+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | INF2 at the Barbed End of F-Actin with Incoming Profilin-Actin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Actin / Filament / Elongation / Ends / CYTOSOLIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsynapse maturation / adenyl-nucleotide exchange factor activity / negative regulation of actin filament bundle assembly / modification of postsynaptic actin cytoskeleton / positive regulation of actin filament bundle assembly / regulation of actin filament polymerization / Signaling by ROBO receptors / negative regulation of stress fiber assembly / proline-rich region binding / PCP/CE pathway ...synapse maturation / adenyl-nucleotide exchange factor activity / negative regulation of actin filament bundle assembly / modification of postsynaptic actin cytoskeleton / positive regulation of actin filament bundle assembly / regulation of actin filament polymerization / Signaling by ROBO receptors / negative regulation of stress fiber assembly / proline-rich region binding / PCP/CE pathway / cytoskeletal motor activator activity / positive regulation of ruffle assembly / myosin heavy chain binding / regulation of mitochondrial fission / tropomyosin binding / actin filament bundle / troponin I binding / filamentous actin / mesenchyme migration / positive regulation of actin filament polymerization / skeletal muscle myofibril / positive regulation of epithelial cell migration / actin filament bundle assembly / striated muscle thin filament / skeletal muscle thin filament assembly / actin monomer binding / skeletal muscle fiber development / stress fiber / phosphotyrosine residue binding / titin binding / actin filament polymerization / phosphatidylinositol-4,5-bisphosphate binding / filopodium / actin filament / neural tube closure / RHO GTPases Activate Formins / modulation of chemical synaptic transmission / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / small GTPase binding / calcium-dependent protein binding / Platelet degranulation / lamellipodium / actin binding / actin cytoskeleton organization / cell body / blood microparticle / cell cortex / cytoskeleton / protein stabilization / cadherin binding / protein domain specific binding / focal adhesion / hydrolase activity / calcium ion binding / positive regulation of gene expression / regulation of transcription by RNA polymerase II / perinuclear region of cytoplasm / glutamatergic synapse / magnesium ion binding / RNA binding / extracellular exosome / ATP binding / membrane / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
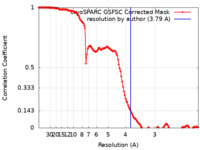
| Method | single particle reconstruction / cryo EM / Resolution: 3.79 Å | |||||||||
 Authors Authors | Palmer NJ / Barrie KR / Dominguez R | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2024 Journal: Nature / Year: 2024Title: Mechanisms of actin filament severing and elongation by formins. Authors: Nicholas J Palmer / Kyle R Barrie / Roberto Dominguez /  Abstract: Humans express 15 formins that play crucial roles in actin-based processes, including cytokinesis, cell motility and mechanotransduction. However, the lack of structures bound to the actin filament ...Humans express 15 formins that play crucial roles in actin-based processes, including cytokinesis, cell motility and mechanotransduction. However, the lack of structures bound to the actin filament (F-actin) has been a major impediment to understanding formin function. Whereas formins are known for their ability to nucleate and elongate F-actin, some formins can additionally depolymerize, sever or bundle F-actin. Two mammalian formins, inverted formin 2 (INF2) and diaphanous 1 (DIA1, encoded by DIAPH1), exemplify this diversity. INF2 shows potent severing activity but elongates weakly whereas DIA1 has potent elongation activity but does not sever. Using cryo-electron microscopy (cryo-EM) we show five structural states of INF2 and two of DIA1 bound to the middle and barbed end of F-actin. INF2 and DIA1 bind differently to these sites, consistent with their distinct activities. The formin-homology 2 and Wiskott-Aldrich syndrome protein-homology 2 (FH2 and WH2, respectively) domains of INF2 are positioned to sever F-actin, whereas DIA1 appears unsuited for severing. These structures also show how profilin-actin is delivered to the fast-growing barbed end, and how this is followed by a transition of the incoming monomer into the F-actin conformation and the release of profilin. Combined, the seven structures presented here provide step-by-step visualization of the mechanisms of F-actin severing and elongation by formins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_44018.map.gz emd_44018.map.gz | 77.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-44018-v30.xml emd-44018-v30.xml emd-44018.xml emd-44018.xml | 26.2 KB 26.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_44018_fsc.xml emd_44018_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_44018.png emd_44018.png | 39.7 KB | ||
