+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Sulfolobus acidocaldarius threads (0406) filament. | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cell surface appendage / N-glycosylation / O-glycosylation / PROTEIN FIBRIL | |||||||||
| Function / homology | Uncharacterized protein Function and homology information Function and homology information | |||||||||
| Biological species |   Sulfolobus acidocaldarius (acidophilic) Sulfolobus acidocaldarius (acidophilic) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 2.65 Å | |||||||||
 Authors Authors | Isupov MN / Gaines M / McLaren M / Daum B | |||||||||
| Funding support | European Union, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Towards a molecular picture of the archaeal cell surface. Authors: Matthew C Gaines / Michail N Isupov / Mathew McLaren / Clara L Mollat / Risat Ul Haque / Jake K Stephenson / Shamphavi Sivabalasarma / Cyril Hanus / Daniel Kattnig / Vicki A M Gold / Sonja ...Authors: Matthew C Gaines / Michail N Isupov / Mathew McLaren / Clara L Mollat / Risat Ul Haque / Jake K Stephenson / Shamphavi Sivabalasarma / Cyril Hanus / Daniel Kattnig / Vicki A M Gold / Sonja Albers / Bertram Daum /    Abstract: Archaea produce various protein filaments with specialised functions. While some archaea produce only one type of filament, the archaeal model species Sulfolobus acidocaldarius generates four. These ...Archaea produce various protein filaments with specialised functions. While some archaea produce only one type of filament, the archaeal model species Sulfolobus acidocaldarius generates four. These include rotary swimming propellers analogous to bacterial flagella (archaella), pili for twitching motility (Aap), adhesive fibres (threads), and filaments facilitating homologous recombination upon UV stress (UV pili). Here, we use cryo-electron microscopy to describe the structure of the S. acidocaldarius archaellum at 2.0 Å resolution, and update the structures of the thread and the Aap pilus at 2.7 Å and 2.6 Å resolution, respectively. We define features unique to archaella of the order Sulfolobales and compare their structure to those of Aap and threads in the context of the S-layer. We define distinct N-glycan patterns in the three filaments and identify a putative O-glycosylation site in the thread. Finally, we ascertain whether N-glycan truncation leads to structural changes in archaella and Aap. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19608.map.gz emd_19608.map.gz | 31.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19608-v30.xml emd-19608-v30.xml emd-19608.xml emd-19608.xml | 14.6 KB 14.6 KB | Display Display |  EMDB header EMDB header |
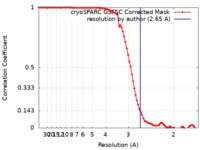
| FSC (resolution estimation) |  emd_19608_fsc.xml emd_19608_fsc.xml | 8.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_19608.png emd_19608.png | 45.4 KB | ||
| Filedesc metadata |  emd-19608.cif.gz emd-19608.cif.gz | 5.8 KB | ||
| Others |  emd_19608_half_map_1.map.gz emd_19608_half_map_1.map.gz emd_19608_half_map_2.map.gz emd_19608_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19608 http://ftp.pdbj.org/pub/emdb/structures/EMD-19608 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19608 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19608 | HTTPS FTP |
-Related structure data
| Related structure data |  8rzlMC  8qx4C  9etsC  9ettC  9ev0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_19608.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19608.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.829 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_19608_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_19608_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Filament 0406
| Entire | Name: Filament 0406 |
|---|---|
| Components |
|
-Supramolecule #1: Filament 0406
| Supramolecule | Name: Filament 0406 / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Sulfolobus acidocaldarius (acidophilic) Sulfolobus acidocaldarius (acidophilic) |
-Macromolecule #1: Sulfolobus acidocaldarius threads (0406) filament.
| Macromolecule | Name: Sulfolobus acidocaldarius threads (0406) filament. / type: protein_or_peptide / ID: 1 / Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Sulfolobus acidocaldarius (acidophilic) Sulfolobus acidocaldarius (acidophilic) |
| Molecular weight | Theoretical: 19.720252 KDa |
| Sequence | String: DVIYYYQGQI TVGNVAPPMY FAIQPNGNAK IGNNSNVPSY INAQPSSGGS GFTAQVNITN ATYNYYFNFM GLAVSKTGYI YLAKVAYSY TATNNPIQNA TLYIMNQQGQ IVYKYKLIVN GVVNSTLPST PLQINSGSYI VSLLIVPYQG TLPKTPSNDL A TITVNFGF SPMTASPPPI PLPSP UniProtKB: Uncharacterized protein |
-Macromolecule #3: alpha-D-mannopyranose
| Macromolecule | Name: alpha-D-mannopyranose / type: ligand / ID: 3 / Number of copies: 5 / Formula: MAN |
|---|---|
| Molecular weight | Theoretical: 180.156 Da |
| Chemical component information |  ChemComp-MAN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 3 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 3 / Number real images: 20579 / Average exposure time: 2.57 sec. / Average electron dose: 43.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8rzl: |
-Atomic model buiding 2
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-8rzl: |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)