+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the F-actin barbed end bound by formin mDia1 | ||||||||||||
 Map data Map data | Sharpened cryo-EM density map of the F-actin barbed end bound by the formin mDia1 | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | actin / formin / Cdc12 / profilin / actin assembly / STRUCTURAL PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationMGMT-mediated DNA damage reversal / negative regulation of neuron projection regeneration / multicellular organismal locomotion / RHOF GTPase cycle / RHOB GTPase cycle / ERBB2 Regulates Cell Motility / RHOC GTPase cycle / RHOD GTPase cycle / methylated-DNA-[protein]-cysteine S-methyltransferase / methylated-DNA-[protein]-cysteine S-methyltransferase activity ...MGMT-mediated DNA damage reversal / negative regulation of neuron projection regeneration / multicellular organismal locomotion / RHOF GTPase cycle / RHOB GTPase cycle / ERBB2 Regulates Cell Motility / RHOC GTPase cycle / RHOD GTPase cycle / methylated-DNA-[protein]-cysteine S-methyltransferase / methylated-DNA-[protein]-cysteine S-methyltransferase activity / RHOA GTPase cycle / actin nucleation / neuron projection retraction / positive regulation of norepinephrine uptake / cellular response to cytochalasin B / regulation of transepithelial transport / DNA-methyltransferase activity / morphogenesis of a polarized epithelium / RHO GTPases Activate Formins / bBAF complex / postsynaptic actin cytoskeleton organization / npBAF complex / nBAF complex / protein localization to adherens junction / brahma complex / profilin binding / postsynaptic actin cytoskeleton / Tat protein binding / structural constituent of postsynaptic actin cytoskeleton / GBAF complex / dense body / Formation of annular gap junctions / regulation of G0 to G1 transition / Gap junction degradation / Cell-extracellular matrix interactions / Folding of actin by CCT/TriC / DNA ligation / apical protein localization / regulation of double-strand break repair / adherens junction assembly / regulation of nucleotide-excision repair / regulation of microtubule-based process / Prefoldin mediated transfer of substrate to CCT/TriC / RSC-type complex / DNA alkylation repair / RHOF GTPase cycle / Regulation of MITF-M-dependent genes involved in pigmentation / Adherens junctions interactions / tight junction / axon midline choice point recognition / Sensory processing of sound by outer hair cells of the cochlea / regulation of norepinephrine uptake / Interaction between L1 and Ankyrins / Sensory processing of sound by inner hair cells of the cochlea / regulation of mitotic metaphase/anaphase transition / SWI/SNF complex / positive regulation of double-strand break repair / regulation of synaptic vesicle endocytosis / positive regulation of T cell differentiation / apical junction complex / establishment or maintenance of cell polarity / regulation of cyclin-dependent protein serine/threonine kinase activity / maintenance of blood-brain barrier / cortical cytoskeleton / positive regulation of stem cell population maintenance / NuA4 histone acetyltransferase complex / nitric-oxide synthase binding / regulation of G1/S transition of mitotic cell cycle / Recycling pathway of L1 / kinesin binding / brush border / calyx of Held / negative regulation of cell differentiation / synaptic vesicle endocytosis / positive regulation of double-strand break repair via homologous recombination / EPH-ephrin mediated repulsion of cells / ephrin receptor signaling pathway / RHO GTPases Activate WASPs and WAVEs / RHO GTPases activate IQGAPs / regulation of protein localization to plasma membrane / positive regulation of myoblast differentiation / cytoskeleton organization / EPHB-mediated forward signaling / substantia nigra development / actin filament polymerization / Neutrophil degranulation / axonogenesis / negative regulation of protein binding / methyltransferase activity / Translocation of SLC2A4 (GLUT4) to the plasma membrane / actin filament / cell motility / RHO GTPases Activate Formins / positive regulation of cell differentiation / adherens junction / sensory perception of sound / FCGR3A-mediated phagocytosis / regulation of transmembrane transporter activity / ruffle membrane / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.49 Å | ||||||||||||
 Authors Authors | Oosterheert W / Boiero Sanders M / Funk J / Prumbaum D / Raunser S / Bieling P | ||||||||||||
| Funding support |  Germany, European Union, 3 items Germany, European Union, 3 items
| ||||||||||||
