[English] 日本語
 Yorodumi
Yorodumi- EMDB-4039: Maltose binding protein genetically fused to dodecameric glutamin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4039 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Maltose binding protein genetically fused to dodecameric glutamine synthetase | |||||||||
 Map data Map data | Maltose-binding protein fused to dodecameric glutamine synthetase | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Fusion protein / chimera / dodecamer / symmetrized construct / ligase | |||||||||
| Function / homology |  Function and homology information Function and homology informationnitrogen utilization / glutamine synthetase / : / glutamine synthetase activity / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing / outer membrane-bounded periplasmic space ...nitrogen utilization / glutamine synthetase / : / glutamine synthetase activity / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing / outer membrane-bounded periplasmic space / ATP binding / membrane / metal ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Salmonella typhi (bacteria) / Salmonella typhi (bacteria) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.2 Å | |||||||||
 Authors Authors | Coscia F / Petosa C | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2016 Journal: Sci Rep / Year: 2016Title: Fusion to a homo-oligomeric scaffold allows cryo-EM analysis of a small protein. Authors: Francesca Coscia / Leandro F Estrozi / Fabienne Hans / Hélène Malet / Marjolaine Noirclerc-Savoye / Guy Schoehn / Carlo Petosa /  Abstract: Recent technical advances have revolutionized the field of cryo-electron microscopy (cryo-EM). However, most monomeric proteins remain too small (<100 kDa) for cryo-EM analysis. To overcome this limitation, we explored a strategy whereby a monomeric target protein is genetically fused to a homo-oligomeric scaffold protein and the junction optimized to allow the target to adopt the scaffold symmetry, thereby generating a chimeric particle suitable for cryo-EM. To demonstrate the concept, we fused maltose-binding protein (MBP), a 40 kDa monomer, to glutamine synthetase, a dodecamer formed by two hexameric rings. Chimeric constructs with different junction lengths were screened by biophysical analysis and negative-stain EM. The optimal construct yielded a cryo-EM reconstruction that revealed the MBP structure at sub-nanometre resolution. These findings illustrate the feasibility of using homo-oligomeric scaffolds to enable cryo-EM analysis of monomeric proteins, paving the way for applying this strategy to challenging structures resistant to crystallographic and NMR analysis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4039.map.gz emd_4039.map.gz | 191.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4039-v30.xml emd-4039-v30.xml emd-4039.xml emd-4039.xml | 14.8 KB 14.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4039.png emd_4039.png | 216 KB | ||
| Filedesc metadata |  emd-4039.cif.gz emd-4039.cif.gz | 6.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4039 http://ftp.pdbj.org/pub/emdb/structures/EMD-4039 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4039 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4039 | HTTPS FTP |
-Related structure data
| Related structure data |  5ldfMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4039.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4039.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Maltose-binding protein fused to dodecameric glutamine synthetase | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8155 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
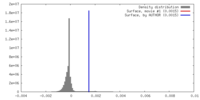
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Maltose-binding protein genetically fused to glutamine synthetase
| Entire | Name: Maltose-binding protein genetically fused to glutamine synthetase |
|---|---|
| Components |
|
-Supramolecule #1: Maltose-binding protein genetically fused to glutamine synthetase
| Supramolecule | Name: Maltose-binding protein genetically fused to glutamine synthetase type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.11 MDa |
-Macromolecule #1: Glutamine synthetase
| Macromolecule | Name: Glutamine synthetase / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO / EC number: glutamine synthetase |
|---|---|
| Source (natural) | Organism:  Salmonella typhi (bacteria) Salmonella typhi (bacteria) |
| Molecular weight | Theoretical: 51.586266 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EHVLTMLNEH EVKFVDLRFT DTKGKEQHVT IPAHQVNAEF FEEGKMFDGS SIGGWKGINE SDMVLMPDAS TAVIDPFFAD STLIIRCDI LEPGTLQGYD RDPRSIAKRA EDYLRATGIA DTVLFGPEPE FFLFDDIRFG ASISGSHVAI DDIEGAWNSS T KYEGGNKG ...String: EHVLTMLNEH EVKFVDLRFT DTKGKEQHVT IPAHQVNAEF FEEGKMFDGS SIGGWKGINE SDMVLMPDAS TAVIDPFFAD STLIIRCDI LEPGTLQGYD RDPRSIAKRA EDYLRATGIA DTVLFGPEPE FFLFDDIRFG ASISGSHVAI DDIEGAWNSS T KYEGGNKG HRPGVKGGYF PVPPVDSAQD IRSEMCLVME QMGLVVEAHH HEVATAGQNE VATRFNTMTK KADEIQIYKY VV HNVAHRF GKTATFMPKP MFGDNGSGMH CHMSLAKNGT NLFSGDKYAG LSEQALYYIG GVIKHAKAIN ALANPTTNSY KRL VPGYEA PVMLAYSARN RSASIRIPVV ASPKARRIEV RFPDPAANPY LCFAALLMAG LDGIKNKIHP GEPMDKNLYD LPPE EAKEI PQVAGSLEEA LNALDLDREF LKAGGVFTDE AIDAYIALRR EEDDRVRMTP HPVEFELYYS V UniProtKB: Glutamine synthetase |
-Macromolecule #2: Maltose-binding periplasmic protein
| Macromolecule | Name: Maltose-binding periplasmic protein / type: protein_or_peptide / ID: 2 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.753152 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KIEEGKLVIW INGDKGYNGL AEVGKKFEKD TGIKVTVEHP DKLEEKFPQV AATGDGPDII FWAHDRFGGY AQSGLLAEIT PDKAFQDKL YPFTWDAVRY NGKLIAYPIA VEALSLIYNK DLLPNPPKTW EEIPALDKEL KAKGKSALMF NLQEPYFTWP L IAADGGYA ...String: KIEEGKLVIW INGDKGYNGL AEVGKKFEKD TGIKVTVEHP DKLEEKFPQV AATGDGPDII FWAHDRFGGY AQSGLLAEIT PDKAFQDKL YPFTWDAVRY NGKLIAYPIA VEALSLIYNK DLLPNPPKTW EEIPALDKEL KAKGKSALMF NLQEPYFTWP L IAADGGYA FKYENGKYDI KDVGVDNAGA KAGLTFLVDL IKNKHMNADT DYSIAEAAFN KGETAMTING PWAWSNIDTS KV NYGVTVL PTFKGQPSKP FVGVLSAGIN AASPNKELAK EFLENYLLTD EGLEAVNKDK PLGAVALKSY EEELAKDPRI AAT MENAQK GEIMPNIPQM SAFWYAVRTA VINAASGRQT VDEALKDAQT RITK UniProtKB: Maltose/maltodextrin-binding periplasmic protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||||||||
| Grid | Model: Quantifoil / Material: COPPER/RHODIUM / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 290 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Details | Polara Top entry |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number real images: 165 / Average exposure time: 6.0 sec. / Average electron dose: 25.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-5ldf: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)