+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-20840 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | FACT_subnucleosome complex 1 | |||||||||
 マップデータ マップデータ | Sharpened map | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | assembly / disassembly / transient / integrity / TRANSCRIPTION / TRANSCRIPTION-DNA complex | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報FACT complex / regulation of chromatin organization / positive regulation of DNA-templated transcription, elongation / nucleosome disassembly / Pausing and recovery of Tat-mediated HIV elongation / Tat-mediated HIV elongation arrest and recovery / HIV elongation arrest and recovery / Pausing and recovery of HIV elongation / nucleosome binding / Tat-mediated elongation of the HIV-1 transcript ...FACT complex / regulation of chromatin organization / positive regulation of DNA-templated transcription, elongation / nucleosome disassembly / Pausing and recovery of Tat-mediated HIV elongation / Tat-mediated HIV elongation arrest and recovery / HIV elongation arrest and recovery / Pausing and recovery of HIV elongation / nucleosome binding / Tat-mediated elongation of the HIV-1 transcript / negative regulation of megakaryocyte differentiation / Formation of HIV-1 elongation complex containing HIV-1 Tat / Formation of HIV elongation complex in the absence of HIV Tat / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / RNA Polymerase II Transcription Elongation / Formation of RNA Pol II elongation complex / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / nucleosomal DNA binding / RNA Polymerase II Pre-transcription Events / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / telomere organization / Meiotic synapsis / Interleukin-7 signaling / RNA Polymerase I Promoter Opening / epigenetic regulation of gene expression / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / Assembly of the ORC complex at the origin of replication / SUMOylation of chromatin organization proteins / DNA methylation / Regulation of endogenous retroelements by KRAB-ZFP proteins / Condensation of Prophase Chromosomes / SIRT1 negatively regulates rRNA expression / HCMV Late Events / Chromatin modifications during the maternal to zygotic transition (MZT) / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / innate immune response in mucosa / PRC2 methylates histones and DNA / Defective pyroptosis / HDACs deacetylate histones / TP53 Regulates Transcription of DNA Repair Genes / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / Transcriptional regulation by small RNAs / transcription elongation by RNA polymerase II / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / NoRC negatively regulates rRNA expression / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / G2/M DNA damage checkpoint / HDMs demethylate histones / B-WICH complex positively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / heterochromatin formation / PKMTs methylate histone lysines / Metalloprotease DUBs / Meiotic recombination / Pre-NOTCH Transcription and Translation / RMTs methylate histone arginines / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / UCH proteinases / antimicrobial humoral immune response mediated by antimicrobial peptide / nucleosome / nucleosome assembly / E3 ubiquitin ligases ubiquitinate target proteins / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / antibacterial humoral response / chromatin organization / RUNX1 regulates transcription of genes involved in differentiation of HSCs / Factors involved in megakaryocyte development and platelet production / HATs acetylate histones / Processing of