+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human PI3KC3-C1 complex | |||||||||
 Map data Map data | Composite map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Autophagy / Lipid kinase / Complex / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationextrinsic component of omegasome membrane / phosphatidylinositol 3-kinase inhibitor activity / regulation of triglyceride metabolic process / extrinsic component of phagophore assembly site membrane / nucleus-vacuole junction / cellular response to aluminum ion / positive regulation of protein lipidation / postsynaptic endosome / Toll Like Receptor 9 (TLR9) Cascade / Synthesis of PIPs at the late endosome membrane ...extrinsic component of omegasome membrane / phosphatidylinositol 3-kinase inhibitor activity / regulation of triglyceride metabolic process / extrinsic component of phagophore assembly site membrane / nucleus-vacuole junction / cellular response to aluminum ion / positive regulation of protein lipidation / postsynaptic endosome / Toll Like Receptor 9 (TLR9) Cascade / Synthesis of PIPs at the late endosome membrane / positive regulation of stress granule assembly / phosphatidylinositol 3-kinase complex, class III / cellular response to oxygen-glucose deprivation / Synthesis of PIPs at the early endosome membrane / phosphatidylinositol 3-kinase complex, class III, type II / phosphatidylinositol 3-kinase complex, class III, type I / host-mediated activation of viral genome replication / response to mitochondrial depolarisation / positive regulation of attachment of mitotic spindle microtubules to kinetochore / presynaptic endosome / mitochondria-associated endoplasmic reticulum membrane contact site / negative regulation of lysosome organization / engulfment of apoptotic cell / Synthesis of PIPs at the Golgi membrane / phosphatidylinositol kinase activity / regulation of protein complex stability / positive regulation of autophagosome assembly / response to L-leucine / early endosome to late endosome transport / negative regulation of autophagosome assembly / cytoplasmic side of mitochondrial outer membrane / protein localization to phagophore assembly site / phosphatidylinositol 3-kinase regulator activity / receptor catabolic process / phagophore assembly site membrane / protein targeting to vacuole / protein targeting to lysosome / endosome organization / SMAD protein signal transduction / late endosome to vacuole transport / pexophagy / phagophore assembly site / Translation of Replicase and Assembly of the Replication Transcription Complex / phosphatidylinositol-3-phosphate biosynthetic process / cellular response to nitrogen starvation / negative regulation of programmed cell death / phosphatidylinositol 3-kinase / lysosome organization / 1-phosphatidylinositol-3-kinase activity / response to iron(II) ion / post-transcriptional regulation of gene expression / mitotic metaphase chromosome alignment / response to vitamin E / cytoplasmic pattern recognition receptor signaling pathway / autophagosome membrane docking / endosome to lysosome transport / Macroautophagy / positive regulation of cardiac muscle hypertrophy / phosphatidylinositol-mediated signaling / RSV-host interactions / p38MAPK cascade / phosphatidylinositol phosphate biosynthetic process / negative regulation of protein phosphorylation / autolysosome / autophagosome membrane / synaptic vesicle endocytosis / PI3K Cascade / axoneme / RHO GTPases Activate NADPH Oxidases / autophagosome assembly / autophagosome maturation / amyloid-beta metabolic process / regulation of macroautophagy / neuron development / cellular defense response / intercellular bridge / cellular response to glucose starvation / phosphatidylinositol 3-kinase binding / mitophagy / positive regulation of intrinsic apoptotic signaling pathway / cellular response to copper ion / phagocytic vesicle / protein-membrane adaptor activity / JNK cascade / positive regulation of autophagy / cellular response to amino acid starvation / autophagosome / cellular response to epidermal growth factor stimulus / regulation of cytokinesis / cellular response to starvation / Antigen Presentation: Folding, assembly and peptide loading of class I MHC / macroautophagy / phosphatidylinositol 3-kinase/protein kinase B signal transduction / trans-Golgi network / response to lead ion / GABA-ergic synapse / protein processing / ISG15 antiviral mechanism / autophagy / circadian rhythm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.96 Å | |||||||||
 Authors Authors | Chen M / Hurley JH | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: bioRxiv / Year: 2023 Journal: bioRxiv / Year: 2023Title: Structure and activation of the human autophagy-initiating ULK1C:PI3KC3-C1 supercomplex Authors: Chen M / Ren X / Cook A / Hurley JH | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40669.map.gz emd_40669.map.gz | 167.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40669-v30.xml emd-40669-v30.xml emd-40669.xml emd-40669.xml | 27 KB 27 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40669.png emd_40669.png | 86.7 KB | ||
| Filedesc metadata |  emd-40669.cif.gz emd-40669.cif.gz | 8.5 KB | ||
| Others |  emd_40669_additional_1.map.gz emd_40669_additional_1.map.gz emd_40669_additional_2.map.gz emd_40669_additional_2.map.gz emd_40669_additional_3.map.gz emd_40669_additional_3.map.gz | 168 MB 164.7 MB 164.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40669 http://ftp.pdbj.org/pub/emdb/structures/EMD-40669 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40669 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40669 | HTTPS FTP |
-Validation report
| Summary document |  emd_40669_validation.pdf.gz emd_40669_validation.pdf.gz | 502.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40669_full_validation.pdf.gz emd_40669_full_validation.pdf.gz | 502.5 KB | Display | |
| Data in XML |  emd_40669_validation.xml.gz emd_40669_validation.xml.gz | 6.9 KB | Display | |
| Data in CIF |  emd_40669_validation.cif.gz emd_40669_validation.cif.gz | 8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40669 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40669 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40669 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40669 | HTTPS FTP |
-Related structure data
| Related structure data |  8sorMC  8soiC  8sqzC  8srmC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40669.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40669.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Composite map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.115 Å | ||||||||||||||||||||||||||||||||||||
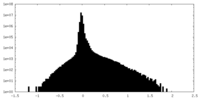
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Consensus map
| File | emd_40669_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Consensus map | ||||||||||||
| Projections & Slices |
| ||||||||||||
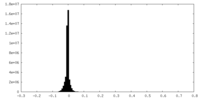
| Density Histograms |
-Additional map: Local refinement map 1/2