| Masks |  emd_44018_msk_1.map emd_44018_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-44018.cif.gz emd-44018.cif.gz | 8.2 KB | ||
| Others |  emd_44018_half_map_1.map.gz emd_44018_half_map_1.map.gz emd_44018_half_map_2.map.gz emd_44018_half_map_2.map.gz | 200.7 MB 200.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-44018 http://ftp.pdbj.org/pub/emdb/structures/EMD-44018 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44018 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-44018 | HTTPS FTP |
-Related structure data
| Related structure data |  9azpMC  9az4C  9azqC  9b03C  9b0kC  9b27C  9b3dC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_44018.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_44018.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_44018_msk_1.map emd_44018_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_44018_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_44018_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Actin
| Entire | Name: Actin |
|---|---|
| Components |
|
-Supramolecule #1: Actin
| Supramolecule | Name: Actin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 264 KDa |
-Supramolecule #2: INF2 Dimer
| Supramolecule | Name: INF2 Dimer / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Actin Filament
| Supramolecule | Name: Actin Filament / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Actin, alpha skeletal muscle
| Macromolecule | Name: Actin, alpha skeletal muscle / type: protein_or_peptide / ID: 1 / Number of copies: 7 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 41.387227 KDa |
| Sequence | String: TTALVCDNGS GLVKAGFAGD DAPRAVFPSI VGRPRHQGVM VGMGQKDSYV GDEAQSKRGI LTLKYPIE(HIC)G IITNWD DME KIWHHTFYNE LRVAPEEHPT LLTEAPLNPK ANREKMTQIM FETFNVPAMY VAIQAVLSLY ASGRTTGIVL DSGDGVT HN VPIYEGYALP ...String: TTALVCDNGS GLVKAGFAGD DAPRAVFPSI VGRPRHQGVM VGMGQKDSYV GDEAQSKRGI LTLKYPIE(HIC)G IITNWD DME KIWHHTFYNE LRVAPEEHPT LLTEAPLNPK ANREKMTQIM FETFNVPAMY VAIQAVLSLY ASGRTTGIVL DSGDGVT HN VPIYEGYALP HAIMRLDLAG RDLTDYLMKI LTERGYSFVT TAEREIVRDI KEKLCYVALD FENEMATAAS SSSLEKSY E LPDGQVITIG NERFRCPETL FQPSFIGMES AGIHETTYNS IMKCDIDIRK DLYANNVMSG GTTMYPGIAD RMQKEITAL APSTMKIKII APPERKYSVW IGGSILASLS TFQQMWITKQ EYDEAGPSIV HRKCF UniProtKB: Actin, alpha skeletal muscle |
-Macromolecule #2: Inverted formin-2
| Macromolecule | Name: Inverted formin-2 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 135.778828 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSVKEGAQRK WAALKEKLGP QDSDPTEANL ESADPELCIR LLQMPSVVNY SGLRKRLEGS DGGWMVQFLE QSGLDLLLEA LARLSGRGV ARISDALLQL TCVSCVRAVM NSRQGIEYIL SNQGYVRQLS QALDTSNVMV KKQVFELLAA LCIYSPEGHV L TLDALDHY ...String: MSVKEGAQRK WAALKEKLGP QDSDPTEANL ESADPELCIR LLQMPSVVNY SGLRKRLEGS DGGWMVQFLE QSGLDLLLEA LARLSGRGV ARISDALLQL TCVSCVRAVM NSRQGIEYIL SNQGYVRQLS QALDTSNVMV KKQVFELLAA LCIYSPEGHV L TLDALDHY KTVCSQQYRF SIVMNELSGS DNVPYVVTLL SVINAVILGP EDLRARTQLR NEFIGLQLLD VLARLRDLED AD LLIQLEA FEEAKAEDEE ELLRVSGGVD MSSHQEVFAS LFHKVSCSPV SAQLLSVLQG LLHLEPTLRS SQLLWEALES LVN RAVLLA SDAQECTLEE VVERLLSVKG RPRPSPLVKA HKSVQANLDQ SQRGSSPQNT TTPKPSVEGQ QPAAAAACEP VDHA QSESI LKVSQPRALE QQASTPPPPP PPPLLPGSSA EPPPPPPPPP LPSVGAKALP TAPPPPPLPG LGAMAPPAPP LPPPL PGSC EFLPPPPPPL PGLGCPPPPP PLLPGMGWGP PPPPPPLLPC TCSPPVAGGM EEVIVAQVDH GLGSAWVPSH RRVNPP TLR MKKLNWQKLP SNVAREHNSM WASLSSPDAE AVEPDFSSIE RLFSFPAAKP KEPTMVAPRA RKEPKEITFL DAKKSLN LN IFLKQFKCSN EEVAAMIRAG DTTKFDVEVL KQLLKLLPEK HEIENLRAFT EERAKLASAD HFYLLLLAIP CYQLRIEC M LLCEGAAAVL DMVRPKAQLV LAACESLLTS RQLPIFCQLI LRIGNFLNYG SHTGDADGFK ISTLLKLTET KSQQNRVTL LHHVLEEAEK SHPDLLQLPR DLEQPSQAAG INLEIIRSEA SSNLKKLLET ERKVSASVAE VQEQYTERLQ ASISAFRALD ELFEAIEQK QRELADYLCE DAQQLSLEDT FSTMKAFRDL FLRALKENKD RKEQAAKAER RKQQLAEEEA RRPRGEDGKP V RKGPGKQE EVCVIDALLA DIRKGFQLRK TARGRGDTDG GSKAASMDPP RATEPVATSN PAGDPVGSTR CPASEPGLDA TT ASESRGW DLVDAVTPGP QPTLEQLEEG GPRPLERRSS WYVDASDVLT TEDPQCPQPL EGAWPVTLGD AQALKPLKFS SNQ PPAAGS SRQDAKDPTS LLGVLQAEAD STSEGLEDAV HSRGARPPAA GPGGDEDEDE EDTAPESALD TSLDKSFSED AVTD SSGSG TLPRARGRAS KGTGKRRKKR PSRSQEEVPP DSDDNKTKKL CVIQ UniProtKB: Inverted formin-2 |
-Macromolecule #3: Profilin-1
| Macromolecule | Name: Profilin-1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.940021 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AGWNAYIDNL MADGTCQDAA IVGYKDSPSV WAAVPGKTFV NITPAEVGVL VGKDRSSFYV NGLTLGGQKC SVIRDSLLQD GEFSMDLRT KSTGGAPTFN VTVTKTDKTL VLLMGKEGVH GGLINKKCYE MASHLRRSQY UniProtKB: Profilin-1 |
-Macromolecule #4: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 6 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 7 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 1 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.45 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 100 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 120 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 41926 / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller






































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)