[English] 日本語
 Yorodumi
Yorodumi- EMDB-25413: Structure of the NADH-bound human COQ7:COQ9 complex by single-par... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the NADH-bound human COQ7:COQ9 complex by single-particle electron cryo-microscopy | |||||||||
 Map data Map data | PostProcess Map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | hydroxylase / complex / lipid / enzyme / NADH / coenzymeQ / isoprene / OPP / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology information3-demethoxyubiquinol 3-hydroxylase / 3-demethoxyubiquinol 3-hydroxylase activity / extrinsic component of mitochondrial inner membrane / Ubiquinol biosynthesis / ubiquinone biosynthesis complex / ubiquinone biosynthetic process / oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen / regulation of reactive oxygen species metabolic process / mitochondrial electron transport, NADH to ubiquinone / determination of adult lifespan ...3-demethoxyubiquinol 3-hydroxylase / 3-demethoxyubiquinol 3-hydroxylase activity / extrinsic component of mitochondrial inner membrane / Ubiquinol biosynthesis / ubiquinone biosynthesis complex / ubiquinone biosynthetic process / oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NAD(P)H as one donor, and incorporation of one atom of oxygen / regulation of reactive oxygen species metabolic process / mitochondrial electron transport, NADH to ubiquinone / determination of adult lifespan / chromosome / regulation of gene expression / mitochondrial inner membrane / lipid binding / chromatin binding / negative regulation of transcription by RNA polymerase II / protein homodimerization activity / positive regulation of transcription by RNA polymerase II / mitochondrion / nucleus / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
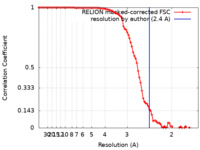
| Method | single particle reconstruction / cryo EM / Resolution: 2.4 Å | |||||||||
 Authors Authors | Aydin H / Frost A | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2022 Journal: Mol Cell / Year: 2022Title: Structure and functionality of a multimeric human COQ7:COQ9 complex. Authors: Mateusz Manicki / Halil Aydin / Luciano A Abriata / Katherine A Overmyer / Rachel M Guerra / Joshua J Coon / Matteo Dal Peraro / Adam Frost / David J Pagliarini /   Abstract: Coenzyme Q (CoQ) is a redox-active lipid essential for core metabolic pathways and antioxidant defense. CoQ is synthesized upon the mitochondrial inner membrane by an ill-defined "complex Q" ...Coenzyme Q (CoQ) is a redox-active lipid essential for core metabolic pathways and antioxidant defense. CoQ is synthesized upon the mitochondrial inner membrane by an ill-defined "complex Q" metabolon. Here, we present structure-function analyses of a lipid-, substrate-, and NADH-bound complex comprising two complex Q subunits: the hydroxylase COQ7 and the lipid-binding protein COQ9. We reveal that COQ7 adopts a ferritin-like fold with a hydrophobic channel whose substrate-binding capacity is enhanced by COQ9. Using molecular dynamics, we further show that two COQ7:COQ9 heterodimers form a curved tetramer that deforms the membrane, potentially opening a pathway for the CoQ intermediates to translocate from the bilayer to the proteins' lipid-binding sites. Two such tetramers assemble into a soluble octamer with a pseudo-bilayer of lipids captured within. Together, these observations indicate that COQ7 and COQ9 cooperate to access hydrophobic precursors within the membrane and coordinate subsequent synthesis steps toward producing CoQ. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25413.map.gz emd_25413.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25413-v30.xml emd-25413-v30.xml emd-25413.xml emd-25413.xml | 18.7 KB 18.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25413_fsc.xml emd_25413_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_25413.png emd_25413.png | 19.8 KB | ||
| Masks |  emd_25413_msk_1.map emd_25413_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-25413.cif.gz emd-25413.cif.gz | 6 KB | ||
| Others |  emd_25413_additional_1.map.gz emd_25413_additional_1.map.gz emd_25413_half_map_1.map.gz emd_25413_half_map_1.map.gz emd_25413_half_map_2.map.gz emd_25413_half_map_2.map.gz | 47.9 MB 48.2 MB 48.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25413 http://ftp.pdbj.org/pub/emdb/structures/EMD-25413 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25413 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25413 | HTTPS FTP |
-Validation report
| Summary document |  emd_25413_validation.pdf.gz emd_25413_validation.pdf.gz | 957 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25413_full_validation.pdf.gz emd_25413_full_validation.pdf.gz | 956.6 KB | Display | |