| File | emd_40669_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local refinement map 1/2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
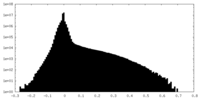
| Density Histograms |
-Additional map: Local refinement map 2/2
| File | emd_40669_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local refinement map 2/2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
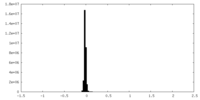
| Density Histograms |
- Sample components
Sample components
-Entire : Human autophagy initiation PI3KC3-C1 complex
| Entire | Name: Human autophagy initiation PI3KC3-C1 complex |
|---|---|
| Components |
|
-Supramolecule #1: Human autophagy initiation PI3KC3-C1 complex
| Supramolecule | Name: Human autophagy initiation PI3KC3-C1 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 362 KDa |
-Macromolecule #1: Phosphatidylinositol 3-kinase catalytic subunit type 3
| Macromolecule | Name: Phosphatidylinositol 3-kinase catalytic subunit type 3 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: phosphatidylinositol 3-kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 101.680328 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGEAEKFHYI YSCDLDINVQ LKIGSLEGKR EQKSYKAVLE DPMLKFSGLY QETCSDLYVT CQVFAEGKPL ALPVRTSYKA FSTRWNWNE WLKLPVKYPD LPRNAQVALT IWDVYGPGKA VPVGGTTVSL FGKYGMFRQG MHDLKVWPNV EADGSEPTKT P GRTSSTLS ...String: MGEAEKFHYI YSCDLDINVQ LKIGSLEGKR EQKSYKAVLE DPMLKFSGLY QETCSDLYVT CQVFAEGKPL ALPVRTSYKA FSTRWNWNE WLKLPVKYPD LPRNAQVALT IWDVYGPGKA VPVGGTTVSL FGKYGMFRQG MHDLKVWPNV EADGSEPTKT P GRTSSTLS EDQMSRLAKL TKAHRQGHMV KVDWLDRLTF REIEMINESE KRSSNFMYLM VEFRCVKCDD KEYGIVYYEK DG DESSPIL TSFELVKVPD PQMSMENLVE SKHHKLARSL RSGPSDHDLK PNAATRDQLN IIVSYPPTKQ LTYEEQDLVW KFR YYLTNQ EKALTKFLKC VNWDLPQEAK QALELLGKWK PMDVEDSLEL LSSHYTNPTV RRYAVARLRQ ADDEDLLMYL LQLV QALKY ENFDDIKNGL EPTKKDSQSS VSENVSNSGI NSAEIDSSQI ITSPLPSVSS PPPASKTKEV PDGENLEQDL CTFLI SRAC KNSTLANYLY WYVIVECEDQ DTQQRDPKTH EMYLNVMRRF SQALLKGDKS VRVMRSLLAA QQTFVDRLVH LMKAVQ RES GNRKKKNERL QALLGDNEKM NLSDVELIPL PLEPQVKIRG IIPETATLFK SALMPAQLFF KTEDGGKYPV IFKHGDD LR QDQLILQIIS LMDKLLRKEN LDLKLTPYKV LATSTKHGFM QFIQSVPVAE VLDTEGSIQN FFRKYAPSEN GPNGISAE V MDTYVKSCAG YCVITYILGV GDRHLDNLLL TKTGKLFHID FGYILGRDPK PLPPPMKLNK EMVEGMGGTQ SEQYQEFRK QCYTAFLHLR RYSNLILNLF SLMVDANIPD IALEPDKTVK KVQDKFRLDL SDEEAVHYMQ SLIDESVHAL FAAVVEQIHK FAQYWRK UniProtKB: Phosphatidylinositol 3-kinase catalytic subunit type 3 |
-Macromolecule #2: Beclin 1-associated autophagy-related key regulator
| Macromolecule | Name: Beclin 1-associated autophagy-related key regulator / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.387266 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MASPSGKGAR ALEAPGCGPR PLARDLVDSV DDAEGLYVAV ERCPLCNTTR RRLTCAKCVQ SGDFVYFDGR DRERFIDKKE RLSRLKSKQ EEFQKEVLKA MEGKWITDQL RWKIMSCKMR IEQLKQTICK GNEEMEKNSE GLLKTKEKNQ KLYSRAQRHQ E KKEKIQRH ...String: MASPSGKGAR ALEAPGCGPR PLARDLVDSV DDAEGLYVAV ERCPLCNTTR RRLTCAKCVQ SGDFVYFDGR DRERFIDKKE RLSRLKSKQ EEFQKEVLKA MEGKWITDQL RWKIMSCKMR IEQLKQTICK GNEEMEKNSE GLLKTKEKNQ KLYSRAQRHQ E KKEKIQRH NRKLGDLVEK KTIDLRSHYE RLANLRRSHI LELTSVIFPI EEVKTGVRDP ADVSSESDSA MTSSTVSKLA EA RRTTYLS GRWVCDDHNG DTSISITGPW ISLPNNGDYS AYYSWVEEKK TTQGPDMEQS NPAYTISAAL CYATQLVNIL SHI LDVNLP KKLCNSEFCG ENLSKQKFTR AVKKLNANIL YLCFSQHVNL DQLQPLHTLR NLMYLVSPSS EHLGRSGPFE VRAD LEESM EFVDPGVAGE SDESGDERVS DEETDLGTDW ENLPSPRFCD IPSQSVEVSQ SQSTQASPPI ASSSAGGMIS SAAAS VTSW FKAYTGHR UniProtKB: Beclin 1-associated autophagy-related key regulator |
