+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5185 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM Helical Reconstruction of TMV | |||||||||
 Map data Map data | Helically symmetrized volume of Tobacco Mosaic Virus. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TMV / helical / RNA | |||||||||
| Function / homology | Tobacco mosaic virus-like, coat protein / Tobacco mosaic virus-like, coat protein / Tobacco mosaic virus-like, coat protein superfamily / Virus coat protein (TMV like) / helical viral capsid / structural molecule activity / identical protein binding / Capsid protein Function and homology information Function and homology information | |||||||||
| Biological species | unidentified (others) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Ge P / Zhou ZH | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2011 Journal: Proc Natl Acad Sci U S A / Year: 2011Title: Hydrogen-bonding networks and RNA bases revealed by cryo electron microscopy suggest a triggering mechanism for calcium switches. Authors: Peng Ge / Z Hong Zhou /  Abstract: Helical assemblies such as filamentous viruses, flagella, and F-actin represent an important category of structures in biology. As the first discovered virus, tobacco mosaic virus (TMV) was at the ...Helical assemblies such as filamentous viruses, flagella, and F-actin represent an important category of structures in biology. As the first discovered virus, tobacco mosaic virus (TMV) was at the center of virus research. Previously, the structure of TMV was solved at atomic detail by X-ray fiber diffraction but only for its dormant or high-calcium-concentration state, not its low-calcium-concentration state, which is relevant to viral assembly and disassembly inside host cells. Here we report a helical reconstruction of TMV in its calcium-free, metastable assembling state at 3.3 Å resolution by cryo electron microscopy, revealing both protein side chains and RNA bases. An atomic model was built de novo showing marked differences from the high-calcium, dormant-state structure. Although it could be argued that there might be inaccuracies in the latter structure derived from X-ray fiber diffraction, these differences can be interpreted as conformational changes effected by calcium-driven switches, a common regulatory mechanism in plant viruses. Our comparisons of the structures of the low- and high-calcium states indicate that hydrogen bonds formed by Asp116 and Arg92 in the place of the calcium ion of the dormant (high-calcium) state might trigger allosteric changes in the RNA base-binding pockets of the coat protein. In turn, the coat protein-RNA interactions in our structure favor an adenine-X-guanine (A*G) motif over the G*A motif of the dormant state, thus offering an explanation underlying viral assembly initiation by an AAG motif. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5185.map.gz emd_5185.map.gz | 483.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5185-v30.xml emd-5185-v30.xml emd-5185.xml emd-5185.xml | 9.8 KB 9.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5185_1.png emd_5185_1.png | 224.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5185 http://ftp.pdbj.org/pub/emdb/structures/EMD-5185 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5185 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5185 | HTTPS FTP |
-Validation report
| Summary document |  emd_5185_validation.pdf.gz emd_5185_validation.pdf.gz | 378.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5185_full_validation.pdf.gz emd_5185_full_validation.pdf.gz | 378.5 KB | Display | |
| Data in XML |  emd_5185_validation.xml.gz emd_5185_validation.xml.gz | 8.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5185 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5185 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5185 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5185 | HTTPS FTP |
-Related structure data
| Related structure data |  3j06MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5185.map.gz / Format: CCP4 / Size: 500 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5185.map.gz / Format: CCP4 / Size: 500 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Helically symmetrized volume of Tobacco Mosaic Virus. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.869 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
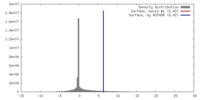
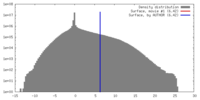
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Tobacco Mosaic Virus (Vulgare)
| Entire | Name:  Tobacco Mosaic Virus (Vulgare) Tobacco Mosaic Virus (Vulgare) |
|---|---|
| Components |
|
-Supramolecule #1000: Tobacco Mosaic Virus (Vulgare)
| Supramolecule | Name: Tobacco Mosaic Virus (Vulgare) / type: sample / ID: 1000 / Oligomeric state: helix / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 37.8 MDa |
-Macromolecule #1: Coat Protein
| Macromolecule | Name: Coat Protein / type: protein_or_peptide / ID: 1 / Name.synonym: Coat Protein / Number of copies: 2160 / Oligomeric state: helix / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism: unidentified (others) / Strain: vulgare |
| Molecular weight | Theoretical: 17.3 MDa |
| Sequence | GO: helical viral capsid / InterPro: Tobacco mosaic virus-like, coat protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: 10mM Tris, 130mM NaCl, pH 7.4 |
|---|---|
| Grid | Details: Quantifoil 1.3/1.2 400 mesh |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 93 K / Instrument: FEI VITROBOT MARK IV Details: Vitrification instrument: FEI Vitrobot Mark IV. 2 uL sample Method: 10 second wait time, 1-2 second blot time, 1-2 second drain time, blot force 1 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Average: 90 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 250,000 times magnification Legacy - Electron beam tilt params: 0 |
| Date | Jun 15, 2009 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 0.869 µm / Number real images: 386 / Average electron dose: 25 e/Å2 / Bits/pixel: 16 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 73000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.86 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 3.3 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: IHRSR, EMAN |
|---|---|
| CTF correction | Details: EMAN |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)