DNA double-strand break ends / gene expression / histone binding / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence / Estrogen-dependent gene expression / Regulation of TP53 Activity through Phosphorylation / DNA replication / transcription by RNA polymerase II / chromosome, telomeric region / Ub-specific processing proteases / defense response to Gram-positive bacterium / cadherin binding / protein heterodimerization activity / negative regulation of cell population proliferation / Amyloid fiber formation / DNA repair / nucleolus 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  | |||||||||
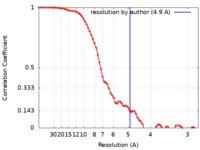
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 4.9 Å | |||||||||
 データ登録者 データ登録者 | Zhou K / Tan YZ | |||||||||
| 資金援助 |  米国, 1件 米国, 1件
| |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2020 ジャーナル: Nature / 年: 2020タイトル: FACT caught in the act of manipulating the nucleosome. 著者: Yang Liu / Keda Zhou / Naifu Zhang / Hui Wei / Yong Zi Tan / Zhening Zhang / Bridget Carragher / Clinton S Potter / Sheena D'Arcy / Karolin Luger /  要旨: The organization of genomic DNA into nucleosomes profoundly affects all DNA-related processes in eukaryotes. The histone chaperone known as 'facilitates chromatin transcription' (FACT) (consisting of ...The organization of genomic DNA into nucleosomes profoundly affects all DNA-related processes in eukaryotes. The histone chaperone known as 'facilitates chromatin transcription' (FACT) (consisting of subunits SPT16 and SSRP1) promotes both disassembly and reassembly of nucleosomes during gene transcription, DNA replication and DNA repair. However, the mechanism by which FACT causes these opposing outcomes is unknown. Here we report two cryo-electron-microscopic structures of human FACT in complex with partially assembled subnucleosomes, with supporting biochemical and hydrogen-deuterium exchange data. We find that FACT is engaged in extensive interactions with nucleosomal DNA and all histone variants. The large DNA-binding surface on FACT appears to be protected by the carboxy-terminal domains of both of its subunits, and this inhibition is released by interaction with H2A-H2B, allowing FACT-H2A-H2B to dock onto a complex containing DNA and histones H3 and H4 (ref. ). SPT16 binds nucleosomal DNA and tethers H2A-H2B through its carboxy-terminal domain by acting as a placeholder for DNA. SSRP1 also contributes to DNA binding, and can assume two conformations, depending on whether a second H2A-H2B dimer is present. Our data suggest a compelling mechanism for how FACT maintains chromatin integrity during polymerase passage, by facilitating removal of the H2A-H2B dimer, stabilizing intermediate subnucleosomal states and promoting nucleosome reassembly. Our findings reconcile discrepancies regarding the many roles of FACT and underscore the dynamic interactions between histone chaperones and nucleosomes. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_20840.map.gz emd_20840.map.gz | 28.6 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-20840-v30.xml emd-20840-v30.xml emd-20840.xml emd-20840.xml | 26.8 KB 26.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_20840_fsc.xml emd_20840_fsc.xml | 7.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_20840.png emd_20840.png | 150.7 KB | ||
| マスクデータ |  emd_20840_msk_1.map emd_20840_msk_1.map | 30.5 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-20840.cif.gz emd-20840.cif.gz | 7.7 KB | ||
