+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | C3HR3_9r_shift4: Extendable repeat protein fiber | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Oligomer / repeat protein / DE NOVO PROTEIN | |||||||||
| Biological species | unidentified (others) / synthetic construct (others) | |||||||||
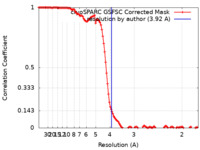
| Method | helical reconstruction / cryo EM / Resolution: 3.92 Å | |||||||||
 Authors Authors | Bethel NP / Borst AJ | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Chem / Year: 2023 Journal: Nat Chem / Year: 2023Title: Precisely patterned nanofibres made from extendable protein multiplexes. Authors: Neville P Bethel / Andrew J Borst / Fabio Parmeggiani / Matthew J Bick / T J Brunette / Hannah Nguyen / Alex Kang / Asim K Bera / Lauren Carter / Marcos C Miranda / Ryan D Kibler / Mila Lamb ...Authors: Neville P Bethel / Andrew J Borst / Fabio Parmeggiani / Matthew J Bick / T J Brunette / Hannah Nguyen / Alex Kang / Asim K Bera / Lauren Carter / Marcos C Miranda / Ryan D Kibler / Mila Lamb / Xinting Li / Banumathi Sankaran / David Baker /   Abstract: Molecular systems with coincident cyclic and superhelical symmetry axes have considerable advantages for materials design as they can be readily lengthened or shortened by changing the length of the ...Molecular systems with coincident cyclic and superhelical symmetry axes have considerable advantages for materials design as they can be readily lengthened or shortened by changing the length of the constituent monomers. Among proteins, alpha-helical coiled coils have such symmetric, extendable architectures, but are limited by the relatively fixed geometry and flexibility of the helical protomers. Here we describe a systematic approach to generating modular and rigid repeat protein oligomers with coincident C to C and superhelical symmetry axes that can be readily extended by repeat propagation. From these building blocks, we demonstrate that a wide range of unbounded fibres can be systematically designed by introducing hydrophilic surface patches that force staggering of the monomers; the geometry of such fibres can be precisely tuned by varying the number of repeat units in the monomer and the placement of the hydrophilic patches. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29856.map.gz emd_29856.map.gz | 87 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29856-v30.xml emd-29856-v30.xml emd-29856.xml emd-29856.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29856_fsc.xml emd_29856_fsc.xml | 11.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_29856.png emd_29856.png | 55.3 KB | ||
| Filedesc metadata |  emd-29856.cif.gz emd-29856.cif.gz | 6.1 KB | ||
| Others |  emd_29856_additional_1.map.gz emd_29856_additional_1.map.gz emd_29856_half_map_1.map.gz emd_29856_half_map_1.map.gz emd_29856_half_map_2.map.gz emd_29856_half_map_2.map.gz | 14.1 MB 157 MB 157 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29856 http://ftp.pdbj.org/pub/emdb/structures/EMD-29856 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29856 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29856 | HTTPS FTP |
-Related structure data
| Related structure data |  8g8iMC  8eovC  8eoxC  8eozC  8erwC  8ga9C  8gaaC  8gaqC C: citing same article ( M: atomic model generated by this map |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_29856.map.gz / Format: CCP4 / Size: 169.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29856.map.gz / Format: CCP4 / Size: 169.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.885 Å | ||||||||||||||||||||||||||||||||||||
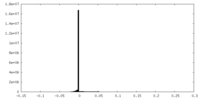
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unsharpened map
| File | emd_29856_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
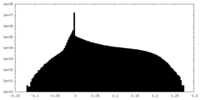
| Density Histograms |
-Half map: Half map A
| File | emd_29856_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
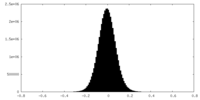
| Density Histograms |
-Half map: Half map B
| File | emd_29856_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
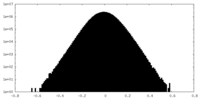
| Density Histograms |
- Sample components
Sample components
-Entire : C3HR3_8r
| Entire | Name: C3HR3_8r |
|---|---|
| Components |
|
-Supramolecule #1: C3HR3_8r
| Supramolecule | Name: C3HR3_8r / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 141 KDa |
-Macromolecule #1: C3HR3_9r_shift4
| Macromolecule | Name: C3HR3_9r_shift4 / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 55.922121 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVYERLKELI EENPEIQEIL ELWKFIGRED VVEKLFEVIK WAVEEGGNNM VFISLLKEIL SNPEIFEILL KWVDIGREDV VKKLFSVIK QAVEDGGNNM VFLSLLKELL SNPEIFEILL LWVEINREDV VKKFFDVIKW AVERGGNNPK FLSLLKLILS N PEIFKILL ...String: MVYERLKELI EENPEIQEIL ELWKFIGRED VVEKLFEVIK WAVEEGGNNM VFISLLKEIL SNPEIFEILL KWVDIGREDV VKKLFSVIK QAVEDGGNNM VFLSLLKELL SNPEIFEILL LWVEINREDV VKKFFDVIKW AVERGGNNPK FLSLLKLILS N PEIFKILL LWVYENKEEV VKKLFDVLLY AVKEGGDNEK FLELLKEILS NPEIFEILLE WVEENKEDVV KKLFDVIKYA VE RGGDNEK FYELLKLLLS NPEIFKILLE WVEINKEDVV KKFFDVIKWA VEEGGDNPVF YRLLKLILSN PEIFKILLEW VDI NNEDVV KKLFSVIKQA VEDGGNNMVF IDLLELILSN PEIFEILLLW VHINREDVVK KLFDVIKWAV EDGGNNMVFI HLLR KLLSN PEIFEILLLW VDIGREEVVK KFFEKVLEAV KEGGNDMIEI KKLEEILLDP EKFEELLLEV GSLEHHHHHH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)