+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of arginine oxidase from Pseudomonas sp. TRU 7192 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | OXIDOREDUCTASE | |||||||||
| Biological species |  Pseudomonas sp. (bacteria) Pseudomonas sp. (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.34 Å | |||||||||
 Authors Authors | Yamaguchi H / Numoto N / Suzuki H / Nishikawa K / Kamegawa A / Takahashi K / Sugiki M / Fujiyoshi Y | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: J Biochem / Year: 2025 Journal: J Biochem / Year: 2025Title: Open and closed structures of L-arginine oxidase by cryo-electron microscopy and X-ray crystallography. Authors: Hiroki Yamaguchi / Kazutoshi Takahashi / Nobutaka Numoto / Hiroshi Suzuki / Moemi Tatsumi / Akiko Kamegawa / Kouki Nishikawa / Yasuhisa Asano / Toshimi Mizukoshi / Hiroshi Miyano / Yoshinori ...Authors: Hiroki Yamaguchi / Kazutoshi Takahashi / Nobutaka Numoto / Hiroshi Suzuki / Moemi Tatsumi / Akiko Kamegawa / Kouki Nishikawa / Yasuhisa Asano / Toshimi Mizukoshi / Hiroshi Miyano / Yoshinori Fujiyoshi / Masayuki Sugiki /  Abstract: L-arginine oxidase (AROD, EC 1.4.3.25) is an oxidoreductase that catalyses the deamination of L-arginine, with flavin adenine dinucleotide (FAD) as a cofactor. Recently identified AROD from ...L-arginine oxidase (AROD, EC 1.4.3.25) is an oxidoreductase that catalyses the deamination of L-arginine, with flavin adenine dinucleotide (FAD) as a cofactor. Recently identified AROD from Pseudomonas sp. TPU 7192 (PT-AROD) demonstrates high selectivity for L-arginine. This enzyme is useful for accurate assays of L-arginine in biological samples. The structural characteristics of the FAD-dependent AROD, however, remain unknown. Here, we report the structure of PT-AROD at a resolution of 2.3 Å by cryo-electron microscopy. PT-AROD adopts an octameric structure with D4 symmetry, which is consistent with its molecular weight in solution, estimated by mass photometry. Comparative analysis of this structure with that determined using X-ray crystallography reveals open and closed forms of the lid-like loop at the entrance to the substrate pocket. Furthermore, mutation of Glu493, located at the substrate binding site, diminishes substrate selectivity, suggesting that this residue contributes significantly to the high selectivity of PT-AROD. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_36635.map.gz emd_36635.map.gz | 33.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-36635-v30.xml emd-36635-v30.xml emd-36635.xml emd-36635.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_36635_fsc.xml emd_36635_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_36635.png emd_36635.png | 117.7 KB | ||
| Masks |  emd_36635_msk_1.map emd_36635_msk_1.map | 36.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-36635.cif.gz emd-36635.cif.gz | 6.6 KB | ||
| Others |  emd_36635_half_map_1.map.gz emd_36635_half_map_1.map.gz emd_36635_half_map_2.map.gz emd_36635_half_map_2.map.gz | 28.9 MB 28.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-36635 http://ftp.pdbj.org/pub/emdb/structures/EMD-36635 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36635 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-36635 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_36635.map.gz / Format: CCP4 / Size: 36.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_36635.map.gz / Format: CCP4 / Size: 36.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.96 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_36635_msk_1.map emd_36635_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
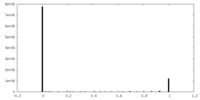
| Density Histograms |
-Half map: #2
| File | emd_36635_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_36635_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : homooctamer of arginine oxidase
| Entire | Name: homooctamer of arginine oxidase |
|---|---|
| Components |
|
-Supramolecule #1: homooctamer of arginine oxidase
| Supramolecule | Name: homooctamer of arginine oxidase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Pseudomonas sp. (bacteria) / Strain: TPU 7192 Pseudomonas sp. (bacteria) / Strain: TPU 7192 |
| Molecular weight | Theoretical: 540 KDa |
-Macromolecule #1: Amine oxidoreductase
| Macromolecule | Name: Amine oxidoreductase / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas sp. (bacteria) / Strain: TPU 7192 Pseudomonas sp. (bacteria) / Strain: TPU 7192 |
| Molecular weight | Theoretical: 67.393164 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHMSQ TQPLDVAIIG GGVSGTYSAW RLQEAQGDHQ RIQLFEYSDR IGGRLFSINL PGLPNVVAEV GGMRWMPATK DNTGGHVMV DKLVGELKLE SKNFPMGSNL PDKDPVGAKD NLFYLRGERF RLRDFTEAPD KIPYKLAWSE RGYGPEDLQV K VMHNIYPG ...String: MHHHHHHMSQ TQPLDVAIIG GGVSGTYSAW RLQEAQGDHQ RIQLFEYSDR IGGRLFSINL PGLPNVVAEV GGMRWMPATK DNTGGHVMV DKLVGELKLE SKNFPMGSNL PDKDPVGAKD NLFYLRGERF RLRDFTEAPD KIPYKLAWSE RGYGPEDLQV K VMHNIYPG FDKLSLAEQM QVKVFGKEIW RYGFWDLLYR VLSNEGYQFM KDAGGYEANV ANASAVTQLP ATEYSDKTVF LA LKKGFQA LPLTLAKRFA EVPGGLIAGE QRIRMNRRLA SVQFSDDTEY PYRLHFQATR TVDGKTSDVP GAEEIIHARQ VIL ALPRRS LELIQSPLFD DPWLKENIDS VLVQSAFKLF LAYEQPWWRS QGLVAGRSVT DLPIRQCYYM GTECEQDGGE KTLN SLLMA SYNDIGTVPF WKGLEDGAPF EGYQPKSLQG RIDANEVVPK MQYQISEEMV RIAQRQVTSL HDQIELPAPY SAVYH AWDA DPFGGGWHEW KANYRLDLII QRMRHPVQEQ EVYIVGEAYS YGQGWVEGAL TTAESTLQDF FGLPRPAWLP EAYQLL PAP APVDIDNPPA LACTDCKKTL TEVTEFAYTG IKA |
-Macromolecule #2: FLAVIN-ADENINE DINUCLEOTIDE
| Macromolecule | Name: FLAVIN-ADENINE DINUCLEOTIDE / type: ligand / ID: 2 / Number of copies: 1 / Formula: FAD |
|---|---|
| Molecular weight | Theoretical: 785.55 Da |
| Chemical component information |  ChemComp-FAD: |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 29 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: Quantifoil R2/2 / Material: MOLYBDENUM / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA KF80 |
- Electron microscopy
Electron microscopy
| Microscope | JEOL CRYO ARM 300 |
|---|---|
| Specialist optics | Energy filter - Name: In-column Omega Filter / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 5035 / Average exposure time: 8.0 sec. / Average electron dose: 69.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 3.4 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | Chain - Source name: AlphaFold / Chain - Initial model type: in silico model |
|---|---|
| Output model |  PDB-8jt7: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)