[English] 日本語
 Yorodumi
Yorodumi- EMDB-22295: Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22295 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV-1 Env BG505 DS-SOSIP.664 | |||||||||
 Map data Map data | Sharpened Map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | SHIV / V1V2 / V2-Apex / SOSIP / Vaccine / PGT145 / IMMUNE SYSTEM / IMMUNE SYSTEM-Viral Protein complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell ...symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / membrane / identical protein binding Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Gorman J / Kwong PD | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Recapitulation of HIV-1 Env-antibody coevolution in macaques leading to neutralization breadth. Authors: Ryan S Roark / Hui Li / Wilton B Williams / Hema Chug / Rosemarie D Mason / Jason Gorman / Shuyi Wang / Fang-Hua Lee / Juliette Rando / Mattia Bonsignori / Kwan-Ki Hwang / Kevin O Saunders / ...Authors: Ryan S Roark / Hui Li / Wilton B Williams / Hema Chug / Rosemarie D Mason / Jason Gorman / Shuyi Wang / Fang-Hua Lee / Juliette Rando / Mattia Bonsignori / Kwan-Ki Hwang / Kevin O Saunders / Kevin Wiehe / M Anthony Moody / Peter T Hraber / Kshitij Wagh / Elena E Giorgi / Ronnie M Russell / Frederic Bibollet-Ruche / Weimin Liu / Jesse Connell / Andrew G Smith / Julia DeVoto / Alexander I Murphy / Jessica Smith / Wenge Ding / Chengyan Zhao / Neha Chohan / Maho Okumura / Christina Rosario / Yu Ding / Emily Lindemuth / Anya M Bauer / Katharine J Bar / David Ambrozak / Cara W Chao / Gwo-Yu Chuang / Hui Geng / Bob C Lin / Mark K Louder / Richard Nguyen / Baoshan Zhang / Mark G Lewis / Donald D Raymond / Nicole A Doria-Rose / Chaim A Schramm / Daniel C Douek / Mario Roederer / Thomas B Kepler / Garnett Kelsoe / John R Mascola / Peter D Kwong / Bette T Korber / Stephen C Harrison / Barton F Haynes / Beatrice H Hahn / George M Shaw /  Abstract: Neutralizing antibodies elicited by HIV-1 coevolve with viral envelope proteins (Env) in distinctive patterns, in some cases acquiring substantial breadth. We report that primary HIV-1 envelope ...Neutralizing antibodies elicited by HIV-1 coevolve with viral envelope proteins (Env) in distinctive patterns, in some cases acquiring substantial breadth. We report that primary HIV-1 envelope proteins-when expressed by simian-human immunodeficiency viruses in rhesus macaques-elicited patterns of Env-antibody coevolution very similar to those in humans, including conserved immunogenetic, structural, and chemical solutions to epitope recognition and precise Env-amino acid substitutions, insertions, and deletions leading to virus persistence. The structure of one rhesus antibody, capable of neutralizing 49% of a 208-strain panel, revealed a V2 apex mode of recognition like that of human broadly neutralizing antibodies (bNAbs) PGT145 and PCT64-35S. Another rhesus antibody bound the CD4 binding site by CD4 mimicry, mirroring human bNAbs 8ANC131, CH235, and VRC01. Virus-antibody coevolution in macaques can thus recapitulate developmental features of human bNAbs, thereby guiding HIV-1 immunogen design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22295.map.gz emd_22295.map.gz | 85.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22295-v30.xml emd-22295-v30.xml emd-22295.xml emd-22295.xml | 27.9 KB 27.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_22295.png emd_22295.png | 131.7 KB | ||
| Masks |  emd_22295_msk_1.map emd_22295_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-22295.cif.gz emd-22295.cif.gz | 7.5 KB | ||
| Others |  emd_22295_additional_1.map.gz emd_22295_additional_1.map.gz emd_22295_additional_2.map.gz emd_22295_additional_2.map.gz emd_22295_half_map_1.map.gz emd_22295_half_map_1.map.gz emd_22295_half_map_2.map.gz emd_22295_half_map_2.map.gz | 24.5 MB 8.5 MB 84.5 MB 84.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22295 http://ftp.pdbj.org/pub/emdb/structures/EMD-22295 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22295 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22295 | HTTPS FTP |
-Related structure data
| Related structure data |  6xrtMC  6xcjC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22295.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22295.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened Map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
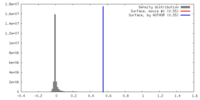
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_22295_msk_1.map emd_22295_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
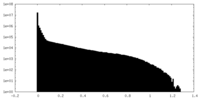
| Density Histograms |
-Additional map: Unsharpened Map
| File | emd_22295_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened Map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Density modified map
| File | emd_22295_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Density modified map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_22295_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_22295_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV...