 Citation Citation |  Journal: Science / Year: 2024 Journal: Science / Year: 2024Title: Molecular mechanism of actin filament elongation by formins. Authors: Wout Oosterheert / Micaela Boiero Sanders / Johanna Funk / Daniel Prumbaum / Stefan Raunser / Peter Bieling /  Abstract: Formins control the assembly of actin filaments (F-actin) that drive cell morphogenesis and motility in eukaryotes. However, their molecular interaction with F-actin and their mechanism of action ...Formins control the assembly of actin filaments (F-actin) that drive cell morphogenesis and motility in eukaryotes. However, their molecular interaction with F-actin and their mechanism of action remain unclear. In this work, we present high-resolution cryo-electron microscopy structures of F-actin barbed ends bound by three distinct formins, revealing a common asymmetric formin conformation imposed by the filament. Formation of new intersubunit contacts during actin polymerization sterically displaces formin and triggers its translocation. This "undock-and-lock" mechanism explains how actin-filament growth is coordinated with formin movement. Filament elongation speeds are controlled by the positioning and stability of actin-formin interfaces, which distinguish fast and slow formins. Furthermore, we provide a structure of the actin-formin-profilin ring complex, which resolves how profilin is rapidly released from the barbed end during filament elongation. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19503.map.gz emd_19503.map.gz | 398.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19503-v30.xml emd-19503-v30.xml emd-19503.xml emd-19503.xml | 27.3 KB 27.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19503_fsc.xml emd_19503_fsc.xml | 15.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_19503.png emd_19503.png | 95.5 KB | ||
| Masks |  emd_19503_msk_1.map emd_19503_msk_1.map | 421.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19503.cif.gz emd-19503.cif.gz | 8 KB | ||
| Others |  emd_19503_additional_1.map.gz emd_19503_additional_1.map.gz emd_19503_half_map_1.map.gz emd_19503_half_map_1.map.gz emd_19503_half_map_2.map.gz emd_19503_half_map_2.map.gz | 209.9 MB 390.9 MB 390.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19503 http://ftp.pdbj.org/pub/emdb/structures/EMD-19503 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19503 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19503 | HTTPS FTP |
-Validation report
| Summary document |  emd_19503_validation.pdf.gz emd_19503_validation.pdf.gz | 1.2 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19503_full_validation.pdf.gz emd_19503_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  emd_19503_validation.xml.gz emd_19503_validation.xml.gz | 25.4 KB | Display | |
| Data in CIF |  emd_19503_validation.cif.gz emd_19503_validation.cif.gz | 33 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19503 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19503 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19503 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19503 | HTTPS FTP |
-Related structure data
| Related structure data |  8ru2MC  8rttC  8rtyC  8ru0C  8rv2C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19503.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19503.map.gz / Format: CCP4 / Size: 421.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened cryo-EM density map of the F-actin barbed end bound by the formin mDia1 | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19503_msk_1.map emd_19503_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
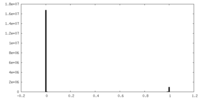
| Density Histograms |
-Additional map: 3D-refined, unsharpened cryo-EM density map of the F-actin...
| File | emd_19503_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D-refined, unsharpened cryo-EM density map of the F-actin barbed end bound by the formin mDia1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
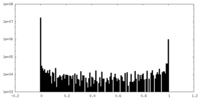
| Density Histograms |
-Half map: Unfiltered half map 1 of the F-actin barbed...
| File | emd_19503_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered half map 1 of the F-actin barbed end bound by the formin mDia1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
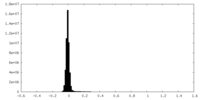
| Density Histograms |
-Half map: Unfiltered half map 2 of the F-actin barbed...
| File | emd_19503_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unfiltered half map 2 of the F-actin barbed end bound by the formin mDia1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
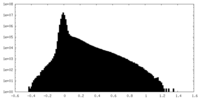
| Density Histograms |
- Sample components
Sample components
-Entire : mDia1-bound F-actin barbed end.
| Entire | Name: mDia1-bound F-actin barbed end. |
|---|---|
| Components |
|
-Supramolecule #1: mDia1-bound F-actin barbed end.
