+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-20934 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | SWI/SNF nucleosome complex with ADP-BeFx | ||||||||||||
 マップデータ マップデータ | SWI/SNF nucleosome complex assembled in the presence of ADP-BeFx | ||||||||||||
 試料 試料 |
| ||||||||||||
 キーワード キーワード | SWI/SNF / chromatin remodeler / TRANSCRIPTION / TRANSCRIPTION-DNA complex | ||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報carbon catabolite activation of transcription from RNA polymerase II promoter / RHO GTPases activate IQGAPs / RHO GTPases Activate WASPs and WAVEs / Regulation of actin dynamics for phagocytic cup formation / positive regulation of cell adhesion involved in single-species biofilm formation / positive regulation of mating type switching / positive regulation of invasive growth in response to glucose limitation / aggrephagy / Platelet degranulation / rDNA binding ...carbon catabolite activation of transcription from RNA polymerase II promoter / RHO GTPases activate IQGAPs / RHO GTPases Activate WASPs and WAVEs / Regulation of actin dynamics for phagocytic cup formation / positive regulation of cell adhesion involved in single-species biofilm formation / positive regulation of mating type switching / positive regulation of invasive growth in response to glucose limitation / aggrephagy / Platelet degranulation / rDNA binding / HDACs deacetylate histones / nucleosome disassembly / DNA strand invasion / RSC-type complex / ATP-dependent chromatin remodeler activity / SWI/SNF complex / nuclear chromosome / NuA4 histone acetyltransferase complex / positive regulation of transcription by RNA polymerase I / anatomical structure morphogenesis / ATP-dependent activity, acting on DNA / nucleosomal DNA binding / cellular response to amino acid starvation / helicase activity / maturation of LSU-rRNA / chromosome segregation / transcription elongation by RNA polymerase II / nucleotide-excision repair / 加水分解酵素; 酸無水物に作用; 酸無水物に作用・細胞または細胞小器官の運動に関与 / double-strand break repair via homologous recombination / lysine-acetylated histone binding / DNA-templated DNA replication / structural constituent of chromatin / nucleosome / double-strand break repair / nucleosome assembly / chromatin organization / histone binding / RNA polymerase II-specific DNA-binding transcription factor binding / transcription cis-regulatory region binding / hydrolase activity / chromatin remodeling / protein heterodimerization activity / chromatin binding / chromatin / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / structural molecule activity / positive regulation of DNA-templated transcription / positive regulation of transcription by RNA polymerase II / DNA binding / nucleoplasm / ATP binding / nucleus / metal ion binding / cytosol 類似検索 - 分子機能 | ||||||||||||
| 生物種 |    | ||||||||||||
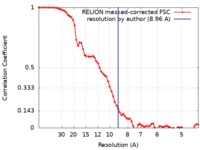
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 8.96 Å | ||||||||||||
 データ登録者 データ登録者 | He Y / Han Y | ||||||||||||
| 資金援助 |  米国, 3件 米国, 3件
| ||||||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2020 ジャーナル: Nature / 年: 2020タイトル: Cryo-EM structure of SWI/SNF complex bound to a nucleosome. 著者: Yan Han / Alexis A Reyes / Sara Malik / Yuan He /  要旨: The chromatin-remodelling complex SWI/SNF is highly conserved and has critical roles in various cellular processes, including transcription and DNA-damage repair. It hydrolyses ATP to remodel ...The chromatin-remodelling complex SWI/SNF is highly conserved and has critical roles in various cellular processes, including transcription and DNA-damage repair. It hydrolyses ATP to remodel chromatin structure by sliding and evicting histone octamers, creating DNA regions that become accessible to other essential factors. However, our mechanistic understanding of the remodelling activity is hindered by the lack of a high-resolution structure of complexes from this family. Here we report the cryo-electron microscopy structure of Saccharomyces cerevisiae SWI/SNF bound to a nucleosome, at near-atomic resolution. In the structure, the actin-related protein (Arp) module is sandwiched