+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of E.coli transcription elongation complex | ||||||||||||||||||
 Map data Map data | W6_refine122 | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Transcription elongation / TRANSCRIPTION-DNA-RNA complex | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationRNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly / bacterial-type flagellum-dependent cell motility / nitrate assimilation / transcription elongation factor complex / regulation of DNA-templated transcription elongation ...RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / cytosolic DNA-directed RNA polymerase complex / regulation of DNA-templated transcription initiation / bacterial-type flagellum assembly / bacterial-type flagellum-dependent cell motility / nitrate assimilation / transcription elongation factor complex / regulation of DNA-templated transcription elongation / transcription antitermination / DNA-templated transcription initiation / cell motility / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / response to heat / protein-containing complex assembly / intracellular iron ion homeostasis / protein dimerization activity / response to antibiotic / magnesium ion binding / DNA binding / zinc ion binding / membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |   | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.42 Å | ||||||||||||||||||
 Authors Authors | Duan W / Serganov A | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Control of transcription elongation and DNA repair by alarmone ppGpp. Authors: Jacob W Weaver / Sergey Proshkin / Wenqian Duan / Vitaly Epshtein / Manjunath Gowder / Binod K Bharati / Elena Afanaseva / Alexander Mironov / Alexander Serganov / Evgeny Nudler /   Abstract: Second messenger (p)ppGpp (collectively guanosine tetraphosphate and guanosine pentaphosphate) mediates bacterial adaptation to nutritional stress by modulating transcription initiation. More ...Second messenger (p)ppGpp (collectively guanosine tetraphosphate and guanosine pentaphosphate) mediates bacterial adaptation to nutritional stress by modulating transcription initiation. More recently, ppGpp has been implicated in coupling transcription and DNA repair; however, the mechanism of ppGpp engagement remained elusive. Here we present structural, biochemical and genetic evidence that ppGpp controls Escherichia coli RNA polymerase (RNAP) during elongation via a specific site that is nonfunctional during initiation. Structure-guided mutagenesis renders the elongation (but not initiation) complex unresponsive to ppGpp and increases bacterial sensitivity to genotoxic agents and ultraviolet radiation. Thus, ppGpp binds RNAP at sites with distinct functions in initiation and elongation, with the latter being important for promoting DNA repair. Our data provide insights on the molecular mechanism of ppGpp-mediated adaptation during stress, and further highlight the intricate relationships between genome stability, stress responses and transcription. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29491.map.gz emd_29491.map.gz | 230.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29491-v30.xml emd-29491-v30.xml emd-29491.xml emd-29491.xml | 28.4 KB 28.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_29491.png emd_29491.png | 51.7 KB | ||
| Filedesc metadata |  emd-29491.cif.gz emd-29491.cif.gz | 9 KB | ||
| Others |  emd_29491_half_map_1.map.gz emd_29491_half_map_1.map.gz emd_29491_half_map_2.map.gz emd_29491_half_map_2.map.gz | 225.9 MB 225.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29491 http://ftp.pdbj.org/pub/emdb/structures/EMD-29491 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29491 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29491 | HTTPS FTP |
-Validation report
| Summary document |  emd_29491_validation.pdf.gz emd_29491_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29491_full_validation.pdf.gz emd_29491_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_29491_validation.xml.gz emd_29491_validation.xml.gz | 15.4 KB | Display | |
| Data in CIF |  emd_29491_validation.cif.gz emd_29491_validation.cif.gz | 18.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29491 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29491 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29491 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29491 | HTTPS FTP |
-Related structure data
| Related structure data |  8fvrMC  8fvwC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29491.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29491.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | W6_refine122 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_29491_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
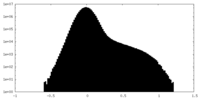
| Projections & Slices |
| ||||||||||||
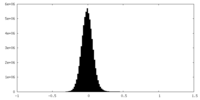
| Density Histograms |
-Half map: #1
| File | emd_29491_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
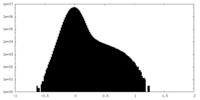
| Projections & Slices |
| ||||||||||||
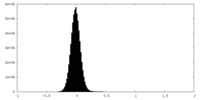
| Density Histograms |
- Sample components
Sample components
+Entire : E. coli transcription elongation complex (EC)
+Supramolecule #1: E. coli transcription elongation complex (EC)
+Macromolecule #1: DNA-directed RNA polymerase subunit alpha
+Macromolecule #2: DNA-directed RNA polymerase subunit beta
+Macromolecule #3: DNA-directed RNA polymerase subunit beta'
+Macromolecule #4: DNA-directed RNA polymerase subunit omega
+Macromolecule #5: DNA (53-MER)
+Macromolecule #7: DNA (53-MER)
+Macromolecule #6: RNA (16-MER)
+Macromolecule #8: ZINC ION
+Macromolecule #9: MAGNESIUM ION
+Macromolecule #10: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6.5 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM HEPES, pH 8.0, 150 mM NaCl, 10 mM MgCl2, 10 uM ZnCl2, 5 mM DTT |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 260.0 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 79.95 K / Max: 79.95 K |
| Alignment procedure | Coma free - Residual tilt: 0.05 mrad |
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number grids imaged: 1 / Number real images: 3298 / Average exposure time: 2.0 sec. / Average electron dose: 59.07 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8fvr: |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X