| Supramolecule | Name: mDia1-bound F-actin barbed end. / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 Details: Human beta-actin and mouse mDia1 were purified separately. Both proteins were mixed to assemble the complex prior to cryo-EM grid preparation. |
|---|
-Supramolecule #2: Actin filament
| Supramolecule | Name: Actin filament / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Mouse mDia1 (FH1FH2C domain)
| Supramolecule | Name: Mouse mDia1 (FH1FH2C domain) / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Actin, cytoplasmic 1, N-terminally processed
| Macromolecule | Name: Actin, cytoplasmic 1, N-terminally processed / type: protein_or_peptide / ID: 1 Details: Human beta-actin was recombinantly purified from BTI-Tnao38 cells. Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.632422 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: DDDIAALVVD NGSGMCKAGF AGDDAPRAVF PSIVGRPRHQ GVMVGMGQKD SYVGDEAQSK RGILTLKYPI E(HIC)GIVT NWD DMEKIWHHTF YNELRVAPEE HPVLLTEAPL NPKANREKMT QIMFETFNTP AMYVAIQAVL SLYASGRTTG IVMDSGD GV THTVPIYEGY ...String: DDDIAALVVD NGSGMCKAGF AGDDAPRAVF PSIVGRPRHQ GVMVGMGQKD SYVGDEAQSK RGILTLKYPI E(HIC)GIVT NWD DMEKIWHHTF YNELRVAPEE HPVLLTEAPL NPKANREKMT QIMFETFNTP AMYVAIQAVL SLYASGRTTG IVMDSGD GV THTVPIYEGY ALPHAILRLD LAGRDLTDYL MKILTERGYS FTTTAEREIV RDIKEKLCYV ALDFEQEMAT AASSSSLE K SYELPDGQVI TIGNERFRCP EALFQPSFLG MESAGIHETT FNSIMKCDVD IRKDLYANTV LSGGTTMYPG IADRMQKEI TALAPSTMKI KIIAPPERKY SVWIGGSILA SLSTFQQMWI SKQEYDESGP SIVHRKCF UniProtKB: Actin, cytoplasmic 1 |
-Macromolecule #2: Methylated-DNA--protein-cysteine methyltransferase,Protein diapha...
| Macromolecule | Name: Methylated-DNA--protein-cysteine methyltransferase,Protein diaphanous homolog 1 type: protein_or_peptide / ID: 2 / Details: Has a N-terminal snap-tag. / Number of copies: 2 / Enantiomer: LEVO EC number: methylated-DNA-[protein]-cysteine S-methyltransferase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 86.469258 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASTMDIKLT GEFAMDKDCE MKRTTLDSPL GKLELSGCEQ GLHEIKLLGK GTSAADAVEV PAPAAVLGGP EPLMQATAWL NAYFHQPEA IEEFPVPALH HPVFQQESFT RQVLWKLLKV VKFGEVISYQ QLAALAGNPA ATAAVKTALS GNPVPILIPC H RVVSSSGA ...String: MASTMDIKLT GEFAMDKDCE MKRTTLDSPL GKLELSGCEQ GLHEIKLLGK GTSAADAVEV PAPAAVLGGP EPLMQATAWL NAYFHQPEA IEEFPVPALH HPVFQQESFT RQVLWKLLKV VKFGEVISYQ QLAALAGNPA ATAAVKTALS GNPVPILIPC H RVVSSSGA VGGYEGGLAV KEWLLAHEGH RLGKPGLGPA GGSPGGGSGG SEMASLSAVV VAPSVSSSAA VPPAPPLPGD SG TVIPPPP PGMGVPPPPP FGFGVPAAPV LPFGLTPKKV YKPEVQLRRP NWSKFVAEDL SQDCFWTKVK EDRFENNELF AKL TLAFSA QTKTSKAKKD QEGGEEKKSV QKKKVKELKV LDSKTAQNLS IFLGSFRMPY QEIKNVILEV NEAVLTESMI QNLI KQMPE PEQLKMLSEL KEEYDDLAES EQFGVVMGTV PRLRPRLNAI LFKLQFSEQV ENIKPEIVSV TAACEELRKS ENFSS LLEL TLLVGNYMNA GSRNAGAFGF NISFLCKLRD TKSADQKMTL LHFLAELCEN DHPEVLKFPD ELAHVEKASR VSAENL QKS LDQMKKQIAD VERDVQNFPA ATDEKDKFVE KMTSFVKDAQ EQYNKLRMMH SNMETLYKEL GDYFVFDPKK LSVEEFF MD LHNFRNMFLQ AVKENQKRRE TEEKMRRAKL AKEKAEKERL EKQQKREQLI DMNAEGDETG VMDSLLEALQ SGAAFRRK R GPRQVNRKAG CAVTSLLASE LTKDDAMAPG PVKVPKKSEG VPTILEEAKE LVGRASHHHH HH UniProtKB: Methylated-DNA--protein-cysteine methyltransferase, Protein diaphanous homolog 1, Protein diaphanous homolog 1 |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 3 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 3 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.1 Component:
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: GOLD / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 286 K / Instrument: FEI VITROBOT MARK IV / Details: 3 seconds, force 0.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: Titan Krios G2 microscope (Thermo Fisher Scientific) with an in-column Cs-corrector. Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 15 eV / Details: Gatan energy filter. |
| Details | 300 kV Titan Krios G2 microscope (Thermo Fisher Scientific) with an in-column Cs-corrector. |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 2 / Number real images: 38913 / Average electron dose: 67.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Details | Refinement performed using phenix real-space refine | |||||||||
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT | |||||||||
| Output model |  PDB-8ru2: |
 Movie
Movie Controller
Controller


































 Z
Z Y
Y X
X