[English] 日本語
 Yorodumi
Yorodumi- EMDB-35234: Cryo-EM structure of Acipimox bound human hydroxy-carboxylic acid... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Acipimox bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer | |||||||||
 Map data Map data | sharpening map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / G-Protein / MEMBRANE PROTEIN / acipimox / signaling | |||||||||
| Function / homology |  Function and homology information Function and homology informationneutrophil apoptotic process / nicotinic acid receptor activity / Hydroxycarboxylic acid-binding receptors / positive regulation of neutrophil apoptotic process / Class A/1 (Rhodopsin-like receptors) / positive regulation of adiponectin secretion / negative regulation of lipid catabolic process / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration ...neutrophil apoptotic process / nicotinic acid receptor activity / Hydroxycarboxylic acid-binding receptors / positive regulation of neutrophil apoptotic process / Class A/1 (Rhodopsin-like receptors) / positive regulation of adiponectin secretion / negative regulation of lipid catabolic process / adenylate cyclase inhibitor activity / positive regulation of protein localization to cell cortex / T cell migration / positive regulation of relaxation of smooth muscle / Adenylate cyclase inhibitory pathway / D2 dopamine receptor binding / adenylate cyclase-inhibiting serotonin receptor signaling pathway / G protein-coupled serotonin receptor binding / cellular response to forskolin / regulation of mitotic spindle organization / chemokine-mediated signaling pathway / Regulation of insulin secretion / neuropeptide signaling pathway / response to prostaglandin E / positive regulation of cholesterol biosynthetic process / negative regulation of insulin secretion / G protein-coupled receptor binding / response to peptide hormone / centriolar satellite / G-protein beta/gamma-subunit complex binding / adenylate cyclase-modulating G protein-coupled receptor signaling pathway / adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G protein-coupled acetylcholine receptor signaling pathway / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / Glucagon signaling in metabolic regulation / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through CDC42 / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / cell junction / G beta:gamma signalling through BTK / photoreceptor disc membrane / ADP signalling through P2Y purinoceptor 12 / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / GDP binding / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / cellular response to catecholamine stimulus / ADP signalling through P2Y purinoceptor 1 / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / GPER1 signaling / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / G alpha (12/13) signalling events / Inactivation, recovery and regulation of the phototransduction cascade / G-protein beta-subunit binding / extracellular vesicle / sensory perception of taste / sperm principal piece / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / adenylate cyclase-activating G protein-coupled receptor signaling pathway / retina development in camera-type eye / GTPase binding / fibroblast proliferation / G protein activity / midbody / Ca2+ pathway / cell cortex / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / Ras protein signal transduction / Extra-nuclear estrogen signaling / cell population proliferation / ciliary basal body / G protein-coupled receptor signaling pathway / cell division / lysosomal membrane / GTPase activity / centrosome / synapse / GTP binding / protein-containing complex binding / nucleolus / magnesium ion binding / Golgi apparatus / signal transduction Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.77 Å | |||||||||
 Authors Authors | Park JH / Ishimoto N / Park SY | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural basis for ligand recognition and signaling of hydroxy-carboxylic acid receptor 2. Authors: Jae-Hyun Park / Kouki Kawakami / Naito Ishimoto / Tatsuya Ikuta / Mio Ohki / Toru Ekimoto / Mitsunori Ikeguchi / Dong-Sun Lee / Young-Ho Lee / Jeremy R H Tame / Asuka Inoue / Sam-Yong Park /   Abstract: Hydroxycarboxylic acid receptors (HCAR1, HCAR2, and HCAR3) transduce G signaling upon biding to molecules such as lactic acid, butyric acid and 3-hydroxyoctanoic acid, which are associated with ...Hydroxycarboxylic acid receptors (HCAR1, HCAR2, and HCAR3) transduce G signaling upon biding to molecules such as lactic acid, butyric acid and 3-hydroxyoctanoic acid, which are associated with lipolytic and atherogenic activity, and neuroinflammation. Although many reports have elucidated the function of HCAR2 and its potential as a therapeutic target for treating not only dyslipidemia but also neuroimmune disorders such as multiple sclerosis and Parkinson's disease, the structural basis of ligand recognition and ligand-induced G-coupling remains unclear. Here we report three cryo-EM structures of the human HCAR2-G signaling complex, each bound with different ligands: niacin, acipimox or GSK256073. All three agonists are held in a deep pocket lined by residues that are not conserved in HCAR1 and HCAR3. A distinct hairpin loop at the HCAR2 N-terminus and extra-cellular loop 2 (ECL2) completely enclose the ligand. These structures also reveal the agonist-induced conformational changes propagated to the G-protein-coupling interface during activation. Collectively, the structures presented here are expected to help in the design of ligands specific for HCAR2, leading to new drugs for the treatment of various diseases such as dyslipidemia and inflammation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35234.map.gz emd_35234.map.gz | 28.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35234-v30.xml emd-35234-v30.xml emd-35234.xml emd-35234.xml | 26.4 KB 26.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_35234_fsc.xml emd_35234_fsc.xml | 6.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_35234.png emd_35234.png | 91.2 KB | ||
| Masks |  emd_35234_msk_1.map emd_35234_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-35234.cif.gz emd-35234.cif.gz | 7.3 KB | ||
| Others |  emd_35234_additional_1.map.gz emd_35234_additional_1.map.gz emd_35234_additional_2.map.gz emd_35234_additional_2.map.gz emd_35234_half_map_1.map.gz emd_35234_half_map_1.map.gz emd_35234_half_map_2.map.gz emd_35234_half_map_2.map.gz | 15.4 MB 27.1 MB 28.3 MB 28.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35234 http://ftp.pdbj.org/pub/emdb/structures/EMD-35234 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35234 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35234 | HTTPS FTP |
-Related structure data
| Related structure data |  8i7vMC  8i7wC  8k5bC  8k5cC  8k5dC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35234.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35234.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpening map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.245 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_35234_msk_1.map emd_35234_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: additional map
| File | emd_35234_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | additional map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: additional map
| File | emd_35234_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | additional map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_35234_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_35234_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Acipomox bound human hydroxy-carboxylic acid receptor 2 in comple...
