+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-25780 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
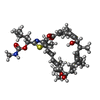
| タイトル | Cryo-EM structure of archazolid A bound to yeast VO V-ATPase | |||||||||
 マップデータ マップデータ | Yeast VO motor with Archazolid A | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | inhibitor / complex / proton pump / ATPase / MEMBRANE PROTEIN | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報cell wall mannoprotein biosynthetic process / ATPase-coupled ion transmembrane transporter activity / protein localization to vacuolar membrane / cellular response to alkaline pH / Insulin receptor recycling / Transferrin endocytosis and recycling / polyphosphate metabolic process / ROS and RNS production in phagocytes / Amino acids regulate mTORC1 / Golgi lumen acidification ...cell wall mannoprotein biosynthetic process / ATPase-coupled ion transmembrane transporter activity / protein localization to vacuolar membrane / cellular response to alkaline pH / Insulin receptor recycling / Transferrin endocytosis and recycling / polyphosphate metabolic process / ROS and RNS production in phagocytes / Amino acids regulate mTORC1 / Golgi lumen acidification / P-type proton-exporting transporter activity / vacuolar transport / vacuolar proton-transporting V-type ATPase, V0 domain / endosomal lumen acidification / vacuole organization / protein targeting to vacuole / proton-transporting V-type ATPase complex / fungal-type vacuole / vacuolar proton-transporting V-type ATPase complex / cellular hyperosmotic response / vacuolar acidification / fungal-type vacuole membrane / phosphatidylinositol-3,5-bisphosphate binding / proton transmembrane transporter activity / proton-transporting ATPase activity, rotational mechanism / intracellular copper ion homeostasis / RNA endonuclease activity / Neutrophil degranulation / proton transmembrane transport / cell periphery / transmembrane transport / endocytosis / ATPase binding / protein-containing complex assembly / intracellular iron ion homeostasis / membrane raft / Golgi membrane / endoplasmic reticulum membrane / membrane 類似検索 - 分子機能 | |||||||||
| 生物種 |  | |||||||||
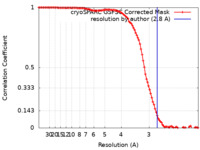
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.8 Å | |||||||||
 データ登録者 データ登録者 | Keon KA / Rubinstein JL | |||||||||
| 資金援助 |  カナダ, 1件 カナダ, 1件
| |||||||||
 引用 引用 |  ジャーナル: ACS Chem Biol / 年: 2022 ジャーナル: ACS Chem Biol / 年: 2022タイトル: Cryo-EM of the Yeast V Complex Reveals Distinct Binding Sites for Macrolide V-ATPase Inhibitors. 著者: Kristine A Keon / Samir Benlekbir / Susanne H Kirsch / Rolf Müller / John L Rubinstein /   要旨: Vacuolar-type adenosine triphosphatases (V-ATPases) are proton pumps found in almost all eukaryotic cells. These enzymes consist of a soluble catalytic V region that hydrolyzes ATP and a membrane- ...Vacuolar-type adenosine triphosphatases (V-ATPases) are proton pumps found in almost all eukaryotic cells. These enzymes consist of a soluble catalytic V region that hydrolyzes ATP and a membrane-embedded V region responsible for proton translocation. V-ATPase activity leads to acidification of endosomes, phagosomes, lysosomes, secretory vesicles, and the trans-Golgi network, with extracellular acidification occurring in some specialized cells. Small-molecule inhibitors of V-ATPase have played a crucial role in elucidating numerous aspects of cell biology by blocking acidification of intracellular compartments, while therapeutic use of V-ATPase inhibitors has been proposed for the treatment of cancer, osteoporosis, and some infections. Here, we determine structures of the isolated V complex from bound to two well-known macrolide inhibitors: bafilomycin A1 and archazolid A. The structures reveal different binding sites for the inhibitors on the surface of the proton-carrying c ring, with only a small amount of overlap between the two sites. Binding of both inhibitors is mediated primarily through van der Waals interactions in shallow pockets and suggests that the inhibitors block rotation of the ring. Together, these structures indicate the existence of a large chemical space available for V-ATPase inhibitors that block acidification by binding the c ring. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_25780.map.gz emd_25780.map.gz | 97.1 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-25780-v30.xml emd-25780-v30.xml emd-25780.xml emd-25780.xml | 20.5 KB 20.5 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_25780_fsc.xml emd_25780_fsc.xml | 11.2 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_25780.png emd_25780.png | 79.3 KB | ||
| Filedesc metadata |  emd-25780.cif.gz emd-25780.cif.gz | 7.3 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25780 http://ftp.pdbj.org/pub/emdb/structures/EMD-25780 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25780 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25780 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_25780.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_25780.map.gz / 形式: CCP4 / 大きさ: 103 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Yeast VO motor with Archazolid A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : Cryo-EM structure of archazolid A bound to yeast VO V-ATPase
+超分子 #1: Cryo-EM structure of archazolid A bound to yeast VO V-ATPase
+分子 #1: V-type proton ATPase subunit c'
+分子 #2: V-type proton ATPase subunit c''
+分子 #3: V0 assembly protein 1
+分子 #4: V-type proton ATPase subunit e
+分子 #5: V-type proton ATPase subunit c
+分子 #6: Yeast V-ATPase subunit f
+分子 #7: V-type proton ATPase subunit d
+分子 #8: V-type proton ATPase subunit a, vacuolar isoform
+分子 #9: Archazolid A
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON IV (4k x 4k) 平均電子線量: 42.0 e/Å2 / 詳細: Prototype Falcon 4i camera |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD 最大 デフォーカス(公称値): 2.5500000000000003 µm 最小 デフォーカス(公称値): 0.37 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)