between the ATPase and the rest of the complex, with the Snf2 helicase-SANT associated (HSA) domain connecting all modules. The body contains an assembly scaffold composed of conserved subunits Snf12 (also known as SMARCD or BAF60), Snf5 (also known as SMARCB1, BAF47 or INI1) and an asymmetric dimer of Swi3 (also known as SMARCC, BAF155 or BAF170). Another conserved subunit, Swi1 (also known as ARID1 or BAF250), resides in the core of SWI/SNF, acting as a molecular hub. We also observed interactions between Snf5 and the histones at the acidic patch, which could serve as an anchor during active DNA translocation. Our structure enables us to map and rationalize a subset of cancer-related mutations in the human SWI/SNF complex and to propose a model for how SWI/SNF recognizes and remodels the +1 nucleosome to generate nucleosome-depleted regions during gene activation. | ||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_20934.map.gz emd_20934.map.gz | 2.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-20934-v30.xml emd-20934-v30.xml emd-20934.xml emd-20934.xml | 38.8 KB 38.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_20934_fsc.xml emd_20934_fsc.xml | 6.9 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_20934.png emd_20934.png | 76.2 KB | ||
| Filedesc metadata |  emd-20934.cif.gz emd-20934.cif.gz | 11.9 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20934 http://ftp.pdbj.org/pub/emdb/structures/EMD-20934 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20934 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20934 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_20934_validation.pdf.gz emd_20934_validation.pdf.gz | 388.5 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_20934_full_validation.pdf.gz emd_20934_full_validation.pdf.gz | 388 KB | 表示 | |
| XML形式データ |  emd_20934_validation.xml.gz emd_20934_validation.xml.gz | 9.5 KB | 表示 | |
| CIF形式データ |  emd_20934_validation.cif.gz emd_20934_validation.cif.gz | 12.3 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20934 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20934 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20934 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20934 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_20934.map.gz / 形式: CCP4 / 大きさ: 27 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_20934.map.gz / 形式: CCP4 / 大きさ: 27 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | SWI/SNF nucleosome complex assembled in the presence of ADP-BeFx | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 2.24 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : SWI/SNF nucleosome complex with ADP-BeFx
+超分子 #1: SWI/SNF nucleosome complex with ADP-BeFx
+分子 #1: Histone H3.2
+分子 #2: Histone H4
+分子 #3: Histone H2A type 1
+分子 #4: Histone H2B 1.1
+分子 #7: Actin-related protein 7
+分子 #8: Actin-like protein ARP9
+分子 #9: Regulator of Ty1 transposition protein 102
+分子 #10: Transcription regulatory protein SNF2
+分子 #11: SWI/SNF chromatin-remodeling complex subunit SWI1
+分子 #12: SWI/SNF chromatin-remodeling complex subunit SNF5
+分子 #13: SWI/SNF complex subunit SWI3
+分子 #14: Transcription regulatory protein SNF12
+分子 #15: Transcription regulatory protein SNF6
+分子 #16: Unknown protein
+分子 #17: SWI/SNF global transcription activator complex subunit SWP82
+分子 #5: 601 sequence bottom strand
+分子 #6: 601 sequence top strand
+分子 #18: PHOSPHATE ION
+分子 #19: ADENOSINE-5'-DIPHOSPHATE
+分子 #20: BERYLLIUM TRIFLUORIDE ION
+分子 #21: MAGNESIUM ION
+分子 #22: water
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.9 詳細: 10 mM HEPES, pH 7.9, 10 mM MgCl2, 50 mM KCl, 1 mM DTT, 5% glycerol, 0.05% NP-40 |
|---|---|
| グリッド | 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: CONTINUOUS / 詳細: unspecified |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | JEOL 3200FS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: COUNTING / 平均電子線量: 76.5 e/Å2 |
| 電子線 | 加速電圧: 200 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 試料ステージ | 試料ホルダーモデル: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER ホルダー冷却材: NITROGEN |
 ムービー
ムービー コントローラー
コントローラー