| その他 |  emd_20840_additional.map.gz emd_20840_additional.map.gz emd_20840_half_map_1.map.gz emd_20840_half_map_1.map.gz emd_20840_half_map_2.map.gz emd_20840_half_map_2.map.gz | 9.6 MB 28.3 MB 28.3 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20840 http://ftp.pdbj.org/pub/emdb/structures/EMD-20840 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20840 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20840 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_20840_validation.pdf.gz emd_20840_validation.pdf.gz | 1.1 MB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_20840_full_validation.pdf.gz emd_20840_full_validation.pdf.gz | 1.1 MB | 表示 | |
| XML形式データ |  emd_20840_validation.xml.gz emd_20840_validation.xml.gz | 13 KB | 表示 | |
| CIF形式データ |  emd_20840_validation.cif.gz emd_20840_validation.cif.gz | 18 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20840 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20840 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20840 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20840 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  6upkMC  6uplC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | |
| 電子顕微鏡画像生データ |  EMPIAR-10333 (タイトル: Single Particle Cryo-EM Reconstructions of Human FACT in Complex with Partially Assembled Sub-nucleosomes EMPIAR-10333 (タイトル: Single Particle Cryo-EM Reconstructions of Human FACT in Complex with Partially Assembled Sub-nucleosomesData size: 2.2 TB Data #1: Unaligned multi-frame micrographs [micrographs - multiframe] Data #2: Aligned and dose-weighted micrographs [micrographs - single frame] Data #3: Final Particle Stacks for Class 1 and 2 with Euler Angles and Shifts [picked particles - single frame - processed]) |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_20840.map.gz / 形式: CCP4 / 大きさ: 30.5 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_20840.map.gz / 形式: CCP4 / 大きさ: 30.5 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.40301 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-マスク #1
| ファイル |  emd_20840_msk_1.map emd_20840_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
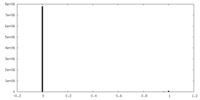
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-追加マップ: unfiltered map
| ファイル | emd_20840_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | unfiltered map | ||||||||||||
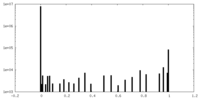
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_20840_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
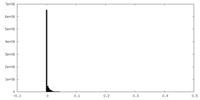
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_20840_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
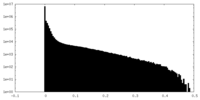
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : FACT_subNucleosome_complex_class1
| 全体 | 名称: FACT_subNucleosome_complex_class1 |
|---|---|
| 要素 |
|
-超分子 #1: FACT_subNucleosome_complex_class1
| 超分子 | 名称: FACT_subNucleosome_complex_class1 / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 320 KDa |
-分子 #1: Histone H3.1
| 分子 | 名称: Histone H3.1 / タイプ: protein_or_peptide / ID: 1 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 15.437167 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MARTKQTARK STGGKAPRKQ LATKAARKSA PATGGVKKPH RYRPGTVALR EIRRYQKSTE LLIRKLPFQR LVREIAQDFK TDLRFQSSA VMALQEACEA YLVGLFEDTN LCAIHAKRVT IMPKDIQLAR RIRGERA UniProtKB: Histone H3.1 |
-分子 #2: Histone H4
| 分子 | 名称: Histone H4 / タイプ: protein_or_peptide / ID: 2 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 11.394426 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-分子 #3: Histone H2A
