+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4189 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of deformed wing virus carrying the GFP gene | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Deformed wing virus / Picornavirales / Iflaviridae / Iflavirus / viral protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationhost cell membrane / viral capsid / host cell cytoplasm / RNA helicase activity / viral RNA genome replication / cysteine-type endopeptidase activity / RNA-dependent RNA polymerase activity / DNA-templated transcription / structural molecule activity / proteolysis ...host cell membrane / viral capsid / host cell cytoplasm / RNA helicase activity / viral RNA genome replication / cysteine-type endopeptidase activity / RNA-dependent RNA polymerase activity / DNA-templated transcription / structural molecule activity / proteolysis / RNA binding / ATP binding / membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Deformed wing virus Deformed wing virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Skubnik K / Plevka P | |||||||||
| Funding support | 2 items
| |||||||||
 Citation Citation |  Journal: Curr Opin Virol / Year: 2020 Journal: Curr Opin Virol / Year: 2020Title: Virion structures and genome delivery of honeybee viruses. Authors: Michaela Procházková / Karel Škubník / Tibor Füzik / Liya Mukhamedova / Antonín Přidal / Pavel Plevka /  Abstract: The western honeybee is the primary pollinator of numerous food crops. Furthermore, honeybees are essential for ecosystem stability by sustaining the diversity and abundance of wild flowering plants. ...The western honeybee is the primary pollinator of numerous food crops. Furthermore, honeybees are essential for ecosystem stability by sustaining the diversity and abundance of wild flowering plants. However, the worldwide population of honeybees is under pressure from environmental stress and pathogens. Viruses from the families Iflaviridae and Dicistroviridae, together with their vector, the parasitic mite Varroa destructor, are the major threat to the world's honeybees. Dicistroviruses and iflaviruses have capsids with icosahedral symmetries. Acidic pH triggers the genome release of both dicistroviruses and iflaviruses. The capsids of iflaviruses expand, whereas those of dicistroviruses remain compact until the genome release. Furthermore, dicistroviruses use inner capsid proteins, whereas iflaviruses employ protruding domains or minor capsid proteins from the virion surface to penetrate membranes and deliver their genomes into the cell cytoplasm. The structural characterization of the infection process opens up possibilities for the development of antiviral compounds. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4189.map.gz emd_4189.map.gz | 62.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4189-v30.xml emd-4189-v30.xml emd-4189.xml emd-4189.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4189_fsc.xml emd_4189_fsc.xml | 18 KB | Display |  FSC data file FSC data file |
| Images |  emd_4189.png emd_4189.png | 253.7 KB | ||
| Filedesc metadata |  emd-4189.cif.gz emd-4189.cif.gz | 6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4189 http://ftp.pdbj.org/pub/emdb/structures/EMD-4189 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4189 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4189 | HTTPS FTP |
-Validation report
| Summary document |  emd_4189_validation.pdf.gz emd_4189_validation.pdf.gz | 305.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_4189_full_validation.pdf.gz emd_4189_full_validation.pdf.gz | 305.1 KB | Display | |
| Data in XML |  emd_4189_validation.xml.gz emd_4189_validation.xml.gz | 16.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4189 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4189 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4189 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-4189 | HTTPS FTP |
