[English] 日本語
 Yorodumi
Yorodumi- EMDB-34862: Cryo-EM Structures and Translocation Mechanism of Crenarchaeota R... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Sulfolobus acidocaldarius ribosome small subunit / RIBOSOME | |||||||||
| Function / homology |  Function and homology information Function and homology informationribonuclease P activity / tRNA 5'-leader removal / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA / positive regulation of apoptotic signaling pathway / ribosomal small subunit biogenesis / ribosome biogenesis / small ribosomal subunit rRNA binding / ribosomal small subunit assembly / small ribosomal subunit ...ribonuclease P activity / tRNA 5'-leader removal / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA / positive regulation of apoptotic signaling pathway / ribosomal small subunit biogenesis / ribosome biogenesis / small ribosomal subunit rRNA binding / ribosomal small subunit assembly / small ribosomal subunit / cytosolic small ribosomal subunit / cytoplasmic translation / tRNA binding / rRNA binding / ribosome / structural constituent of ribosome / ribonucleoprotein complex / translation / mRNA binding / RNA binding / zinc ion binding / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Sulfolobus acidocaldarius DSM 639 (acidophilic) Sulfolobus acidocaldarius DSM 639 (acidophilic) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.14 Å | |||||||||
 Authors Authors | Wang YH / Zhou J | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2023 Journal: Nucleic Acids Res / Year: 2023Title: Cryo-electron microscopy structure and translocation mechanism of the crenarchaeal ribosome. Authors: Ying-Hui Wang / Hong Dai / Ling Zhang / Yun Wu / Jingfen Wang / Chen Wang / Cai-Huang Xu / Hai Hou / Bing Yang / Yongqun Zhu / Xing Zhang / Jie Zhou /  Abstract: Archaeal ribosomes have many domain-specific features; however, our understanding of these structures is limited. We present 10 cryo-electron microscopy (cryo-EM) structures of the archaeal ribosome ...Archaeal ribosomes have many domain-specific features; however, our understanding of these structures is limited. We present 10 cryo-electron microscopy (cryo-EM) structures of the archaeal ribosome from crenarchaeota Sulfolobus acidocaldarius (Sac) at 2.7-5.7 Å resolution. We observed unstable conformations of H68 and h44 of ribosomal RNA (rRNA) in the subunit structures, which may interfere with subunit association. These subunit structures provided models for 12 rRNA expansion segments and 3 novel r-proteins. Furthermore, the 50S-aRF1 complex structure showed the unique domain orientation of aRF1, possibly explaining P-site transfer RNA (tRNA) release after translation termination. Sac 70S complexes were captured in seven distinct steps of the tRNA translocation reaction, confirming conserved structural features during archaeal ribosome translocation. In aEF2-engaged 70S ribosome complexes, 3D classification of cryo-EM data based on 30S head domain identified two new translocation intermediates with 30S head domain tilted 5-6° enabling its disengagement from the translocated tRNA and its release post-translocation. Additionally, we observed conformational changes to aEF2 during ribosome binding and switching from three different states. Our structural and biochemical data provide new insights into archaeal translation and ribosome translocation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_34862.map.gz emd_34862.map.gz | 10.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-34862-v30.xml emd-34862-v30.xml emd-34862.xml emd-34862.xml | 41.1 KB 41.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_34862.png emd_34862.png | 134.4 KB | ||
| Filedesc metadata |  emd-34862.cif.gz emd-34862.cif.gz | 9 KB | ||
| Others |  emd_34862_half_map_1.map.gz emd_34862_half_map_1.map.gz emd_34862_half_map_2.map.gz emd_34862_half_map_2.map.gz | 126.8 MB 127.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-34862 http://ftp.pdbj.org/pub/emdb/structures/EMD-34862 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34862 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-34862 | HTTPS FTP |
-Validation report
| Summary document |  emd_34862_validation.pdf.gz emd_34862_validation.pdf.gz | 826.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_34862_full_validation.pdf.gz emd_34862_full_validation.pdf.gz | 826.5 KB | Display | |
| Data in XML |  emd_34862_validation.xml.gz emd_34862_validation.xml.gz | 15.4 KB | Display | |
| Data in CIF |  emd_34862_validation.cif.gz emd_34862_validation.cif.gz | 18.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34862 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34862 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34862 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-34862 | HTTPS FTP |
-Related structure data
| Related structure data |  8hkxMC  8hkuC  8hkvC  8hkyC  8hkzC  8hl1C  8hl2C  8hl3C  8hl4C  8hl5C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_34862.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_34862.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_34862_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
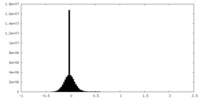
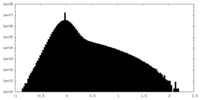
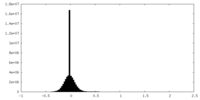
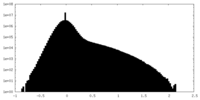
| Density Histograms |
-Half map: #1
| File | emd_34862_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Sulfolobus acidocaldarius ribosome small sbunit
+Supramolecule #1: Sulfolobus acidocaldarius ribosome small sbunit
+Macromolecule #1: 16s rRNA (1491-MER)
+Macromolecule #2: 30S ribosomal protein S2
+Macromolecule #3: 30S ribosomal protein S4e
+Macromolecule #4: 30S ribosomal protein S4
+Macromolecule #5: 30S ribosomal protein S5
+Macromolecule #6: 30S ribosomal protein S6e
+Macromolecule #7: 30S ribosomal protein S8e
+Macromolecule #8: 30S ribosomal protein S8
+Macromolecule #9: 30S ribosomal protein S11
+Macromolecule #10: 30S ribosomal protein S12
+Macromolecule #11: 30S ribosomal protein S15
+Macromolecule #12: 30S ribosomal protein S17
+Macromolecule #13: 30S ribosomal protein S24e
+Macromolecule #14: 30S ribosomal protein S27e
+Macromolecule #15: 30S ribosomal protein S3Ae
+Macromolecule #16: 30S ribosomal protein S3
+Macromolecule #17: 30S ribosomal protein S7
+Macromolecule #18: 30S ribosomal protein S9
+Macromolecule #19: 30S ribosomal protein S10
+Macromolecule #20: 30S ribosomal protein S13
+Macromolecule #21: 30S ribosomal protein S14 type Z
+Macromolecule #22: 30S ribosomal protein S17e
+Macromolecule #23: 30S ribosomal protein S19e
+Macromolecule #24: 30S ribosomal protein S19
+Macromolecule #25: 30S ribosomal protein S28e
+Macromolecule #26: 50S ribosomal protein L7Ae
+Macromolecule #27: 30S ribosomal protein S27ae
+Macromolecule #28: 30S ribosomal protein
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 26.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID:  3f1e |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.14 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 98366 |
| Initial angle assignment | Type: NOT APPLICABLE |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)