[English] 日本語
 Yorodumi
Yorodumi- EMDB-5423: Filaments from Ignicoccus hospitalis Show Diversity of Packing in... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5423 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Filaments from Ignicoccus hospitalis Show Diversity of Packing in Proteins Containing N-terminal Type IV Pilin Helices | |||||||||
 Map data Map data | Reconstruction of adhesion filament | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | helical polymers / Type IV pili | |||||||||
| Function / homology | Flagellin/pilin, N-terminal / Archaeal Type IV pilin N-terminal domain-containing protein Function and homology information Function and homology information | |||||||||
| Biological species |   Ignicoccus hospitalis (archaea) Ignicoccus hospitalis (archaea) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 7.5 Å | |||||||||
 Authors Authors | Yu S / Goforth C / Meyer C / Rachel R / Wirth R / Schroeder G / Egelman EH | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2012 Journal: J Mol Biol / Year: 2012Title: Filaments from Ignicoccus hospitalis show diversity of packing in proteins containing N-terminal type IV pilin helices. Authors: Xiong Yu / Charles Goforth / Carolin Meyer / Reinhard Rachel / Reinhard Wirth / Gunnar F Schröder / Edward H Egelman /  Abstract: Bacterial motility is driven by the rotation of flagellar filaments that supercoil. The supercoiling involves the switching of coiled-coil protofilaments between two different states. In archaea, the ...Bacterial motility is driven by the rotation of flagellar filaments that supercoil. The supercoiling involves the switching of coiled-coil protofilaments between two different states. In archaea, the flagellar filaments responsible for motility are formed by proteins with distinct homology in their N-terminal portion to bacterial Type IV pilins. The bacterial pilins have a single N-terminal hydrophobic α-helix, not the coiled coil found in flagellin. We have used electron cryo-microscopy to study the adhesion filaments from the archaeon Ignicoccus hospitalis. While I. hospitalis is non-motile, these filaments make transitions between rigid stretches and curved regions and appear morphologically similar to true archaeal flagellar filaments. A resolution of ~7.5Å allows us to unambiguously build a model for the packing of these N-terminal α-helices, and this packing is different from several bacterial Type IV pili whose structure has been analyzed by electron microscopy and modeling. Our results show that the mechanism responsible for the supercoiling of bacterial flagellar filaments cannot apply to archaeal filaments. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5423.map.gz emd_5423.map.gz | 10.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5423-v30.xml emd-5423-v30.xml emd-5423.xml emd-5423.xml | 8.4 KB 8.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5423_1.jpg emd_5423_1.jpg | 78.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5423 http://ftp.pdbj.org/pub/emdb/structures/EMD-5423 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5423 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5423 | HTTPS FTP |
-Related structure data
| Related structure data |  3j1rMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5423.map.gz / Format: CCP4 / Size: 10.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5423.map.gz / Format: CCP4 / Size: 10.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of adhesion filament | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.25 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
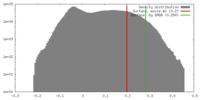
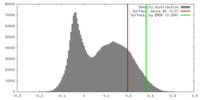
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Adhesion filament
| Entire | Name: Adhesion filament |
|---|---|
| Components |
|
-Supramolecule #1000: Adhesion filament
| Supramolecule | Name: Adhesion filament / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Adhesion filament
| Supramolecule | Name: Adhesion filament / type: organelle_or_cellular_component / ID: 1 / Oligomeric state: helical filament / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:   Ignicoccus hospitalis (archaea) Ignicoccus hospitalis (archaea) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Jan 1, 2011 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON COOLSCAN / Number real images: 17 / Bits/pixel: 14 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.4 µm / Nominal defocus min: 1.7 µm / Nominal magnification: 55000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | The filaments were reconstructed using IHRSR. |
|---|---|
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 5.3 Å Applied symmetry - Helical parameters - Δ&Phi: 106.65 ° Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.5 Å / Resolution method: OTHER / Software - Name: Spider, IHRSR Details: Each image was multiplied by the CTF. The final volume was amplitude-corrected in Fourier space by dividing by the sum of the squared CTFs. |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)