-Related structure data
| Related structure data |  6f5jMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4189.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4189.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.063 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
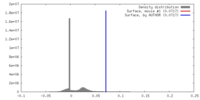
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Deformed wing virus
| Entire | Name:   Deformed wing virus Deformed wing virus |
|---|---|
| Components |
|
-Supramolecule #1: Deformed wing virus
| Supramolecule | Name: Deformed wing virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / Details: Virus was purified from honeybee pupae. / NCBI-ID: 198112 / Sci species name: Deformed wing virus / Virus type: VIRION / Virus isolate: OTHER / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / Diameter: 390.0 Å / T number (triangulation number): 3 |
-Macromolecule #1: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Deformed wing virus Deformed wing virus |
| Molecular weight | Theoretical: 28.679273 KDa |
| Sequence | String: GEESRNTTVL DTTTTLQSSG FGRAFFGEAF NDLKTLMRRY QLYGQLLLSV TTDKDIDHCM FTFPCLPQGL ALDIGSAGSP HEIFNRCRD GIIPLIASGY RFYRGDLRYK IVFPSNVNSN IWVQHRPDRR LEGWSAAKIV NCDAVSTGQG VYNHGYASHI Q ITRVNNVI ...String: GEESRNTTVL DTTTTLQSSG FGRAFFGEAF NDLKTLMRRY QLYGQLLLSV TTDKDIDHCM FTFPCLPQGL ALDIGSAGSP HEIFNRCRD GIIPLIASGY RFYRGDLRYK IVFPSNVNSN IWVQHRPDRR LEGWSAAKIV NCDAVSTGQG VYNHGYASHI Q ITRVNNVI ELEVPFYNAT CYNYLQAFNA SSAASSYAVS LGEISVGFQA TSDDIASIVN KPVTIYYSIG DGMQFSQWVG YQ PMMILDQ LPAPVVRAVP E UniProtKB: Genome polyprotein |
-Macromolecule #2: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Deformed wing virus Deformed wing virus |
| Molecular weight | Theoretical: 28.3609 KDa |
| Sequence | String: MDNPNPGPDG EGEVELEKDS NVVLTTQRDP STSIPAPVSV KWSRWTSNDV VDDYATITSR WYQIAEFVWS KDDPFDKELA RLILPRALL SSIEANSDAI CDVPNTIPFK VHAYWRGDME VRVQINSNKF QVGQLQATWY YSDHENLNIS SKRSVYGFSQ M DHALISAS ...String: MDNPNPGPDG EGEVELEKDS NVVLTTQRDP STSIPAPVSV KWSRWTSNDV VDDYATITSR WYQIAEFVWS KDDPFDKELA RLILPRALL SSIEANSDAI CDVPNTIPFK VHAYWRGDME VRVQINSNKF QVGQLQATWY YSDHENLNIS SKRSVYGFSQ M DHALISAS ASNEAKLVIP FKHVYPFLPT RIVPDWTTGI LDMGALNIRV IAPLRMSATG PTTCNVVVFI KLNNSEFTGT SS GKFYASQ IRAKPE UniProtKB: Genome polyprotein |
-Macromolecule #3: Genome polyprotein
| Macromolecule | Name: Genome polyprotein / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Deformed wing virus Deformed wing virus |
| Molecular weight | Theoretical: 46.697582 KDa |
| Sequence | String: DNPSYQQSPR HFVPTGMHSL ALGTNLVEPL HALRLDAAGT TQHPVGCAPD EDMTVSSIAS RYGLIRRVQW KKDHAKGSLL LQLDADPFV EQRIEGTNPI SLYWFAPVGV VSSMFMQWRG SLEYRFDIIA SQFHTGRLIV GYVPGLTASL QLQMDYMKLK S SSYVVFDL ...String: DNPSYQQSPR HFVPTGMHSL ALGTNLVEPL HALRLDAAGT TQHPVGCAPD EDMTVSSIAS RYGLIRRVQW KKDHAKGSLL LQLDADPFV EQRIEGTNPI SLYWFAPVGV VSSMFMQWRG SLEYRFDIIA SQFHTGRLIV GYVPGLTASL QLQMDYMKLK S SSYVVFDL QESNSFTFEV PYVSYRPWWV RKYGGNYLPS STDAPSTLFM YVQVPLIPME AVSDTIDINV YVRGGSSFEV CV PVQPSLG LNWNTDFILR NDEEYRAKTG YAPYYAGVWH SFNNSNSLVF RWGSASDQIA QWPTISVPRG ELAFLRIKDG KQA AVGTQP WRTMVVWPSG HGYNIGIPTY NAERARQLAQ HLYGGGSLTD EKAKQLFVPA NQQGPGKVSN GNPVWEVMRA PLAT QRAHI QDFEFIEAIP E UniProtKB: Genome polyprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL |
|---|---|
| Buffer | pH: 7.4 Details: Dulbeccos Phosphate Buffered Saline D8537 sigma aldrich |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: NITROGEN / Pretreatment - Pressure: 0.007 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 298 K / Instrument: FEI VITROBOT MARK IV |
| Details | Virus was dissolved in PBS buffer. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Frames/image: 2-16 / Number grids imaged: 1 / Average exposure time: 1.0 sec. / Average electron dose: 21.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated magnification: 74235 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-6f5j: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)