[English] 日本語
 Yorodumi
Yorodumi- EMDB-3761: Cryo-EM structure of the TOM core complex from Neurospora crassa -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3761 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the TOM core complex from Neurospora crassa | |||||||||
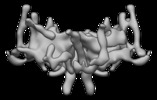
 Map data Map data | TOM core complex consisting of Tom40, Tom22, Tom5, Tom6 and Tom7 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TOM-Complex / Protein Import / Mitochondria / Cryo-EM / PROTEIN TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationmitochondrial outer membrane translocase complex / protein insertion into mitochondrial outer membrane / porin activity / pore complex / protein import into mitochondrial matrix / protein transmembrane transporter activity / monoatomic ion transport Similarity search - Function | |||||||||
| Biological species |  Neurospora crassa (fungus) / Neurospora crassa (fungus) /  Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) | |||||||||
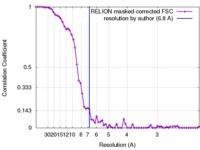
| Method | single particle reconstruction / cryo EM / Resolution: 6.8 Å | |||||||||
 Authors Authors | Bausewein T / Mills DJ | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2017 Journal: Cell / Year: 2017Title: Cryo-EM Structure of the TOM Core Complex from Neurospora crassa. Authors: Thomas Bausewein / Deryck J Mills / Julian D Langer / Beate Nitschke / Stephan Nussberger / Werner Kühlbrandt /  Abstract: The TOM complex is the main entry gate for protein precursors from the cytosol into mitochondria. We have determined the structure of the TOM core complex by cryoelectron microscopy (cryo-EM). The ...The TOM complex is the main entry gate for protein precursors from the cytosol into mitochondria. We have determined the structure of the TOM core complex by cryoelectron microscopy (cryo-EM). The complex is a 148 kDa symmetrical dimer of ten membrane protein subunits that create a shallow funnel on the cytoplasmic membrane surface. In the core of the dimer, the β-barrels of the Tom40 pore form two identical preprotein conduits. Each Tom40 pore is surrounded by the transmembrane segments of the α-helical subunits Tom5, Tom6, and Tom7. Tom22, the central preprotein receptor, connects the two Tom40 pores at the dimer interface. Our structure offers detailed insights into the molecular architecture of the mitochondrial preprotein import machinery. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3761.map.gz emd_3761.map.gz | 21.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3761-v30.xml emd-3761-v30.xml emd-3761.xml emd-3761.xml | 9.5 KB 9.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3761_fsc.xml emd_3761_fsc.xml | 6.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_3761.png emd_3761.png | 73.3 KB | ||
| Filedesc metadata |  emd-3761.cif.gz emd-3761.cif.gz | 4.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3761 http://ftp.pdbj.org/pub/emdb/structures/EMD-3761 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3761 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3761 | HTTPS FTP |
-Related structure data
| Related structure data |  5o8oMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_3761.map.gz / Format: CCP4 / Size: 23 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3761.map.gz / Format: CCP4 / Size: 23 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TOM core complex consisting of Tom40, Tom22, Tom5, Tom6 and Tom7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : TOM core complex consisting of Tom40, Tom22, Tom5, Tom6 and Tom7
| Entire | Name: TOM core complex consisting of Tom40, Tom22, Tom5, Tom6 and Tom7 |
|---|---|
| Components |
|
-Supramolecule #1: TOM core complex consisting of Tom40, Tom22, Tom5, Tom6 and Tom7
| Supramolecule | Name: TOM core complex consisting of Tom40, Tom22, Tom5, Tom6 and Tom7 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Neurospora crassa (fungus) / Strain: GR-107 Neurospora crassa (fungus) / Strain: GR-107 |
-Macromolecule #1: Mitochondrial import receptor subunit tom40
| Macromolecule | Name: Mitochondrial import receptor subunit tom40 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus)Strain: ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987 |
| Molecular weight | Theoretical: 38.184797 KDa |
| Sequence | String: MASFSTESPL AMLRDNAIYS SLSDAFNAFQ ERRKQFGLSN PGTIETIARE VQRDTLLTNY MFSGLRADVT KAFSLAPLFQ VSHQFAMGE RLNPYAFAAL YGTNQIFAQG NLDNEGALST RFNYRWGDRT ITKTQFSIGG GQDMAQFEHE HLGDDFSASL K AINPSFLD ...String: MASFSTESPL AMLRDNAIYS SLSDAFNAFQ ERRKQFGLSN PGTIETIARE VQRDTLLTNY MFSGLRADVT KAFSLAPLFQ VSHQFAMGE RLNPYAFAAL YGTNQIFAQG NLDNEGALST RFNYRWGDRT ITKTQFSIGG GQDMAQFEHE HLGDDFSASL K AINPSFLD GGLTGIFVGD YLQAVTPRLG LGLQAVWQRQ GLTQGPDTAI SYFARYKAGD WVASAQLQAQ GALNTSFWKK LT DRVQAGV DMTLSVAPSQ SMMGGLTKEG ITTFGAKYDF RMSTFRAQID SKGKLSCLLE KRLGAAPVTL TFAADVDHVT QQA KLGMSV SIEASDVDLQ EQQEGAQSLN IPF UniProtKB: Mitochondrial import receptor subunit tom40 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.2 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-5o8o: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)