+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of miniGq-coupled hM3Dq in complex with DCZ | |||||||||
 Map data Map data | CryoEM structure of Gq-coupled hM3Dq in complex with DCZ | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GPCR / Deschloroclozapine / DCZ / active state / hM3Dq / hM3R / DREADD / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationsaliva secretion / phospholipase C-activating G protein-coupled acetylcholine receptor signaling pathway / Acetylcholine regulates insulin secretion / Muscarinic acetylcholine receptors / G protein-coupled acetylcholine receptor activity / positive regulation of smooth muscle contraction / regulation of smooth muscle contraction / phosphatidylinositol-4,5-bisphosphate phospholipase C activity / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway / acetylcholine binding ...saliva secretion / phospholipase C-activating G protein-coupled acetylcholine receptor signaling pathway / Acetylcholine regulates insulin secretion / Muscarinic acetylcholine receptors / G protein-coupled acetylcholine receptor activity / positive regulation of smooth muscle contraction / regulation of smooth muscle contraction / phosphatidylinositol-4,5-bisphosphate phospholipase C activity / adenylate cyclase-inhibiting G protein-coupled acetylcholine receptor signaling pathway / acetylcholine binding / acetylcholine receptor signaling pathway / ligand-gated ion channel signaling pathway / smooth muscle contraction / G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger / basal plasma membrane / calcium-mediated signaling / positive regulation of insulin secretion / protein modification process / Olfactory Signaling Pathway / Activation of the phototransduction cascade / G beta:gamma signalling through PLC beta / Presynaptic function of Kainate receptors / Thromboxane signalling through TP receptor / G protein-coupled acetylcholine receptor signaling pathway / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / G-protein activation / G beta:gamma signalling through CDC42 / Prostacyclin signalling through prostacyclin receptor / Glucagon signaling in metabolic regulation / G beta:gamma signalling through BTK / Synthesis, secretion, and inactivation of Glucagon-like Peptide-1 (GLP-1) / ADP signalling through P2Y purinoceptor 12 / photoreceptor disc membrane / Sensory perception of sweet, bitter, and umami (glutamate) taste / Glucagon-type ligand receptors / Adrenaline,noradrenaline inhibits insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / G alpha (z) signalling events / ADP signalling through P2Y purinoceptor 1 / cellular response to catecholamine stimulus / ADORA2B mediated anti-inflammatory cytokines production / G beta:gamma signalling through PI3Kgamma / adenylate cyclase-activating dopamine receptor signaling pathway / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / nervous system development / GPER1 signaling / signaling receptor activity / G-protein beta-subunit binding / cellular response to prostaglandin E stimulus / heterotrimeric G-protein complex / Inactivation, recovery and regulation of the phototransduction cascade / G alpha (12/13) signalling events / extracellular vesicle / sensory perception of taste / Thrombin signalling through proteinase activated receptors (PARs) / signaling receptor complex adaptor activity / retina development in camera-type eye / GTPase binding / Ca2+ pathway / fibroblast proliferation / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G alpha (i) signalling events / basolateral plasma membrane / G alpha (s) signalling events / phospholipase C-activating G protein-coupled receptor signaling pathway / G alpha (q) signalling events / chemical synaptic transmission / postsynaptic membrane / Ras protein signal transduction / Extra-nuclear estrogen signaling / cell population proliferation / G protein-coupled receptor signaling pathway / lysosomal membrane / GTPase activity / dendrite / synapse / endoplasmic reticulum membrane / protein-containing complex binding / signal transduction / extracellular exosome / membrane / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
