[English] 日本語
 Yorodumi
Yorodumi- EMDB-26346: Cryo-EM structure of human CST bound to DNA polymerase alpha-prim... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human CST bound to DNA polymerase alpha-primase in a recruitment state | |||||||||
 Map data Map data | Human CST-Pol alpha/Primase bound to ssDNA in a recruitment state. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Fill-in / Telomere / Replication / DNA BINDING PROTEIN / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationCST complex / ribonucleotide binding / DNA primase AEP / DNA replication initiation / telomerase inhibitor activity / DNA/RNA hybrid binding / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / telomere maintenance via telomere lengthening / Telomere C-strand synthesis initiation / alpha DNA polymerase:primase complex ...CST complex / ribonucleotide binding / DNA primase AEP / DNA replication initiation / telomerase inhibitor activity / DNA/RNA hybrid binding / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / telomere maintenance via telomere lengthening / Telomere C-strand synthesis initiation / alpha DNA polymerase:primase complex / regulation of type I interferon production / Polymerase switching / Processive synthesis on the lagging strand / single-stranded telomeric DNA binding / Removal of the Flap Intermediate / lagging strand elongation / intermediate filament cytoskeleton / G-rich strand telomeric DNA binding / telomere capping / DNA replication, synthesis of primer / mitotic DNA replication initiation / Polymerase switching on the C-strand of the telomere / bone marrow development / DNA strand elongation involved in DNA replication / hematopoietic stem cell proliferation / G1/S-Specific Transcription / leading strand elongation / telomeric repeat DNA binding / DNA synthesis involved in DNA repair / negative regulation of telomere maintenance via telomerase / DNA replication origin binding / replicative senescence / Activation of the pre-replicative complex / DNA replication initiation / spleen development / regulation of G2/M transition of mitotic cell cycle / telomere maintenance / positive regulation of DNA replication / thymus development / Defective pyroptosis / double-strand break repair via nonhomologous end joining / positive regulation of fibroblast proliferation / multicellular organism growth / fibrillar center / nuclear matrix / protein import into nucleus / DNA-directed RNA polymerase activity / nuclear envelope / single-stranded DNA binding / 4 iron, 4 sulfur cluster binding / DNA-directed DNA polymerase / DNA-directed DNA polymerase activity / chromosome, telomeric region / DNA replication / ciliary basal body / nucleotide binding / DNA repair / chromatin binding / DNA damage response / protein kinase binding / chromatin / nucleolus / magnesium ion binding / DNA binding / zinc ion binding / nucleoplasm / membrane / metal ion binding / nucleus / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
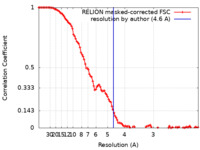
| Method | single particle reconstruction / cryo EM / Resolution: 4.6 Å | |||||||||
 Authors Authors | Cai SW / Zinder JC / Svetlov V / Bush MW / Nudler E / Walz T / de Lange T | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: Cryo-EM structure of the human CST-Polα/primase complex in a recruitment state. Authors: Sarah W Cai / John C Zinder / Vladimir Svetlov / Martin W Bush / Evgeny Nudler / Thomas Walz / Titia de Lange /  Abstract: The CST-Polα/primase complex is essential for telomere maintenance and functions to counteract resection at double-strand breaks. We report a 4.6-Å resolution cryo-EM structure of human CST- ...The CST-Polα/primase complex is essential for telomere maintenance and functions to counteract resection at double-strand breaks. We report a 4.6-Å resolution cryo-EM structure of human CST-Polα/primase, captured prior to catalysis in a recruitment state stabilized by chemical cross-linking. Our structure reveals an evolutionarily conserved interaction between the C-terminal domain of the catalytic POLA1 subunit and an N-terminal expansion in metazoan CTC1. Cross-linking mass spectrometry and negative-stain EM analysis provide insight into CST binding by the flexible POLA1 N-terminus. Finally, Coats plus syndrome disease mutations previously characterized to disrupt formation of the CST-Polα/primase complex map to protein-protein interfaces observed in the recruitment state. Together, our results shed light on the architecture and stoichiometry of the metazoan fill-in machinery. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26346.map.gz emd_26346.map.gz | 95.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26346-v30.xml emd-26346-v30.xml emd-26346.xml emd-26346.xml | 27.8 KB 27.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_26346_fsc.xml emd_26346_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_26346.png emd_26346.png | 96.1 KB | ||
| Filedesc metadata |  emd-26346.cif.gz emd-26346.cif.gz | 9.1 KB | ||
| Others |  emd_26346_half_map_1.map.gz emd_26346_half_map_1.map.gz emd_26346_half_map_2.map.gz emd_26346_half_map_2.map.gz | 80.9 MB 80.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26346 http://ftp.pdbj.org/pub/emdb/structures/EMD-26346 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26346 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26346 | HTTPS FTP |
-Related structure data
| Related structure data |  7u5cMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
| EM raw data |  EMPIAR-11131 (Title: Single particle cryo-EM of the human CST•Polα/Primase (POLA1ΔN) complex in a recruitment state EMPIAR-11131 (Title: Single particle cryo-EM of the human CST•Polα/Primase (POLA1ΔN) complex in a recruitment stateData size: 869.0 Data #1: Motion corrected micrographs of CST-Pola/Primase deltaN complex in recruitment state [micrographs - single frame]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26346.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26346.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human CST-Pol alpha/Primase bound to ssDNA in a recruitment state. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_26346_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_26346_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Human CST bound to DNA polymerase alpha(POLA1 delta N)-primase in...
+Supramolecule #1: Human CST bound to DNA polymerase alpha(POLA1 delta N)-primase in...
+Macromolecule #1: DNA primase small subunit
+Macromolecule #2: DNA primase large subunit
+Macromolecule #3: DNA polymerase alpha catalytic subunit
+Macromolecule #4: DNA polymerase alpha subunit B
+Macromolecule #5: CST complex subunit CTC1
+Macromolecule #6: CST complex subunit STN1
+Macromolecule #7: CST complex subunit TEN1
+Macromolecule #8: canonical telomeric DNA sequence
+Macromolecule #9: ZINC ION
+Macromolecule #10: IRON/SULFUR CLUSTER
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.075 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Support film - Material: GRAPHENE OXIDE / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV |
| Details | The sample was cross-linked with glutaraldehyde (GraFix) |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)

































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)