-Macromolecule #3: Beclin-1
| Macromolecule | Name: Beclin-1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 51.953102 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEGSKTSNNS TMQVSFVCQR CSQPLKLDTS FKILDRVTIQ ELTAPLLTTA QAKPGETQEE ETNSGEEPFI ETPRQDGVSR RFIPPARMM STESANSFTL IGEASDGGTM ENLSRRLKVT GDLFDIMSGQ TDVDHPLCEE CTDTLLDQLD TQLNVTENEC Q NYKRCLEI ...String: MEGSKTSNNS TMQVSFVCQR CSQPLKLDTS FKILDRVTIQ ELTAPLLTTA QAKPGETQEE ETNSGEEPFI ETPRQDGVSR RFIPPARMM STESANSFTL IGEASDGGTM ENLSRRLKVT GDLFDIMSGQ TDVDHPLCEE CTDTLLDQLD TQLNVTENEC Q NYKRCLEI LEQMNEDDSE QLQMELKELA LEEERLIQEL EDVEKNRKIV AENLEKVQAE AERLDQEEAQ YQREYSEFKR QQ LELDDEL KSVENQMRYA QTQLDKLKKT NVFNATFHIW HSGQFGTINN FRLGRLPSVP VEWNEINAAW GQTVLLLHAL ANK MGLKFQ RYRLVPYGNH SYLESLTDKS KELPLYCSGG LRFFWDNKFD HAMVAFLDCV QQFKEEVEKG ETRFCLPYRM DVEK GKIED TGGSGGSYSI KTQFNSEEQW TKALKFMLTN LKWGLAWVSS QFYNK UniProtKB: Beclin-1 |
-Macromolecule #4: Phosphoinositide 3-kinase regulatory subunit 4
| Macromolecule | Name: Phosphoinositide 3-kinase regulatory subunit 4 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO / EC number: non-specific serine/threonine protein kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 153.293797 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGNQLAGIAP SQILSVESYF SDIHDFEYDK SLGSTRFFKV ARAKHREGLV VVKVFAIQDP TLPLTSYKQE LEELKIRLNS AQNCLPFQK ASEKASEKAA MLFRQYVRDN LYDRISTRPF LNNIEKRWIA FQILTAVDQA HKSGVRHGDI KTENVMVTSW N WVLLTDFA ...String: MGNQLAGIAP SQILSVESYF SDIHDFEYDK SLGSTRFFKV ARAKHREGLV VVKVFAIQDP TLPLTSYKQE LEELKIRLNS AQNCLPFQK ASEKASEKAA MLFRQYVRDN LYDRISTRPF LNNIEKRWIA FQILTAVDQA HKSGVRHGDI KTENVMVTSW N WVLLTDFA SFKPTYLPED NPADFNYFFD TSRRRTCYIA PERFVDGGMF ATELEYMRDP STPLVDLNSN QRTRGELKRA MD IFSAGCV IAELFTEGVP LFDLSQLLAY RNGHFFPEQV LNKIEDHSIR ELVTQMIHRE PDKRLEAEDY LKQQRGNAFP EIF YTFLQP YMAQFAKETF LSADERILVI RKDLGNIIHN LCGHDLPEKA EGEPKENGLV ILVSVITSCL QTLKYCDSKL AALE LILHL APRLSVEILL DRITPYLLHF SNDSVPRVRA EALRTLTKVL ALVKEVPRND INIYPEYILP GIAHLAQDDA TIVRL AYAE NIALLAETAL RFLELVQLKN LNMENDPNNE EIDEVTHPNG NYDTELQALH EMVQQKVVTL LSDPENIVKQ TLMENG ITR LCVFFGRQKA NDVLLSHMIT FLNDKNDWHL RGAFFDSIVG VAAYVGWQSS SILKPLLQQG LSDAEEFVIV KALYALT CM CQLGLLQKPH VYEFASDIAP FLCHPNLWIR YGAVGFITVV ARQISTADVY CKLMPYLDPY ITQPIIQIER KLVLLSVL K EPVSRSIFDY ALRSKDITSL FRHLHMRQKK RNGSLPDCPP PEDPAIAQLL KKLLSQGMTE EEEDKLLALK DFMMKSNKA KANIVDQSHL HDSSQKGVID LAALGITGRQ VDLVKTKQEP DDKRARKHVK QDSNVNEEWK SMFGSLDPPN MPQALPKGSD QEVIQTGKP PRSESSAGIC VPLSTSSQVP EVTTVQNKKP VIPVLSSTIL PSTYQIRITT CKTELQQLIQ QKREQCNAER I AKQMMENA EWESKPPPPG WRPKGLLVAH LHEHKSAVNR IRVSDEHSLF ATCSNDGTVK IWNSQKMEGK TTTTRSILTY SR IGGRVKT LTFCQGSHYL AIASDNGAVQ LLGIEASKLP KSPKIHPLQS RILDQKEDGC VVDMHHFNSG AQSVLAYATV NGS LVGWDL RSSSNAWTLK HDLKSGLITS FAVDIHQCWL CIGTSSGTMA CWDMRFQLPI SSHCHPSRAR IRRLSMHPLY QSWV IAAVQ GNNEVSMWDM ETGDRRFTLW ASSAPPLSEL QPSPHSVHGI YCSPADGNPI LLTAGSDMKI RFWDLAYPER SYVVA GSTS SPSVSYYRKI IEGTEVVQEI QNKQKVGPSD DTPRRGPESL PVGHHDIITD VATFQTTQGF IVTASRDGIV KVWK UniProtKB: Phosphoinositide 3-kinase regulatory subunit 4 |
-Macromolecule #5: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 1 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.25 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR / Details: 25 mA | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 2243 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 36000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
| Output model |  PDB-8sor: |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)