+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human ULK1C:PI3KC3-C1 supercomplex | |||||||||
 Map data Map data | z-flipped from the original map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Autophagy / Protein kinase / Lipid kinase / Supercomplex / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationextrinsic component of omegasome membrane / phosphatidylinositol 3-kinase inhibitor activity / regulation of triglyceride metabolic process / extrinsic component of phagophore assembly site membrane / nucleus-vacuole junction / cellular response to aluminum ion / Toll Like Receptor 9 (TLR9) Cascade / protein lipidation / omegasome membrane / neuron projection regeneration ...extrinsic component of omegasome membrane / phosphatidylinositol 3-kinase inhibitor activity / regulation of triglyceride metabolic process / extrinsic component of phagophore assembly site membrane / nucleus-vacuole junction / cellular response to aluminum ion / Toll Like Receptor 9 (TLR9) Cascade / protein lipidation / omegasome membrane / neuron projection regeneration / Synthesis of PIPs at the late endosome membrane / regulation of protein lipidation / phosphatidylinositol 3-kinase complex, class III / Synthesis of PIPs at the early endosome membrane / ribophagy / cellular response to oxygen-glucose deprivation / phosphatidylinositol 3-kinase complex, class III, type II / phosphatidylinositol 3-kinase complex, class III, type I / negative regulation of collateral sprouting / positive regulation of stress granule assembly / Atg1/ULK1 kinase complex / mitochondria-associated endoplasmic reticulum membrane contact site / autophagy of peroxisome / glycophagy / response to mitochondrial depolarisation / positive regulation of attachment of mitotic spindle microtubules to kinetochore / cytoplasmic side of mitochondrial outer membrane / negative regulation of lysosome organization / positive regulation by host of viral genome replication / Synthesis of PIPs at the Golgi membrane / receptor catabolic process / positive regulation of autophagosome assembly / engulfment of apoptotic cell / phosphatidylinositol kinase activity / negative regulation of autophagosome assembly / regulation of protein complex stability / phosphatidylinositol 3-kinase regulator activity / protein targeting to vacuole / suppression by virus of host autophagy / protein localization to phagophore assembly site / protein targeting to lysosome / phagophore assembly site membrane / late endosome to vacuole transport / early endosome to late endosome transport / SMAD protein signal transduction / piecemeal microautophagy of the nucleus / RAB GEFs exchange GTP for GDP on RABs / negative regulation of programmed cell death / regulation of tumor necrosis factor-mediated signaling pathway / protein kinase regulator activity / axon extension / TBC/RABGAPs / Translation of Replicase and Assembly of the Replication Transcription Complex / phagophore assembly site / cytoplasmic pattern recognition receptor signaling pathway / autophagy of mitochondrion / response to iron(II) ion / cellular response to nitrogen starvation / reticulophagy / positive regulation of protein targeting to mitochondrion / phosphatidylinositol 3-kinase / phosphatidylinositol-3-phosphate biosynthetic process / autophagosome membrane docking / mitotic metaphase chromosome alignment / post-transcriptional regulation of gene expression / 1-phosphatidylinositol-3-kinase activity / endosome to lysosome transport / Receptor Mediated Mitophagy / lysosome organization / Macroautophagy / RSV-host interactions / response to starvation / positive regulation of cardiac muscle hypertrophy / autolysosome / p38MAPK cascade / phosphatidylinositol-mediated signaling / axoneme / phosphatidylinositol phosphate biosynthetic process / autophagosome membrane / PI3K Cascade / positive regulation of cell size / autophagosome maturation / neuron development / autophagosome assembly / mitophagy / RHO GTPases Activate NADPH Oxidases / response to vitamin E / negative regulation of reactive oxygen species metabolic process / amyloid-beta metabolic process / regulation of macroautophagy / negative regulation of protein-containing complex assembly / cellular defense response / cellular response to nutrient levels / phosphatidylinositol 3-kinase binding / cellular response to glucose starvation / phagocytic vesicle / positive regulation of intrinsic apoptotic signaling pathway / positive regulation of autophagy / protein-membrane adaptor activity / extrinsic apoptotic signaling pathway Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.2 Å | |||||||||
 Authors Authors | Chen M / Hurley JH | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: bioRxiv / Year: 2023 Journal: bioRxiv / Year: 2023Title: Structure and activation of the human autophagy-initiating ULK1C:PI3KC3-C1 supercomplex Authors: Chen M / Ren X / Cook A / Hurley JH | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40738.map.gz emd_40738.map.gz | 777.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40738-v30.xml emd-40738-v30.xml emd-40738.xml emd-40738.xml | 29 KB 29 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40738.png emd_40738.png | 63.8 KB | ||
| Filedesc metadata |  emd-40738.cif.gz emd-40738.cif.gz | 8.6 KB | ||
| Others |  emd_40738_half_map_1.map.gz emd_40738_half_map_1.map.gz emd_40738_half_map_2.map.gz emd_40738_half_map_2.map.gz | 762.1 MB 762.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40738 http://ftp.pdbj.org/pub/emdb/structures/EMD-40738 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40738 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40738 | HTTPS FTP |
-Validation report
| Summary document |  emd_40738_validation.pdf.gz emd_40738_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40738_full_validation.pdf.gz emd_40738_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_40738_validation.xml.gz emd_40738_validation.xml.gz | 20.4 KB | Display | |
| Data in CIF |  emd_40738_validation.cif.gz emd_40738_validation.cif.gz | 24.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40738 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40738 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40738 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40738 | HTTPS FTP |
-Related structure data
| Related structure data |  9c82MC  8soiC  8sorC  8sqzC  8srmC  8srq M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40738.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40738.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | z-flipped from the original map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.115 Å | ||||||||||||||||||||||||||||||||||||
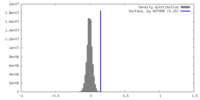
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map A
| File | emd_40738_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
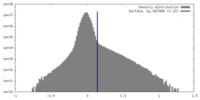
| Density Histograms |
-Half map: half map B
| File | emd_40738_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
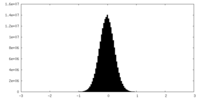
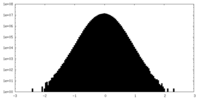
| Density Histograms |
- Sample components
Sample components
+Entire : Supercomplex composed of ULK1 complex and PI3KC3-C1 complex
+Supramolecule #1: Supercomplex composed of ULK1 complex and PI3KC3-C1 complex
+Supramolecule #2: Human autophagy initiation ULK1 complex core
+Supramolecule #3: Human autophagy initiation PI3KC3-C1 complex
+Macromolecule #1: RB1-inducible coiled-coil protein 1
+Macromolecule #2: Serine/threonine-protein kinase ULK1
+Macromolecule #3: Autophagy-related protein 13
+Macromolecule #4: Phosphoinositide 3-kinase regulatory subunit 4
+Macromolecule #5: Phosphatidylinositol 3-kinase catalytic subunit type 3
+Macromolecule #6: Beclin 1-associated autophagy-related key regulator
+Macromolecule #7: Beclin-1-C 35 kDa
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.35 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Details: 25 mA | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 3421 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 36000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)