| Entire | Name: Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV-1 Env BG505 DS-SOSIP |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV...
| Supramolecule | Name: Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV-1 Env BG505 DS-SOSIP type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #2: Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV...
| Supramolecule | Name: Cryo-EM structure of SHIV-elicited RHA1.V2.01 in complex with HIV-1 Env BG505 DS-SOSIP.664 type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Macromolecule #1: Envelope glycoprotein gp160
| Macromolecule | Name: Envelope glycoprotein gp160 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 53.300348 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHTDIIS LWDQSLKPC VKLTPLCVTL QCTNVTNNIT DDMRGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT ...String: AENLWVTVYY GVPVWKDAET TLFCASDAKA YETEKHNVWA THACVPTDPN PQEIHLENVT EEFNMWKNNM VEQMHTDIIS LWDQSLKPC VKLTPLCVTL QCTNVTNNIT DDMRGELKNC SFNMTTELRD KKQKVYSLFY RLDVVQINEN QGNRSNNSNK E YRLINCNT SACTQACPKV SFEPIPIHYC APAGFAILKC KDKKFNGTGP CPSVSTVQCT HGIKPVVSTQ LLLNGSLAEE EV MIRSENI TNNAKNILVQ FNTPVQINCT RPNNNTRKSI RIGPGQAFYA TGDIIGDIRQ AHCNVSKATW NETLGKVVKQ LRK HFGNNT IIRFANSSGG DLEVTTHSFN CGGEFFYCNT SGLFNSTWIS NTSVQGSNST GSNDSITLPC RIKQIINMWQ RIGQ CMYAP PIQGVIRCVS NITGLILTRD GGSTNSTTET FRPGGGDMRD NWRSELYKYK VVKIEPLGVA PTRCKRRVVG R UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #2: HIV-1 Envelope Glycoprotein BG505 SOSIP.664 gp41
| Macromolecule | Name: HIV-1 Envelope Glycoprotein BG505 SOSIP.664 gp41 / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 17.146482 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AVGIGAVFLG FLGAAGSTMG AASMTLTVQA RNLLSGIVQQ QSNLLRAPEA QQHLLKLTVW GIKQLQARVL AVERYLRDQQ LLGIWGCSG KLICCTNVPW NSSWSNRNLS EIWDNMTWLQ WDKEISNYTQ IIYGLLEESQ NQQEKNEQDL LALD UniProtKB: Envelope glycoprotein gp160 |
-Macromolecule #3: VRC01.23 Heavy Chain
| Macromolecule | Name: VRC01.23 Heavy Chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 14.70429 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLRESGPG LVKPSETLVL TCAVSGGGDS FGFHYWNWIR QPPGKGLEWI GHIGGSSGST DFNPSLKSRV TISMDSSRNQ FSLRLKSVT AADTAVYFCA RKGEDFYEDD (TYS)GQYFTAGWF FDLWGPGTPI IISS |
-Macromolecule #4: VRC01.23 Light Chain
| Macromolecule | Name: VRC01.23 Light Chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 11.312331 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QFVLTQPPSV SGAPGQTVTI SCTGRSSNFG GHYVQWYQQL PGTAPRLVIF ENDRRPSGVS DRFSGSQSGA SASLTITGLQ SEDEADYYC QCYDSSVLFG RGTRLT |
-Macromolecule #10: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 10 / Number of copies: 40 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.0 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Formula: PBS |
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 1945 / Average exposure time: 10.0 sec. / Average electron dose: 71.06 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)