| 分子 | 名称: Histone H2A / タイプ: protein_or_peptide / ID: 3 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 14.135523 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSGRGKQGGK ARAKAKSRSS RAGLQFPVGR VHRLLRKGNY AERVGAGAPV YLAAVLEYLT AEILELAGNA ARDNKKTRII PRHLQLAIR NDEELNKLLG RVTIAQGGVL PNIQAVLLPK KTESHHKAKG K UniProtKB: Histone H2A type 1-C |
-分子 #4: Histone H2B
| 分子 | 名称: Histone H2B / タイプ: protein_or_peptide / ID: 4 / コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 13.937213 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MPEPAKSAPA PKKGSKKAVT KAQKKDGKKR KRSRKESYSV YVYKVLKQVH PDTGISSKAM GIMNSFVNDI FERIAGEASR LAHYNKRST ITSREIQTAV RLLLPGELAK HAVSEGTKAV TKYTSSK UniProtKB: Histone H2B type 1-C/E/F/G/I |
-分子 #5: FACT complex subunit SPT16
| 分子 | 名称: FACT complex subunit SPT16 / タイプ: protein_or_peptide / ID: 5 詳細: Amino acids 640-651 cannot be identified due to the resolution limitations. To show the connectivity of the electron density, poly-(UNK) was placed. This was also done for the end of this ...詳細: Amino acids 640-651 cannot be identified due to the resolution limitations. To show the connectivity of the electron density, poly-(UNK) was placed. This was also done for the end of this chain (926-967). The full sequence for the experiment is: MHHHHHHAVTLDKDAYYRRVKRLYSNWRKGEDEYANVDAIVVSVGVDEEIVYAKSTALQTWLFGYELTDTIMVFCDDKII FMASKKKVEFLKQIANTKGNENANGAPAITLLIREKNESNKSSFDKMIEAIKESKNGKKIGVFSKDKFPGEFMKSWNDCL NKEGFDKIDISAVVAYTIAVKEDGELNLMKKAASITSEVFNKFFKERVMEIVDADEKVRHSKLAESVEKAIEEKKYLAGA DPSTVEMCYPPIIQSGGNYNLKFSVVSDKNHMHFGAITCAMGIRFKSYCSNLVRTLMVDPSQEVQENYNFLLQLQEELLK ELRHGVKICDVYNAVMDVVKKQKPELLNKITKNLGFGMGIEFREGSLVINSKNQYKLKKGMVFSINLGFSDLTNKEGKKP EEKTYALFIGDTVLVDEDGPATVLTSVKKKVKNVGIFLKNEDEEEEEEEKDEAEDLLGRGSRAALLTERTRNEMTAEEKR RAHQKELAAQLNEEAKRRLTEQKGEQQIQKARKSNVSYKNPSLMPKEPHIREMKIYIDKKYETVIMPVFGIATPFHIATI KNISMSVEGDYTYLRINFYCPGSALGRNEGNIFPNPEATFVKEITYRASNIKAPGEQTVPALNLQNAFRIIKEVQKRYKT REAEEKEKEGIVKQDSLVINLNRSNPKLKDLYIRPNIAQKRMQGSLEAHVNGFRFTSVRGDKVDILYNNIKHALFQPCDG EMIIVLHFHLKNAIMFGKKRHTDVQFYTEVGEITTDLGKHQHMHDRDDLYAEQMEREMRHKLKTAFKNFIEKVEALTKEE LEFEVPFRDLGFNGAPYRSTCLLQPTSSALVNATEWPPFVVTLDEVELIHFERVQFHLKNFDMVIVYKDYSKKVTMINAI PVASLDPIKEWLNSCDLKYTEGVQSLNWTKIMKTIVDDPEGFFEQGGWSFLEPEGEGSDAEEGDSESEIEDETFNPSEDD YEEEEEDSDEDYSSEAEESDYSKESLGSEEESGKDWDELEEEARKADRESRYEEEEEQSRSMSRKRKASVHSSGRGSNRG SRHSSAPPKKKRK コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 109.574867 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MHHHHHHAVT LDKDAYYRRV KRLYSNWRKG EDEYANVDAI VVSVGVDEEI VYAKSTALQT WLFGYELTDT IMVFCDDKII FMASKKKVE FLKQIANTKG NENANGAPAI TLLIREKNES NKSSFDKMIE AIKESKNGKK IGVFSKDKFP GEFMKSWNDC L NKEGFDKI ...文字列: MHHHHHHAVT LDKDAYYRRV KRLYSNWRKG EDEYANVDAI VVSVGVDEEI VYAKSTALQT WLFGYELTDT IMVFCDDKII FMASKKKVE FLKQIANTKG NENANGAPAI TLLIREKNES NKSSFDKMIE AIKESKNGKK IGVFSKDKFP GEFMKSWNDC L NKEGFDKI DISAVVAYTI AVKEDGELNL MKKAASITSE VFNKFFKERV MEIVDADEKV RHSKLAESVE KAIEEKKYLA GA DPSTVEM CYPPIIQSGG NYNLKFSVVS DKNHMHFGAI TCAMGIRFKS YCSNLVRTLM VDPSQEVQEN YNFLLQLQEE LLK ELRHGV KICDVYNAVM DVVKKQKPEL LNKITKNLGF GMGIEFREGS LVINSKNQYK LKKGMVFSIN LGFSDLTNKE GKKP EEKTY ALFIGDTVLV DEDGPATVLT SVKKKVKNVG IFLKNEDEEE EEEEKDEAED LLGRGSRAAL LTERTRNEMT AEEKR RAHQ KELAAQLNEE AKRRLTEQKG EQQIQKARKS NVSYKNPSLM PKEPHIREMK IYIDKKYETV IMPVFGIATP FHIATI KNI SMSVEGDYTY LRINFYCPGS ALGRNEGNIF PNPEATFVKE ITYRASNIKA PGEQTVPALN LQNAFRIIKE VQKRYK (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)V KQDSLVINLN RSNPKLKDL YIRPNIAQKR MQGSLEAHVN GFRFTSVRGD KVDILYNNIK HALFQPCDGE MIIVLHFHLK NAIMFGKKRH TDVQFYTEVG EITTDLGKH QHMHDRDDLY AEQMEREMRH KLKTAFKNFI EKVEALTKEE LEFEVPFRDL GFNGAPYRST CLLQPTSSAL V NATEWPPF VVTLDEVELI HFERVQFHLK NFDMVIVYKD YSKKVTMINA IPVASLDPIK EWLNSCDLKY TEGVQSLNWT KI MKTIVDD PEGFFEQGGW SFL(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
-分子 #6: FACT complex subunit SSRP1
