[English] 日本語
 Yorodumi
Yorodumi- EMDB-22242: Full-length Hsc82 in complex with Aha1 in the presence of AMP-PNP -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22242 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Full-length Hsc82 in complex with Aha1 in the presence of AMP-PNP | ||||||||||||||||||
 Map data Map data | Hsc82 in complex with Aha1 in the presence of AMP-PNP | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Co-chaperone / activator / CHAPERONE | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationTetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation / eNOS activation / Extra-nuclear estrogen signaling / HSP90 chaperone cycle for steroid hormone receptors (SHR) in the presence of ligand / VEGFR2 mediated vascular permeability / HSF1-dependent transactivation / HSF1 activation / response to oxygen levels / box C/D snoRNP assembly / ATPase activator activity ...Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation / eNOS activation / Extra-nuclear estrogen signaling / HSP90 chaperone cycle for steroid hormone receptors (SHR) in the presence of ligand / VEGFR2 mediated vascular permeability / HSF1-dependent transactivation / HSF1 activation / response to oxygen levels / box C/D snoRNP assembly / ATPase activator activity / proteasome assembly / Neutrophil degranulation / telomere maintenance / ATP-dependent protein folding chaperone / protein import into nucleus / : / cellular response to heat / protein-folding chaperone binding / protein folding / protein stabilization / perinuclear region of cytoplasm / ATP hydrolysis activity / protein-containing complex / mitochondrion / ATP binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
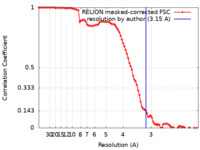
| Method | single particle reconstruction / cryo EM / Resolution: 3.15 Å | ||||||||||||||||||
 Authors Authors | Liu YX / Sun M | ||||||||||||||||||
| Funding support |  United States, 5 items United States, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structures reveal a multistep mechanism of Hsp90 activation by co-chaperone Aha1 Authors: Liu YX / Sun M / Myasnikov AG / Elnatan D / Agard DA | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22242.map.gz emd_22242.map.gz | 59.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22242-v30.xml emd-22242-v30.xml emd-22242.xml emd-22242.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22242_fsc.xml emd_22242_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_22242.png emd_22242.png | 79.3 KB | ||
| Filedesc metadata |  emd-22242.cif.gz emd-22242.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22242 http://ftp.pdbj.org/pub/emdb/structures/EMD-22242 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22242 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22242 | HTTPS FTP |
-Related structure data
| Related structure data |  6xlfMC  6xlbC  6xlcC  6xldC  6xleC  6xlgC  6xlhC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22242.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22242.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Hsc82 in complex with Aha1 in the presence of AMP-PNP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.059 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
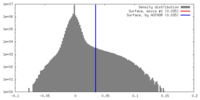
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Full-length Hsc82 in complex with Aha1 in the presence of AMP-PNP
| Entire | Name: Full-length Hsc82 in complex with Aha1 in the presence of AMP-PNP |
|---|---|
| Components |
|
-Supramolecule #1: Full-length Hsc82 in complex with Aha1 in the presence of AMP-PNP
| Supramolecule | Name: Full-length Hsc82 in complex with Aha1 in the presence of AMP-PNP type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
-Macromolecule #1: ATP-dependent molecular chaperone HSC82
| Macromolecule | Name: ATP-dependent molecular chaperone HSC82 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 81.003594 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAGETFEFQA EITQLMSLII NTVYSNKEIF LRELISNASD ALDKIRYQAL SDPKQLETEP DLFIRITPKP EEKVLEIRDS GIGMTKAEL INNLGTIAKS GTKAFMEALS AGADVSMIGQ FGVGFYSLFL VADRVQVISK NNEDEQYIWE SNAGGSFTVT L DEVNERIG ...String: MAGETFEFQA EITQLMSLII NTVYSNKEIF LRELISNASD ALDKIRYQAL SDPKQLETEP DLFIRITPKP EEKVLEIRDS GIGMTKAEL INNLGTIAKS GTKAFMEALS AGADVSMIGQ FGVGFYSLFL VADRVQVISK NNEDEQYIWE SNAGGSFTVT L DEVNERIG RGTVLRLFLK DDQLEYLEEK RIKEVIKRHS EFVAYPIQLL VTKEVEKEVP IPEEEKKDEE KKDEDDKKPK LE EVDEEEE EKKPKTKKVK EEVQELEELN KTKPLWTRNP SDITQEEYNA FYKSISNDWE DPLYVKHFSV EGQLEFRAIL FIP KRAPFD LFESKKKKNN IKLYVRRVFI TDEAEDLIPE WLSFVKGVVD SEDLPLNLSR EMLQQNKIMK VIRKNIVKKL IEAF NEIAE DSEQFDKFYS AFAKNIKLGV HEDTQNRAAL AKLLRYNSTK SVDELTSLTD YVTRMPEHQK NIYYITGESL KAVEK SPFL DALKAKNFEV LFLTDPIDEY AFTQLKEFEG KTLVDITKDF ELEETDEEKA EREKEIKEYE PLTKALKDIL GDQVEK VVV SYKLLDAPAA IRTGQFGWSA NMERIMKAQA LRDSSMSSYM SSKKTFEISP KSPIIKELKK RVDEGGAQDK TVKDLTN LL FETALLTSGF SLEEPTSFAS RINRLISLGL NIDEDEETET APEASTEAPV EEVPADTEME EVD UniProtKB: ATP-dependent molecular chaperone HSC82 |
-Macromolecule #2: Hsp90 co-chaperone AHA1
| Macromolecule | Name: Hsp90 co-chaperone AHA1 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 39.486422 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVVNNPNNWH WVDKNCIGWA KEYFKQKLVG VEAGSVKDKK YAKIKSVSSI EGDCEVNQRK GKVISLFDLK ITVLIEGHVD SKDGSALPF EGSINVPEVA FDSEASSYQF DISIFKETSE LSEAKPLIRS ELLPKLRQIF QQFGKDLLAT HGNDIQVPES Q VKSNYTRG ...String: MVVNNPNNWH WVDKNCIGWA KEYFKQKLVG VEAGSVKDKK YAKIKSVSSI EGDCEVNQRK GKVISLFDLK ITVLIEGHVD SKDGSALPF EGSINVPEVA FDSEASSYQF DISIFKETSE LSEAKPLIRS ELLPKLRQIF QQFGKDLLAT HGNDIQVPES Q VKSNYTRG NQKSSFTEIK DSASKPKKNA LPSSTSTSAP VSSTNKVPQN GSGNSTSIYL EPTFNVPSSE LYETFLDKQR IL AWTRSAQ FFNSGPKLET KEKFELFGGN VISELVSCEK DKKLVFHWKL KDWSAPFNST IEMTFHESQE FHETKLQVKW TGI PVGEED RVRANFEEYY VRSIKLTFGF GAVL UniProtKB: Hsp90 co-chaperone AHA1 |
-Macromolecule #3: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER / type: ligand / ID: 3 / Number of copies: 2 / Formula: ANP |
|---|---|
| Molecular weight | Theoretical: 506.196 Da |
| Chemical component information |  ChemComp-ANP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 72.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-6xlf: |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)