+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CRYO-EM STRUCTURE OF IMPORTIN ALPHA1/BETA HETERODIMER | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Importins / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationRNA import into nucleus / Inhibition of nitric oxide production / mitotic chromosome movement towards spindle pole / Sensing of DNA Double Strand Breaks / endoplasmic reticulum tubular network / regulation of DNA recombination / establishment of mitotic spindle localization / entry of viral genome into host nucleus through nuclear pore complex via importin / positive regulation of viral life cycle / astral microtubule organization ...RNA import into nucleus / Inhibition of nitric oxide production / mitotic chromosome movement towards spindle pole / Sensing of DNA Double Strand Breaks / endoplasmic reticulum tubular network / regulation of DNA recombination / establishment of mitotic spindle localization / entry of viral genome into host nucleus through nuclear pore complex via importin / positive regulation of viral life cycle / astral microtubule organization / Transport of Ribonucleoproteins into the Host Nucleus / Regulation of cholesterol biosynthesis by SREBP (SREBF) / importin-alpha family protein binding / NS1 Mediated Effects on Host Pathways / NLS-dependent protein nuclear import complex / Apoptosis induced DNA fragmentation / Initiation of Nuclear Envelope (NE) Reformation / Nuclear import of Rev protein / ribosomal protein import into nucleus / Postmitotic nuclear pore complex (NPC) reformation / nuclear import signal receptor activity / nuclear localization sequence binding / NLS-bearing protein import into nucleus / CREB1 phosphorylation through the activation of CaMKII/CaMKK/CaMKIV cascasde / mitotic metaphase chromosome alignment / CaMK IV-mediated phosphorylation of CREB / DNA metabolic process / Maturation of hRSV A proteins / mitotic spindle assembly / nuclear pore / positive regulation of type I interferon production / Assembly of the ORC complex at the origin of replication / Hsp90 protein binding / positive regulation of cholesterol biosynthetic process / specific granule lumen / ISG15 antiviral mechanism / small GTPase binding / histone deacetylase binding / cytoplasmic stress granule / protein import into nucleus / SARS-CoV-1 activates/modulates innate immune responses / Interferon alpha/beta signaling / nuclear envelope / host cell / nuclear membrane / Estrogen-dependent gene expression / ficolin-1-rich granule lumen / Golgi membrane / protein domain specific binding / Neutrophil degranulation / endoplasmic reticulum membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / enzyme binding / RNA binding / extracellular exosome / extracellular region / zinc ion binding / nucleoplasm / membrane / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.95 Å | |||||||||
 Authors Authors | Ko Y / Cingolani G | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2024 Journal: Sci Adv / Year: 2024Title: Structural basis for nuclear import of hepatitis B virus (HBV) nucleocapsid core. Authors: Ruoyu Yang / Ying-Hui Ko / Fenglin Li / Ravi K Lokareddy / Chun-Feng David Hou / Christine Kim / Shelby Klein / Santiago Antolínez / Juan F Marín / Carolina Pérez-Segura / Martin F ...Authors: Ruoyu Yang / Ying-Hui Ko / Fenglin Li / Ravi K Lokareddy / Chun-Feng David Hou / Christine Kim / Shelby Klein / Santiago Antolínez / Juan F Marín / Carolina Pérez-Segura / Martin F Jarrold / Adam Zlotnick / Jodi A Hadden-Perilla / Gino Cingolani /  Abstract: Nuclear import of the hepatitis B virus (HBV) nucleocapsid is essential for replication that occurs in the nucleus. The ~360-angstrom HBV capsid translocates to the nuclear pore complex (NPC) as an ...Nuclear import of the hepatitis B virus (HBV) nucleocapsid is essential for replication that occurs in the nucleus. The ~360-angstrom HBV capsid translocates to the nuclear pore complex (NPC) as an intact particle, hijacking human importins in a reaction stimulated by host kinases. This paper describes the mechanisms