+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of TRPM3 ion channel in the absence of PIP2 | |||||||||
 Map data Map data | sharpened map with a B-factor of 13.2 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRPM3 / ion channel / PIP2 / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmetal ion transport / monoatomic cation transmembrane transport / monoatomic cation transport / monoatomic cation channel activity / protein tetramerization / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
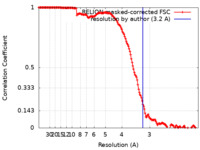
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Zhao C / MacKinnon R | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Neuron / Year: 2023 Journal: Neuron / Year: 2023Title: Structural and functional analyses of a GPCR-inhibited ion channel TRPM3. Authors: Chen Zhao / Roderick MacKinnon /  Abstract: G-protein coupled receptors (GPCRs) govern the physiological response to stimuli by modulating the activity of downstream effectors, including ion channels. TRPM3 is an ion channel inhibited by GPCRs ...G-protein coupled receptors (GPCRs) govern the physiological response to stimuli by modulating the activity of downstream effectors, including ion channels. TRPM3 is an ion channel inhibited by GPCRs through direct interaction with G protein (Gβγ) released upon their activation. This GPCR-TRPM3 signaling pathway contributes to the analgesic effect of morphine. Here, we characterized Gβγ inhibition of TRPM3 using electrophysiology and single particle cryo-electron microscopy (cryo-EM). From electrophysiology, we obtained a half inhibition constant (IC50) of ∼240 nM. Using cryo-EM, we determined structures of mouse TRPM3 expressed in human cells with and without Gβγ and with and without PIP, a lipid required for TRPM3 activity, at resolutions of 2.7-4.7 Å. Gβγ-TRPM3 interfaces vary depending on PIP occupancy; however, in all cases, Gβγ appears loosely attached to TRPM3. The IC50 in electrophysiology experiments raises the possibility that additional unknown factors may stabilize the TRPM3-Gβγ complex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27339.map.gz emd_27339.map.gz | 14.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27339-v30.xml emd-27339-v30.xml emd-27339.xml emd-27339.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27339_fsc.xml emd_27339_fsc.xml | 15.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_27339.png emd_27339.png | 173.4 KB | ||
| Filedesc metadata |  emd-27339.cif.gz emd-27339.cif.gz | 6.4 KB | ||
| Others |  emd_27339_half_map_1.map.gz emd_27339_half_map_1.map.gz emd_27339_half_map_2.map.gz emd_27339_half_map_2.map.gz | 254.4 MB 254.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27339 http://ftp.pdbj.org/pub/emdb/structures/EMD-27339 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27339 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27339 | HTTPS FTP |
-Validation report
| Summary document |  emd_27339_validation.pdf.gz emd_27339_validation.pdf.gz | 827.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27339_full_validation.pdf.gz emd_27339_full_validation.pdf.gz | 826.9 KB | Display | |
| Data in XML |  emd_27339_validation.xml.gz emd_27339_validation.xml.gz | 23.1 KB | Display | |
| Data in CIF |  emd_27339_validation.cif.gz emd_27339_validation.cif.gz | 30.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27339 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27339 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27339 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27339 | HTTPS FTP |
-Related structure data
| Related structure data |  8ddrMC  8ddqC  8ddsC  8ddtC  8dduC  8ddvC  8ddwC  8ddxC  8ed7C  8ed8C  8ed9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27339.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27339.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map with a B-factor of 13.2 | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.046 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map 1
