)(DA)(DT)(DA)(DT) ...String: (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DA) (DT) (DA)(DT)(DA)(DT)(DA)(DT)(DA)(DT) (DA)(DT)(DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 6 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 20.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 4.6 Å Applied symmetry - Helical parameters - Δ&Phi: 38.7 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: BACK PROJECTION / Resolution.type: BY AUTHOR / Resolution: 4.5 Å / Resolution method: OTHER / Software - Name: SPIDER / Number images used: 119495 |
|---|---|
| Startup model | Type of model: OTHER |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)