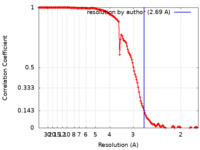
| Method | single particle reconstruction / cryo EM / Resolution: 2.69 Å | |||||||||
 Authors Authors | Zhang S / Fay JF / Roth BL | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Molecular basis for selective activation of DREADD-based chemogenetics. Authors: Shicheng Zhang / Ryan H Gumpper / Xi-Ping Huang / Yongfeng Liu / Brian E Krumm / Can Cao / Jonathan F Fay / Bryan L Roth /  Abstract: Designer receptors exclusively activated by designer drugs (DREADDs) represent a powerful chemogenetic technology for the remote control of neuronal activity and cellular signalling. The muscarinic ...Designer receptors exclusively activated by designer drugs (DREADDs) represent a powerful chemogenetic technology for the remote control of neuronal activity and cellular signalling. The muscarinic receptor-based DREADDs are the most widely used chemogenetic tools in neuroscience research. The G-coupled DREADD (hM3Dq) is used to enhance neuronal activity, whereas the G-coupled DREADD (hM4Di) is utilized to inhibit neuronal activity. Here we report four DREADD-related cryogenic electron microscopy high-resolution structures: a hM3Dq-miniG complex and a hM4Di-miniG complex bound to deschloroclozapine; a hM3Dq-miniG complex bound to clozapine-N-oxide; and a hM3R-miniG complex bound to iperoxo. Complemented with mutagenesis, functional and computational simulation data, our structures reveal key details of the recognition of DREADD chemogenetic actuators and the molecular basis for activation. These findings should accelerate the structure-guided discovery of next-generation chemogenetic tools. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27966.map.gz emd_27966.map.gz | 77.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27966-v30.xml emd-27966-v30.xml emd-27966.xml emd-27966.xml | 27.2 KB 27.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27966_fsc.xml emd_27966_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_27966.png emd_27966.png | 63.1 KB | ||
| Filedesc metadata |  emd-27966.cif.gz emd-27966.cif.gz | 7.3 KB | ||
| Others |  emd_27966_additional_1.map.gz emd_27966_additional_1.map.gz emd_27966_half_map_1.map.gz emd_27966_half_map_1.map.gz emd_27966_half_map_2.map.gz emd_27966_half_map_2.map.gz | 84.9 MB 84.6 MB 84.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27966 http://ftp.pdbj.org/pub/emdb/structures/EMD-27966 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27966 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27966 | HTTPS FTP |
-Related structure data
| Related structure data |  8e9wMC  8e9xC  8e9yC  8e9zC  8ea0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27966.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27966.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of Gq-coupled hM3Dq in complex with DCZ | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Additional Map
| File | emd_27966_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Additional Map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 1
| File | emd_27966_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half Map 2
| File | emd_27966_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Gq-coupled hM3R complex
| Entire | Name: Gq-coupled hM3R complex |
|---|---|
| Components |
|
-Supramolecule #1: Gq-coupled hM3R complex
| Supramolecule | Name: Gq-coupled hM3R complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|
-Supramolecule #2: muscarinic acetylcholine receptor 3, miniGq protein, Guanine nucl...