| Data in XML |  emd_25413_validation.xml.gz emd_25413_validation.xml.gz | 16.2 KB | Display | |
| Data in CIF |  emd_25413_validation.cif.gz emd_25413_validation.cif.gz | 21.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25413 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25413 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25413 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25413 | HTTPS FTP |
-Related structure data
| Related structure data |  7sssMC  7sspC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25413.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25413.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | PostProcess Map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.833 Å | ||||||||||||||||||||||||||||||||||||
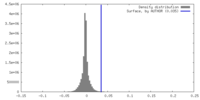
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25413_msk_1.map emd_25413_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
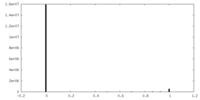
| Density Histograms |
-Additional map: Refine3D map
| File | emd_25413_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Refine3D map | ||||||||||||
| Projections & Slices |
| ||||||||||||
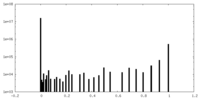
| Density Histograms |
-Half map: Half Map 1
| File | emd_25413_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
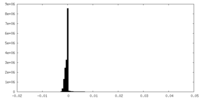
| Density Histograms |
-Half map: Half Map 2
| File | emd_25413_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human COQ7:COQ9 Complex
| Entire | Name: Human COQ7:COQ9 Complex |
|---|---|
| Components |
|
-Supramolecule #1: Human COQ7:COQ9 Complex
| Supramolecule | Name: Human COQ7:COQ9 Complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Ubiquinone biosynthesis protein COQ9, mitochondrial
| Macromolecule | Name: Ubiquinone biosynthesis protein COQ9, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 35.549945 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAAAVSGAL GRAGWRLLQL RCLPVARCRQ ALVPRAFHAS AVGLRSSDEQ KQQPPNSFSQ QHSETQGAEK PDPESSHSPP RYTDQGGEE EEDYESEEQL QHRILTAALE FVPAHGWTAE AIAEGAQSLG LSSAAASMFG KDGSELILHF VTQCNTRLTR V LEEEQKLV ...String: MAAAAVSGAL GRAGWRLLQL RCLPVARCRQ ALVPRAFHAS AVGLRSSDEQ KQQPPNSFSQ QHSETQGAEK PDPESSHSPP RYTDQGGEE EEDYESEEQL QHRILTAALE FVPAHGWTAE AIAEGAQSLG LSSAAASMFG KDGSELILHF VTQCNTRLTR V LEEEQKLV QLGQAEKRKT DQFLRDAVET RLRMLIPYIE HWPRALSILM LPHNIPSSLS LLTSMVDDMW HYAGDQSTDF NW YTRRAML AAIYNTTELV MMQDSSPDFE DTWRFLENRV NDAMNMGHTA KQVKSTGEAL VQGLMGAAVT LKNLTGLNQR R UniProtKB: Ubiquinone biosynthesis protein COQ9, mitochondrial |
-Macromolecule #2: 5-demethoxyubiquinone hydroxylase, mitochondrial
| Macromolecule | Name: 5-demethoxyubiquinone hydroxylase, mitochondrial / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO / EC number: 3-demethoxyubiquinol 3-hydroxylase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.313064 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSCAGAAAAP RLWRLRPGAR RSLSAYGRRT SVRFRSSGMT LDNISRAAVD RIIRVDHAGE YGANRIYAGQ MAVLGRTSVG PVIQKMWDQ EKDHLKKFNE LMVTFRVRPT VLMPLWNVLG FALGAGTALL GKEGAMACTV AVEESIAHHY NNQIRTLMEE D PEKYEELL ...String: MSCAGAAAAP RLWRLRPGAR RSLSAYGRRT SVRFRSSGMT LDNISRAAVD RIIRVDHAGE YGANRIYAGQ MAVLGRTSVG PVIQKMWDQ EKDHLKKFNE LMVTFRVRPT VLMPLWNVLG FALGAGTALL GKEGAMACTV AVEESIAHHY NNQIRTLMEE D PEKYEELL QLIKKFRDEE LEHHDIGLDH DAELAPAYAV LKSIIQAGCR VAIYLSERL UniProtKB: 5-demethoxyubiquinone hydroxylase, mitochondrial |
-Macromolecule #3: (1S)-2-{[(2-AMINOETHOXY)(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY...
| Macromolecule | Name: (1S)-2-{[(2-AMINOETHOXY)(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL STEARATE type: ligand / ID: 3 / Number of copies: 32 / Formula: PEV |
|---|---|
| Molecular weight | Theoretical: 720.012 Da |
| Chemical component information |  ChemComp-PEV: |
-Macromolecule #4: 2-[(2E,6E,10E,14E,18E,22E,26E)-3,7,11,15,19,23,27,31-OCTAMETHYLDO...
| Macromolecule | Name: 2-[(2E,6E,10E,14E,18E,22E,26E)-3,7,11,15,19,23,27,31-OCTAMETHYLDOTRIACONTA-2,6,10,14,18,22,26,30-OCTAENYL]PHENOL type: ligand / ID: 4 / Number of copies: 4 / Formula: 8PP |
|---|---|
| Molecular weight | Theoretical: 639.047 Da |
| Chemical component information |  ChemComp-8PP: |
-Macromolecule #5: NICOTINAMIDE-ADENINE-DINUCLEOTIDE
| Macromolecule | Name: NICOTINAMIDE-ADENINE-DINUCLEOTIDE / type: ligand / ID: 5 / Number of copies: 2 / Formula: NAD |
|---|---|
| Molecular weight | Theoretical: 663.425 Da |
| Chemical component information |  ChemComp-NAD: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 170 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 65.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.2 µm / Nominal defocus min: 0.3 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)