+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | ACE2-SIT1 complex bound with proline | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transporter / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsolute:sodium symporter activity / L-isoleucine import across plasma membrane / L-proline import across plasma membrane / Variant SLC6A20 affecting neurotransmitter transport contributes towards hyperglycinuria (HG) and iminoglycinuria (IG) / Variant SLC6A20 affecting amino acid transport contributes towards hyperglycinuria (HG) and iminoglycinuria (IG) / amino-acid betaine transport / proline import across plasma membrane / L-isoleucine transmembrane transporter activity / proline:sodium symporter activity / L-proline transmembrane transporter activity ...solute:sodium symporter activity / L-isoleucine import across plasma membrane / L-proline import across plasma membrane / Variant SLC6A20 affecting neurotransmitter transport contributes towards hyperglycinuria (HG) and iminoglycinuria (IG) / Variant SLC6A20 affecting amino acid transport contributes towards hyperglycinuria (HG) and iminoglycinuria (IG) / amino-acid betaine transport / proline import across plasma membrane / L-isoleucine transmembrane transporter activity / proline:sodium symporter activity / L-proline transmembrane transporter activity / amino-acid betaine transmembrane transporter activity / glycine transport / glycine import across plasma membrane / proline transport / amino acid import across plasma membrane / neutral L-amino acid transmembrane transporter activity / Amino acid transport across the plasma membrane / amino acid transmembrane transporter activity / SLC-mediated transport of neurotransmitters / positive regulation of amino acid transport / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / amino acid transport / positive regulation of gap junction assembly / tryptophan transport / regulation of systemic arterial blood pressure by renin-angiotensin / maternal process involved in female pregnancy / regulation of cardiac conduction / peptidyl-dipeptidase activity / regulation of vasoconstriction / transporter activator activity / Metabolism of Angiotensinogen to Angiotensins / transport across blood-brain barrier / carboxypeptidase activity / angiotensin maturation / viral life cycle / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / metallocarboxypeptidase activity / positive regulation of cardiac muscle contraction / regulation of cytokine production / blood vessel diameter maintenance / sodium ion transmembrane transport / negative regulation of smooth muscle cell proliferation / brush border membrane / negative regulation of ERK1 and ERK2 cascade / positive regulation of reactive oxygen species metabolic process / metallopeptidase activity / endocytic vesicle membrane / regulation of cell population proliferation / virus receptor activity / regulation of inflammatory response / endopeptidase activity / Induction of Cell-Cell Fusion / Potential therapeutics for SARS / membrane fusion / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / apical plasma membrane / cilium / membrane raft / endoplasmic reticulum lumen / symbiont entry into host cell / cell surface / negative regulation of transcription by RNA polymerase II / : / extracellular exosome / extracellular region / zinc ion binding / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Li YN / Zhang YY / Shen YP / Yan RH | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2023 Journal: Cell Discov / Year: 2023Title: Structural insight into the substrate recognition and transport mechanism of amino acid transporter complex ACE2-BAT1 and ACE2-SIT1. Authors: Yaning Li / Yiming Chen / Yuanyuan Zhang / Yaping Shen / Kangtai Xu / Yaqi Liu / Zilong Wang / Renhong Yan /  | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35254.map.gz emd_35254.map.gz | 85.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35254-v30.xml emd-35254-v30.xml emd-35254.xml emd-35254.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35254.png emd_35254.png | 52.9 KB | ||
| Filedesc metadata |  emd-35254.cif.gz emd-35254.cif.gz | 6.5 KB | ||