| Supramolecule | Name: muscarinic acetylcholine receptor 3, miniGq protein, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Single-chain variable fragment scFv16
| Supramolecule | Name: Single-chain variable fragment scFv16 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #5 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Muscarinic acetylcholine receptor M3
| Macromolecule | Name: Muscarinic acetylcholine receptor M3 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.590023 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AGNFSSPDGT TDDPLGGHTV WQVVFIAFLT GILALVTIIG NILVIVSFKV NKQLKTVNNY FLLSLACADL IIGVISMNLF TTYIIMNRW ALGNLACDLW LAIDCVASNA SVMNLLVISF DRYFSITRPL TYRAKRTTKR AGVMIGLAWV ISFVLWAPAI L FWQYFVGK ...String: AGNFSSPDGT TDDPLGGHTV WQVVFIAFLT GILALVTIIG NILVIVSFKV NKQLKTVNNY FLLSLACADL IIGVISMNLF TTYIIMNRW ALGNLACDLW LAIDCVASNA SVMNLLVISF DRYFSITRPL TYRAKRTTKR AGVMIGLAWV ISFVLWAPAI L FWQYFVGK RTVPPGECFI QFLSEPTITF GTAIAGFYMP VTIMTILYWR IYKETEKRTK ELAGLQASGT EAETENFVHP AK RFALKTR SQITKRKRMS LVKEKKAAQT LSAILLAFII TWTPYNIMVL VNTFCDSCIP KTFWNLGYWL CYINSTVNPV CYA LCNKTF RTTFKMLLLC QCDKKKRRKQ QYQQRQSVIF HKRAPEQAL UniProtKB: Muscarinic acetylcholine receptor M3, Muscarinic acetylcholine receptor M3 |
-Macromolecule #2: miniGq
| Macromolecule | Name: miniGq / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.084832 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSTVSAEDK AAAERSKMID KNLREDGEKA RRTLRLLLLG ADNSGKSTIV KQMRILHGGS GGSGGTSGIF ETKFQVDKVN FHMFDVGGQ RDERRKWIQC FNDVTAIIFV VDSSDYNRLQ EALNDFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK I EDYFPEFA ...String: MGSTVSAEDK AAAERSKMID KNLREDGEKA RRTLRLLLLG ADNSGKSTIV KQMRILHGGS GGSGGTSGIF ETKFQVDKVN FHMFDVGGQ RDERRKWIQC FNDVTAIIFV VDSSDYNRLQ EALNDFKSIW NNRWLRTISV ILFLNKQDLL AEKVLAGKSK I EDYFPEFA RYTTPEDATP EPGEDPRVTR AKYFIRKEFV DISTASGDGR HICYPHFTCA VDTENARRIF NDCKDIILQM NL REYNLV |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 37.285734 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SELDQLRQEA EQLKNQIRDA RKACADATLS QITNNIDPVG RIQMRTRRTL RGHLAKIYAM HWGTDSRLLV SASQDGKLII WDSYTTNKV HAIPLRSSWV MTCAYAPSGN YVACGGLDNI CSIYNLKTRE GNVRVSRELA GHTGYLSCCR FLDDNQIVTS S GDTTCALW ...String: SELDQLRQEA EQLKNQIRDA RKACADATLS QITNNIDPVG RIQMRTRRTL RGHLAKIYAM HWGTDSRLLV SASQDGKLII WDSYTTNKV HAIPLRSSWV MTCAYAPSGN YVACGGLDNI CSIYNLKTRE GNVRVSRELA GHTGYLSCCR FLDDNQIVTS S GDTTCALW DIETGQQTTT FTGHTGDVMS LSLAPDTRLF VSGACDASAK LWDVREGMCR QTFTGHESDI NAICFFPNGN AF ATGSDDA TCRLFDLRAD QELMTYSHDN IICGITSVSF SKSGRLLLAG YDDFNCNVWD ALKADRAGVL AGHDNRVSCL GVT DDGMAV ATGSWDSFLK IWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #4: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.861143 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFCAI L UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Macromolecule #5: scFv16
| Macromolecule | Name: scFv16 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 26.679721 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS ...String: DVQLVESGGG LVQPGGSRKL SCSASGFAFS SFGMHWVRQA PEKGLEWVAY ISSGSGTIYY ADTVKGRFTI SRDDPKNTLF LQMTSLRSE DTAMYYCVRS IYYYGSSPFD FWGQGTTLTV SSGGGGSGGG GSGGGGSDIV MTQATSSVPV TPGESVSISC R SSKSLLHS NGNTYLYWFL QRPGQSPQLL IYRMSNLASG VPDRFSGSGS GTAFTLTISR LEAEDVGVYY CMQHLEYPLT FG AGTKLEL KAAA |
-Macromolecule #6: 11-(4-methylpiperazin-1-yl)-5H-dibenzo[b,e][1,4]diazepine
| Macromolecule | Name: 11-(4-methylpiperazin-1-yl)-5H-dibenzo[b,e][1,4]diazepine type: ligand / ID: 6 / Number of copies: 1 / Formula: WEC |
|---|---|
| Molecular weight | Theoretical: 292.378 Da |
| Chemical component information |  ChemComp-WEC: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 3234 / Average electron dose: 46.1 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.3 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)