| Entire | Name: Acipomox bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer |
|---|---|
| Components |
|
-Supramolecule #1: Acipomox bound human hydroxy-carboxylic acid receptor 2 in comple...
| Supramolecule | Name: Acipomox bound human hydroxy-carboxylic acid receptor 2 in complex with Gi heterotrimer type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 150 KDa |
-Macromolecule #1: Human hydroxycarboxylic acid receptor 2,Hydroxycarboxylic acid re...
| Macromolecule | Name: Human hydroxycarboxylic acid receptor 2,Hydroxycarboxylic acid receptor 2,Hydroxycarboxylic acid receptor 2 type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 54.320297 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPGAPADLED NWETLNDNLK VIEKADNAAQ VKDALTKMRA AALDAQKATP PKLEDKSPDS PEMKDFRHGF DILVGQIDDA LKLANEGKV KEAQAAAEQL KTTRNAYIQK YLEFMNRHHL QDHFLEIDKK NCCVFRDDFI VKVLPPVLGL EFIFGLLGNG L ALWIFCFH ...String: GPGAPADLED NWETLNDNLK VIEKADNAAQ VKDALTKMRA AALDAQKATP PKLEDKSPDS PEMKDFRHGF DILVGQIDDA LKLANEGKV KEAQAAAEQL KTTRNAYIQK YLEFMNRHHL QDHFLEIDKK NCCVFRDDFI VKVLPPVLGL EFIFGLLGNG L ALWIFCFH LKSWKSSRIF LFNLAVADFL LIICLPFLMD NYVRRWDWKF GDIPCRLMLF MLAMNRQGSI IFLTVVAVDR YF RVVHPHH ALNKISNRTA AIISCLLWGI TIGLTVHLLK KKMPIQNGGA NLCSSFSICH TFQWHEAMFL LEFFLPLGII LFC SARIIW SLRQRQMDRH AKIKRAITFI MVVAIVFVIC FLPSVVVRIR IFWLLHTSGT QNCEVYRSVD LAFFITLSFT YMNS MLDPV VYYFSSPSFP NFFSTLINRC LQRKMTGEPD NNRSTSVELT GDPNKTRGAP EALMANSGEP WSPSYLGPTS P UniProtKB: Hydroxycarboxylic acid receptor 2 |
-Macromolecule #2: Guanine nucleotide-binding protein G(i) subunit alpha-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(i) subunit alpha-1 type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.415031 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI ...String: MGCTLSAEDK AAVERSKMID RNLREDGEKA AREVKLLLLG AGESGKSTIV KQMKIIHEAG YSEEECKQYK AVVYSNTIQS IIAIIRAMG RLKIDFGDSA RADDARQLFV LAGAAEEGFM TAELAGVIKR LWKDSGVQAC FNRSREYQLN DSAAYYLNDL D RIAQPNYI PTQQDVLRTR VKTTGIVETH FTFKDLHFKM FDVGGQRSER KKWIHCFEGV TAIIFCVALS DYDLVLAEDE EM NRMHESM KLFDSICNNK WFTDTSIILF LNKKDLFEEK IKKSPLTICY PEYAGSNTYE EAAAYIQCQF EDLNKRKDTK EIY THFTCA TDTKNVQFVF DAVTDVIIKN NLKDCGLF UniProtKB: Guanine nucleotide-binding protein G(i) subunit alpha-1 |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.728152 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GPGSSGSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD ...String: GPGSSGSELD QLRQEAEQLK NQIRDARKAC ADATLSQITN NIDPVGRIQM RTRRTLRGHL AKIYAMHWGT DSRLLVSASQ DGKLIIWDS YTTNKVHAIP LRSSWVMTCA YAPSGNYVAC GGLDNICSIY NLKTREGNVR VSRELAGHTG YLSCCRFLDD N QIVTSSGD TTCALWDIET GQQTTTFTGH TGDVMSLSLA PDTRLFVSGA CDASAKLWDV REGMCRQTFT GHESDINAIC FF PNGNAFA TGSDDATCRL FDLRADQELM TYSHDNIICG ITSVSFSKSG RLLLAGYDDF NCNVWDALKA DRAGVLAGHD NRV SCLGVT DDGMAVATGS WDSFLKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 8.506765 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI LGSAGSAGSA UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.409588 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAALEVLFQ |
-Macromolecule #6: 5-methyl-4-oxidanyl-pyrazin-4-ium-2-carboxylic acid
| Macromolecule | Name: 5-methyl-4-oxidanyl-pyrazin-4-ium-2-carboxylic acid / type: ligand / ID: 6 / Number of copies: 1 / Formula: OJX |
|---|---|
| Molecular weight | Theoretical: 155.131 Da |
| Chemical component information | 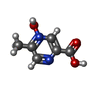 ChemComp-OJX: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Details: 20 mM HEPES pH8.0, 100 mM NaCl, 1 mM MgCl2, 0.5 mM TCEP, 0.001% LMNG, 0.0001% CHS |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.8 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-8i7v: |
 Movie
Movie Controller
Controller



































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)