| 分子 | 名称: FACT complex subunit SSRP1 / タイプ: protein_or_peptide / ID: 6 詳細: The poly-(UNK) was placed to show the electron density between 171-198. The full sequence for the experiment is: ...詳細: The poly-(UNK) was placed to show the electron density between 171-198. The full sequence for the experiment is: MAETLEFNDVYQEVKGSMNDGRLRLSRQGIIFKNSKTGKVDNIQAGELTEGIWRRVALGHGLKLLTKNGHVYKYDGFRES EFEKLSDFFKTHYRLELMEKDLCVKGWNWGTVKFGGQLLSFDIGDQPVFEIPLSNVSQCTTGKNEVTLEFHQNDDAEVSL MEVRFYVPPTQEDGVDPVEAFAQNVLSKADVIQATGDAICIFRELQCLTPRGRYDIRIYPTFLHLHGKTFDYKIPYTTVL RLFLLPHKDQRQMFFVISLDPPIKQGQTRYHFLILLFSKDEDISLTLNMNEEEVEKRFEGRLTKNMSGSLYEMVSRVMKA LVNRKITVPGNFQGHSGAQCITCSYKASSGLLYPLERGFIYVHKPPVHIRFDEISFVNFARGTTTTRSFDFEIETKQGTQ YTFSSIEREEYGKLFDFVNAKKLNIKNRGLKEGMNPSYDEYADSDEDQHDAYLERMKEEGKIREENANDSSDDSGEETDE SFNPGEEEEDVAEEFDSNASASSSSNEGDSDRDEKKRKQLKKAKMAKDRKSRKKPVEVKKGKDPNAPKRPMSAYMLWLNA SREKIKSDHPGISITDLSKKAGEIWKGMSKEKKEEWDRKAEDARRDYEKAMKEYEGGRGESSKRDKSKKKKKVKVKMEKK STPSRGSSSKSSSRQLSESFKSKEFVSSDESSSGENKSKKKRRRSEDSEEEELASTPPSSEDSASGSDE コピー数: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 73.361172 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MAETLEFNDV YQEVKGSMND GRLRLSRQGI IFKNSKTGKV DNIQAGELTE GIWRRVALGH GLKLLTKNGH VYKYDGFRES EFEKLSDFF KTHYRLELME KDLCVKGWNW GTVKFGGQLL SFDIGDQPVF EIPLSNVSQC TTGKNEVTLE FHQNDDAEVS L MEVRFYVP ...文字列: MAETLEFNDV YQEVKGSMND GRLRLSRQGI IFKNSKTGKV DNIQAGELTE GIWRRVALGH GLKLLTKNGH VYKYDGFRES EFEKLSDFF KTHYRLELME KDLCVKGWNW GTVKFGGQLL SFDIGDQPVF EIPLSNVSQC TTGKNEVTLE FHQNDDAEVS L MEVRFYVP PTQ(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)D AICIFRELQC LTPRGRYDI RIYPTFLHLH GKTFDYKIPY TTVLRLFLLP HKDQRQMFFV ISLDPPIKQG QTRYHFLILL FSKDEDISLT LNMNEEEVEK RFEGRLTKN MSGSLYEMVS RVMKALVNRK ITVPGNFQGH SGAQCITCSY KASSGLLYPL ERGFIYVHKP PVHIRFDEIS F VNFARGTT TTRSFDFEIE TKQGTQYTFS SIEREEYGKL FDFVNAKKLN IKNRGLKEGM NPSYDEYADS DEDQHDAYLE RM KEEGKIR EENANDSSDD SGEETDESFN PGEEEEDVAE EFDSNASASS SSNEGDSDRD EKKRKQLKKA KMAKDRKSRK KPV EVKKGK DPNAPKRPMS AYMLWLNASR EKIKSDHPGI SITDLSKKAG EIWKGMSKEK KEEWDRKAED ARRDYEKAMK EYEG GRGES SKRDKSKKKK KVKVKMEKK |
-分子 #7: DNA (79-mer)
| 分子 | 名称: DNA (79-mer) / タイプ: dna / ID: 7 / コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 24.144422 KDa |
| 配列 | 文字列: (DT)(DC)(DG)(DT)(DA)(DG)(DA)(DC)(DA)(DG) (DC)(DT)(DC)(DT)(DA)(DG)(DC)(DA)(DC)(DC) (DG)(DC)(DT)(DT)(DA)(DA)(DA)(DC)(DG) (DC)(DA)(DC)(DG)(DT)(DA)(DC)(DG)(DC)(DG) (DC) (DT)(DG)(DT)(DC)(DC) ...文字列: (DT)(DC)(DG)(DT)(DA)(DG)(DA)(DC)(DA)(DG) (DC)(DT)(DC)(DT)(DA)(DG)(DC)(DA)(DC)(DC) (DG)(DC)(DT)(DT)(DA)(DA)(DA)(DC)(DG) (DC)(DA)(DC)(DG)(DT)(DA)(DC)(DG)(DC)(DG) (DC) (DT)(DG)(DT)(DC)(DC)(DC)(DC)(DC) (DG)(DC)(DG)(DT)(DT)(DT)(DT)(DA)(DA)(DC) (DC)(DG) (DC)(DC)(DA)(DA)(DG)(DG)(DG) (DG)(DA)(DT)(DT)(DA)(DC)(DT)(DC)(DC)(DC) (DT)(DA) |
-分子 #8: DNA (79-mer)
| 分子 | 名称: DNA (79-mer) / タイプ: dna / ID: 8 / コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種:  |
| 分子量 | 理論値: 24.58468 KDa |
| 配列 | 文字列: (DT)(DA)(DG)(DG)(DG)(DA)(DG)(DT)(DA)(DA) (DT)(DC)(DC)(DC)(DC)(DT)(DT)(DG)(DG)(DC) (DG)(DG)(DT)(DT)(DA)(DA)(DA)(DA)(DC) (DG)(DC)(DG)(DG)(DG)(DG)(DG)(DA)(DC)(DA) (DG) (DC)(DG)(DC)(DG)(DT) ...文字列: (DT)(DA)(DG)(DG)(DG)(DA)(DG)(DT)(DA)(DA) (DT)(DC)(DC)(DC)(DC)(DT)(DT)(DG)(DG)(DC) (DG)(DG)(DT)(DT)(DA)(DA)(DA)(DA)(DC) (DG)(DC)(DG)(DG)(DG)(DG)(DG)(DA)(DC)(DA) (DG) (DC)(DG)(DC)(DG)(DT)(DA)(DC)(DG) (DT)(DG)(DC)(DG)(DT)(DT)(DT)(DA)(DA)(DG) (DC)(DG) (DG)(DT)(DG)(DC)(DT)(DA)(DG) (DA)(DG)(DC)(DT)(DG)(DT)(DC)(DT)(DA)(DC) (DG)(DA) |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.5 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 平均電子線量: 7.8 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)