of HBV capsid recognition by importins. We found that importin α1 binds a nuclear localization signal (NLS) at the far end of the HBV coat protein Cp183 carboxyl-terminal domain (CTD). This NLS is exposed to the capsid surface through a pore at the icosahedral quasi-sixfold vertex. Phosphorylation at serine-155, serine-162, and serine-170 promotes CTD compaction but does not affect the affinity for importin α1. The binding of 30 importin α1/β1 augments HBV capsid diameter to ~620 angstroms, close to the maximum size trafficable through the NPC. We propose that phosphorylation favors CTD externalization and prompts its compaction at the capsid surface, exposing the NLS to importins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29936.map.gz emd_29936.map.gz | 16.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29936-v30.xml emd-29936-v30.xml emd-29936.xml emd-29936.xml | 17.8 KB 17.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_29936.png emd_29936.png | 104 KB | ||
| Filedesc metadata |  emd-29936.cif.gz emd-29936.cif.gz | 6.5 KB | ||
| Others |  emd_29936_half_map_1.map.gz emd_29936_half_map_1.map.gz emd_29936_half_map_2.map.gz emd_29936_half_map_2.map.gz | 31.9 MB 31.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29936 http://ftp.pdbj.org/pub/emdb/structures/EMD-29936 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29936 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29936 | HTTPS FTP |
-Related structure data
| Related structure data |  8gcnMC  7umiC  8g5vC  8g6vC  8g8yC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29936.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29936.map.gz / Format: CCP4 / Size: 34.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.872 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_29936_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
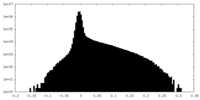
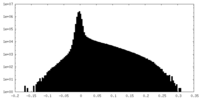
| Density Histograms |
-Half map: #1
| File | emd_29936_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Importin Alpha1/Beta Heterodimer
| Entire | Name: Importin Alpha1/Beta Heterodimer |
|---|---|
| Components |
|
-Supramolecule #1: Importin Alpha1/Beta Heterodimer
| Supramolecule | Name: Importin Alpha1/Beta Heterodimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Importin subunit beta-1
| Macromolecule | Name: Importin subunit beta-1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 97.257812 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MELITILEKT VSPDRLELEA AQKFLERAAV ENLPTFLVEL SRVLANPGNS QVARVAAGLQ IKNSLTSKDP DIKAQYQQRW LAIDANARR EVKNYVLQTL GTETYRPSSA SQCVAGIACA EIPVNQWPEL IPQLVANVTN PNSTEHMKES TLEAIGYICQ D IDPEQLQD ...String: MELITILEKT VSPDRLELEA AQKFLERAAV ENLPTFLVEL SRVLANPGNS QVARVAAGLQ IKNSLTSKDP DIKAQYQQRW LAIDANARR EVKNYVLQTL GTETYRPSSA SQCVAGIACA EIPVNQWPEL IPQLVANVTN PNSTEHMKES TLEAIGYICQ D IDPEQLQD KSNEILTAII QGMRKEEPSN NVKLAATNAL LNSLEFTKAN FDKESERHFI MQVVCEATQC PDTRVRVAAL QN LVKIMSL YYQYMETYMG PALFAITIEA MKSDIDEVAL QGIEFWSNVC DEEMDLAIEA SEAAEQGRPP EHTSKFYAKG ALQ YLVPIL TQTLTKQDEN DDDDDWNPCK AAGVCLMLLA TCCEDDIVPH VLPFIKEHIK NPDWRYRDAA VMAFGCILEG PEPS QLKPL VIQAMPTLIE LMKDPSVVVR DTAAWTVGRI CELLPEAAIN DVYLAPLLQC LIEGLSAEPR VASNVCWAFS SLAEA AYEA ADVADDQEEP ATYCLSSSFE LIVQKLLETT DRPDGHQNNL RSSAYESLME IVKNSAKDCY PAVQKTTLVI MERLQQ VLQ MESHIQSTSD RIQFNDLQSL LCATLQNVLR KVQHQDALQI SDVVMASLLR MFQSTAGSGG VQEDALMAVS TLVEVLG GE FLKYMEAFKP FLGIGLKNYA EYQVCLAAVG LVGDLCRALQ SNIIPFCDEV MQLLLENLGN ENVHRSVKPQ ILSVFGDI A LAIGGEFKKY LEVVLNTLQQ ASQAQVDKSD YDMVDYLNEL RESCLEAYTG IVQGLKGDQE NVHPDVMLVQ PRVEFILSF IDHIAGDEDH TDGVVACAAG LIGDLCTAFG KDVLKLVEAR PMIHELLTEG RRSKTNKAKT LATWATKELR KLKNQA UniProtKB: Importin subunit beta-1 |
-Macromolecule #2: Importin subunit alpha-1
| Macromolecule | Name: Importin subunit alpha-1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 5.296262 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AARLHRFKNK GKDSTEMRRR RIEVNVELRK AKKDDQMLKR RNV UniProtKB: Importin subunit alpha-1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)