[English] 日本語
 Yorodumi
Yorodumi- EMDB-33665: Structure of the Bacterial Ribosome with human tRNA Tyr(GalQ34) a... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the Bacterial Ribosome with human tRNA Tyr(GalQ34) and mRNA(UAC) | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | RIBOSOME / tRNA modifications / decoding | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of cytoplasmic translational initiation / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / misfolded RNA binding / Group I intron splicing / RNA folding / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity ...negative regulation of cytoplasmic translational initiation / transcription antitermination factor activity, RNA binding / ornithine decarboxylase inhibitor activity / misfolded RNA binding / Group I intron splicing / RNA folding / transcriptional attenuation / endoribonuclease inhibitor activity / positive regulation of ribosome biogenesis / RNA-binding transcription regulator activity / four-way junction DNA binding / negative regulation of cytoplasmic translation / DnaA-L2 complex / regulation of mRNA stability / translation repressor activity / negative regulation of translational initiation / negative regulation of DNA-templated DNA replication initiation / mRNA regulatory element binding translation repressor activity / regulation of DNA-templated transcription elongation / positive regulation of RNA splicing / transcription elongation factor complex / response to reactive oxygen species / cytosolic ribosome assembly / ribosome assembly / assembly of large subunit precursor of preribosome / transcription antitermination / DNA endonuclease activity / translational initiation / regulation of cell growth / DNA-templated transcription termination / response to radiation / maintenance of translational fidelity / mRNA 5'-UTR binding / regulation of translation / large ribosomal subunit / ribosomal small subunit assembly / transferase activity / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / response to antibiotic / negative regulation of DNA-templated transcription / hydrolase activity / mRNA binding / DNA binding / RNA binding / zinc ion binding / membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||||||||
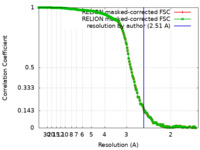
| Method | single particle reconstruction / cryo EM / Resolution: 2.51 Å | ||||||||||||
 Authors Authors | Ishiguro K / Yokoyama T / Shirouzu M / Suzuki T | ||||||||||||
| Funding support |  Japan, 3 items Japan, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell / Year: 2023 Journal: Cell / Year: 2023Title: Glycosylated queuosines in tRNAs optimize translational rate and post-embryonic growth. Authors: Xuewei Zhao / Ding Ma / Kensuke Ishiguro / Hironori Saito / Shinichiro Akichika / Ikuya Matsuzawa / Mari Mito / Toru Irie / Kota Ishibashi / Kimi Wakabayashi / Yuriko Sakaguchi / Takeshi ...Authors: Xuewei Zhao / Ding Ma / Kensuke Ishiguro / Hironori Saito / Shinichiro Akichika / Ikuya Matsuzawa / Mari Mito / Toru Irie / Kota Ishibashi / Kimi Wakabayashi / Yuriko Sakaguchi / Takeshi Yokoyama / Yuichiro Mishima / Mikako Shirouzu / Shintaro Iwasaki / Takeo Suzuki / Tsutomu Suzuki /  Abstract: Transfer RNA (tRNA) modifications are critical for protein synthesis. Queuosine (Q), a 7-deaza-guanosine derivative, is present in tRNA anticodons. In vertebrate tRNAs for Tyr and Asp, Q is further ...Transfer RNA (tRNA) modifications are critical for protein synthesis. Queuosine (Q), a 7-deaza-guanosine derivative, is present in tRNA anticodons. In vertebrate tRNAs for Tyr and Asp, Q is further glycosylated with galactose and mannose to generate galQ and manQ, respectively. However, biogenesis and physiological relevance of Q-glycosylation remain poorly understood. Here, we biochemically identified two RNA glycosylases, QTGAL and QTMAN, and successfully reconstituted Q-glycosylation of tRNAs using nucleotide diphosphate sugars. Ribosome profiling of knockout cells revealed that Q-glycosylation slowed down elongation at cognate codons, UAC and GAC (GAU), respectively. We also found that galactosylation of Q suppresses stop codon readthrough. Moreover, protein aggregates increased in cells lacking Q-glycosylation, indicating that Q-glycosylation contributes to proteostasis. Cryo-EM of human ribosome-tRNA complex revealed the molecular basis of codon recognition regulated by Q-glycosylations. Furthermore, zebrafish qtgal and qtman knockout lines displayed shortened body length, implying that Q-glycosylation is required for post-embryonic growth in vertebrates. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33665.map.gz emd_33665.map.gz | 531.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33665-v30.xml emd-33665-v30.xml emd-33665.xml emd-33665.xml | 94.6 KB 94.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33665_fsc.xml emd_33665_fsc.xml emd_33665_fsc_2.xml emd_33665_fsc_2.xml | 18.7 KB 18.7 KB | Display Display |  FSC data file FSC data file |