| File | emd_27339_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
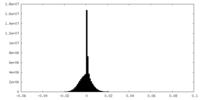
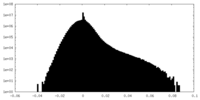
| Density Histograms |
-Half map: half map 2
| File | emd_27339_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
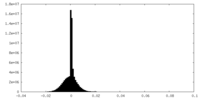
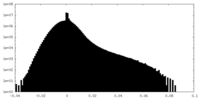
| Density Histograms |
- Sample components
Sample components
-Entire : TRPM3
| Entire | Name: TRPM3 |
|---|---|
| Components |
|
-Supramolecule #1: TRPM3
| Supramolecule | Name: TRPM3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transient receptor potential cation channel, subfamily M, member 3
| Macromolecule | Name: Transient receptor potential cation channel, subfamily M, member 3 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 154.649328 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GKKWRDAGEL ERGCSDREDS AESRRRSRSA SRGRFAESWK RLSSKQGSTK RSGLPAQQTP AQKSWIERAF YKRECVHIIP STKDPHRCC CGRLIGQHVG LTPSISVLQN EKNESRLSRN DIQSEKWSIS KHTQLSPTDA FGTIEFQGGG HSNKAMYVRV S FDTKPDLL ...String: GKKWRDAGEL ERGCSDREDS AESRRRSRSA SRGRFAESWK RLSSKQGSTK RSGLPAQQTP AQKSWIERAF YKRECVHIIP STKDPHRCC CGRLIGQHVG LTPSISVLQN EKNESRLSRN DIQSEKWSIS KHTQLSPTDA FGTIEFQGGG HSNKAMYVRV S FDTKPDLL LHLMTKEWQL ELPKLLISVH GGLQNFELQP KLKQVFGKGL IKAAMTTGAW IFTGGVNTGV IRHVGDALKD HA SKSRGKI CTIGIAPWGI VENQEDLIGR DVVRPYQTMS NPMSKLTVLN SMHSHFILAD NGTTGKYGAE VKLRRQLEKH ISL QKINTR IGQGVPVVAL IVEGGPNVIS IVLEYLRDTP PVPVVVCDGS GRASDILAFG HKYSEEGGLI NESLRDQLLV TIQK TFTYT RTQAQHLFII LMECMKKKEL ITVFRMGSEG HQDIDLAILT ALLKGANASA PDQLSLALAW NRVDIARSQI FIYGQ QWPV GSLEQAMLDA LVLDRVDFVK LLIENGVSMH RFLTISRLEE LYNTRHGPSN TLYHLVRDVK KGNLPPDYRI SLIDIG LVI EYLMGGAYRC NYTRKRFRTL YHNLFGPKRP KALKLLGMED DIPLRRGRKT TKKREEEVDI DLDDPEINHF PFPFHEL MV WAVLMKRQKM ALFFWQHGEE AMAKALVACK LCKAMAHEAS ENDMVDDISQ ELNHNSRDFG QLAVELLDQS YKQDEQLA M KLLTYELKNW SNATCLQLAV AAKHRDFIAH TCSQMLLTDM WMGRLRMRKN SGLKVILGIL LPPSILSLEF KNKDDMPYM TQAQEIHLQE KEPEEPEKPT KEKDEEDMEL TAMLGRSNGE SSRKKDEEEV QSRHRLIPVG RKIYEFYNAP IVKFWFYTLA YIGYLMLFN YIVLVKMERW PSTQEWIVIS YIFTLGIEKM REILMSEPGK LLQKVKVWLQ EYWNVTDLIA ILLFSVGMIL R LQDQPFRS DGRVIYCVNI IYWYIRLLDI FGVNKYLGPY VMMIGKMMID MMYFVIIMLV VLMSFGVARQ AILFPNEEPS WK LAKNIFY MPYWMIYGEV FADQIDPPCG QNETREDGKT IQLPPCKTGA WIVPAIMACY LLVANILLVN LLIAVFNNTF FEV KSISNQ VWKFQRYQLI MTFHERPVLP PPLIIFSHMT MIFQHVCCRW RKHESDQDER DYGLKLFITD DELKKVHDFE EQCI EEYFR EKDDRFNSSN DERIRVTSER VENMSMRLEE VNEREHSMKA SLQTVDIRLA QLEDLIGRMA TALERLTGLE RAESN KIRS RTSSDCTDAA YIVRQSSFNS QEGNTFKLQE SIDPAGEETI SPTSPTLMPR MRSHSFYSV UniProtKB: Transient receptor potential cation channel, subfamily M, member 3 |
-Macromolecule #2: Unidentified segment at the N-terminus of TRPM3
| Macromolecule | Name: Unidentified segment at the N-terminus of TRPM3 / type: protein_or_peptide / ID: 2 Details: a segment at the N-terminus of TRPM3 whose sequence cannot be identified from the cryo-EM density Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.464797 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) |
-Macromolecule #3: 1,2-DIACYL-GLYCEROL-3-SN-PHOSPHATE
| Macromolecule | Name: 1,2-DIACYL-GLYCEROL-3-SN-PHOSPHATE / type: ligand / ID: 3 / Number of copies: 4 / Formula: 3PH |
|---|---|
| Molecular weight | Theoretical: 704.998 Da |
| Chemical component information |  ChemComp-3PH: |
-Macromolecule #4: (3beta,14beta,17beta,25R)-3-[4-methoxy-3-(methoxymethyl)butoxy]sp...
| Macromolecule | Name: (3beta,14beta,17beta,25R)-3-[4-methoxy-3-(methoxymethyl)butoxy]spirost-5-en type: ligand / ID: 4 / Number of copies: 4 / Formula: 9Z9 |
|---|---|
| Molecular weight | Theoretical: 544.805 Da |
| Chemical component information |  ChemComp-9Z9: |
-Macromolecule #5: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 5 / Number of copies: 2 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 IS (4k x 4k) / Average electron dose: 73.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)