| Others |  emd_35254_half_map_1.map.gz emd_35254_half_map_1.map.gz emd_35254_half_map_2.map.gz emd_35254_half_map_2.map.gz | 84.1 MB 84.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35254 http://ftp.pdbj.org/pub/emdb/structures/EMD-35254 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35254 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35254 | HTTPS FTP |
-Related structure data
| Related structure data |  8i91MC  8i92C  8i93C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35254.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35254.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_35254_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_35254_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ACE2-SIT1 complex bound with proline
| Entire | Name: ACE2-SIT1 complex bound with proline |
|---|---|
| Components |
|
-Supramolecule #1: ACE2-SIT1 complex bound with proline
| Supramolecule | Name: ACE2-SIT1 complex bound with proline / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Angiotensin-converting enzyme 2
| Macromolecule | Name: Angiotensin-converting enzyme 2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: angiotensin-converting enzyme 2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 95.379828 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MRSSSSWLLL SLVAVTAAWS HPQFEKQSTI EEQAKTFLDK FNHEAEDLFY QSSLASWNYN TNITEENVQN MNNAGDKWSA FLKEQSTLA QMYPLQEIQN LTVKLQLQAL QQNGSSVLSE DKSKRLNTIL NTMSTIYSTG KVCNPDNPQE CLLLEPGLNE I MANSLDYN ...String: MRSSSSWLLL SLVAVTAAWS HPQFEKQSTI EEQAKTFLDK FNHEAEDLFY QSSLASWNYN TNITEENVQN MNNAGDKWSA FLKEQSTLA QMYPLQEIQN LTVKLQLQAL QQNGSSVLSE DKSKRLNTIL NTMSTIYSTG KVCNPDNPQE CLLLEPGLNE I MANSLDYN ERLWAWESWR SEVGKQLRPL YEEYVVLKNE MARANHYEDY GDYWRGDYEV NGVDGYDYSR GQLIEDVEHT FE EIKPLYE HLHAYVRAKL MNAYPSYISP IGCLPAHLLG DMWGRFWTNL YSLTVPFGQK PNIDVTDAMV DQAWDAQRIF KEA EKFFVS VGLPNMTQGF WENSMLTDPG NVQKAVCHPT AWDLGKGDFR ILMCTKVTMD DFLTAHHEMG HIQYDMAYAA QPFL LRNGA NEGFHEAVGE IMSLSAATPK HLKSIGLLSP DFQEDNETEI NFLLKQALTI VGTLPFTYML EKWRWMVFKG EIPKD QWMK KWWEMKREIV GVVEPVPHDE TYCDPASLFH VSNDYSFIRY YTRTLYQFQF QEALCQAAKH EGPLHKCDIS NSTEAG QKL FNMLRLGKSE PWTLALENVV GAKNMNVRPL LNYFEPLFTW LKDQNKNSFV GWSTDWSPYA DQSIKVRISL KSALGDK AY EWNDNEMYLF RSSVAYAMRQ YFLKVKNQMI LFGEEDVRVA NLKPRISFNF FVTAPKNVSD IIPRTEVEKA IRMSRSRI N DAFRLNDNSL EFLGIQPTLG PPNQPPVSIW LIVFGVVMGV IVVGIVILIF TGIRDRKKKN KARSGENPYA SIDISKGEN NPGFQNTDDV QTSFLEHHHH HHHHHH UniProtKB: Angiotensin-converting enzyme 2 |
-Macromolecule #2: SIT1
| Macromolecule | Name: SIT1 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 68.22482 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MADYKDDDDK SGPDEVDASG RMEKARPLWA NSLQFVFACI SYAVGLGNVW RFPYLCQMYG GGSFLVPYII MLIVEGMPLL YLELAVGQR MRQGSIGAWR TISPYLSGVG VASVVVSFFL SMYYNVINAW AFWYLFHSFQ DPLPWSVCPL NGNHTGYDEE C EKASSTQY ...String: MADYKDDDDK SGPDEVDASG RMEKARPLWA NSLQFVFACI SYAVGLGNVW RFPYLCQMYG GGSFLVPYII MLIVEGMPLL YLELAVGQR MRQGSIGAWR TISPYLSGVG VASVVVSFFL SMYYNVINAW AFWYLFHSFQ DPLPWSVCPL NGNHTGYDEE C EKASSTQY FWYRKTLNIS PSLQENGGVQ WEPALCLLLA WLVVYLCILR GTESTGKVVY FTASLPYCVL IIYLIRGLTL HG ATNGLMY MFTPKIEQLA NPKAWINAAT QIFFSLGLGF GSLIAFASYN EPSNNCQKHA IIVSLINSFT SIFASIVTFS IYG FKATFN YENCLKKVSL LLTNTFDLED GFLTASNLEQ VKGYLASAYP SKYSEMFPQI KNCSLESELD TAVQGTGLAF IVYT EAIKN MEVSQLWSVL YFFMLLMLGI GSMLGNTAAI LTPLTDSKII SSHLPKEAIS GLVCLVNCAI GMVFTMEAGN YWFDI FNDY AATLSLLLIV LVETIAVCYV YGLRRFESDL KAMTGRAVSW YWKVMWAGVS PLLIVSLFVF YLSDYILTGT LKYQAW DAS QGQLVTKDYP AYALAVIGLL VASSTMCIPL AALGTFVQRR LKRGDADPVA UniProtKB: Sodium- and chloride-dependent transporter XTRP3 |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 4 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #5: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #6: PROLINE
| Macromolecule | Name: PROLINE / type: ligand / ID: 6 / Number of copies: 2 / Formula: PRO |
|---|---|
| Molecular weight | Theoretical: 115.13 Da |
| Chemical component information | 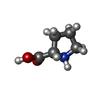 ChemComp-PRO: |
-Macromolecule #7: CHLORIDE ION
| Macromolecule | Name: CHLORIDE ION / type: ligand / ID: 7 / Number of copies: 2 / Formula: CL |
|---|---|
| Molecular weight | Theoretical: 35.453 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Material: GOLD |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.3 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. v4) / Number images used: 992519 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)