| Images |  emd_33665.png emd_33665.png | 119.3 KB | ||
| Filedesc metadata |  emd-33665.cif.gz emd-33665.cif.gz | 16.7 KB | ||
| Others |  emd_33665_additional_1.map.gz emd_33665_additional_1.map.gz emd_33665_half_map_1.map.gz emd_33665_half_map_1.map.gz emd_33665_half_map_2.map.gz emd_33665_half_map_2.map.gz | 455.5 MB 457.1 MB 456.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33665 http://ftp.pdbj.org/pub/emdb/structures/EMD-33665 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33665 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33665 | HTTPS FTP |
-Related structure data
| Related structure data |  7y7hMC  7y7cC  7y7dC  7y7eC  7y7fC  7y7gC  8jdjC  8jdkC  8jdlC  8jdmC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33665.map.gz / Format: CCP4 / Size: 567.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33665.map.gz / Format: CCP4 / Size: 567.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8285 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: before postprocess
| File | emd_33665_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | before postprocess | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_33665_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_33665_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : The complex of E. coli 70S ribosome with mRNA and A-, P- site tRNA
+Supramolecule #1: The complex of E. coli 70S ribosome with mRNA and A-, P- site tRNA
+Supramolecule #2: E. coli 70S ribosome
+Supramolecule #3: mRNA
+Supramolecule #4: A-site tRNA
+Supramolecule #5: P-site tRNA
+Macromolecule #1: 16S rRNA
+Macromolecule #22: 23S rRNA
+Macromolecule #23: 5S rRNA
+Macromolecule #53: mRNA
+Macromolecule #54: P-site tRNA-fMet
+Macromolecule #55: A-site tRNA-Tyr
+Macromolecule #2: 30S ribosomal protein S2
+Macromolecule #3: 30S ribosomal protein S3
+Macromolecule #4: 30S ribosomal protein S4
+Macromolecule #5: 30S ribosomal protein S5
+Macromolecule #6: 30S ribosomal protein S6, fully modified isoform
+Macromolecule #7: 30S ribosomal protein S7
+Macromolecule #8: 30S ribosomal protein S8
+Macromolecule #9: 30S ribosomal protein S9
+Macromolecule #10: 30S ribosomal protein S10
+Macromolecule #11: 30S ribosomal protein S11
+Macromolecule #12: 30S ribosomal protein S12
+Macromolecule #13: 30S ribosomal protein S13
+Macromolecule #14: 30S ribosomal protein S14
+Macromolecule #15: 30S ribosomal protein S15
+Macromolecule #16: 30S ribosomal protein S16
+Macromolecule #17: 30S ribosomal protein S17
+Macromolecule #18: 30S ribosomal protein S18
+Macromolecule #19: 30S ribosomal protein S19
+Macromolecule #20: 30S ribosomal protein S20
+Macromolecule #21: 30S ribosomal protein S21
+Macromolecule #24: 50S ribosomal protein L2
+Macromolecule #25: 50S ribosomal protein L3
+Macromolecule #26: 50S ribosomal protein L4
+Macromolecule #27: 50S ribosomal protein L5
+Macromolecule #28: 50S ribosomal protein L6
+Macromolecule #29: 50S ribosomal protein L9
+Macromolecule #30: 50S ribosomal protein L13
+Macromolecule #31: 50S ribosomal protein L14
+Macromolecule #32: 50S ribosomal protein L15
+Macromolecule #33: 50S ribosomal protein L16
+Macromolecule #34: 50S ribosomal protein L17
+Macromolecule #35: 50S ribosomal protein L18
+Macromolecule #36: 50S ribosomal protein L19
+Macromolecule #37: 50S ribosomal protein L20
+Macromolecule #38: 50S ribosomal protein L21
+Macromolecule #39: 50S ribosomal protein L22
+Macromolecule #40: 50S ribosomal protein L23
+Macromolecule #41: 50S ribosomal protein L24
+Macromolecule #42: 50S ribosomal protein L25
+Macromolecule #43: 50S ribosomal protein L27
+Macromolecule #44: 50S ribosomal protein L28
+Macromolecule #45: 50S ribosomal protein L29
+Macromolecule #46: 50S ribosomal protein L30
+Macromolecule #47: 50S ribosomal protein L32
+Macromolecule #48: 50S ribosomal protein L33
+Macromolecule #49: 50S ribosomal protein L34
+Macromolecule #50: 50S ribosomal protein L35
+Macromolecule #51: 50S ribosomal protein L36
+Macromolecule #52: 50S ribosomal protein L31
+Macromolecule #56: MAGNESIUM ION
+Macromolecule #57: beta-D-galactopyranose
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 Component:
Details: The Buffer pH was adjusted to 7.6 using KOH. | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||
| Details | 100nM ribosomes were incubated with 500nM tRNAs and mRNA |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 7237 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)