[English] 日本語
 Yorodumi
Yorodumi- EMDB-17006: CryoEM Structure INO80core Hexasome complex composite map state1 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM Structure INO80core Hexasome complex composite map state1 | |||||||||||||||
 Map data Map data | ctINO80 hexasome complex composite map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | ATP-dependent chromatin remodeler / DNA BINDING PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationDASH complex / protein transport along microtubule to mitotic spindle pole body / mitotic sister chromatid biorientation / attachment of spindle microtubules to kinetochore / R2TP complex / Swr1 complex / attachment of mitotic spindle microtubules to kinetochore / Ino80 complex / box C/D snoRNP assembly / NuA4 histone acetyltransferase complex ...DASH complex / protein transport along microtubule to mitotic spindle pole body / mitotic sister chromatid biorientation / attachment of spindle microtubules to kinetochore / R2TP complex / Swr1 complex / attachment of mitotic spindle microtubules to kinetochore / Ino80 complex / box C/D snoRNP assembly / NuA4 histone acetyltransferase complex / Replacement of protamines by nucleosomes in the male pronucleus / arachidonate 15-lipoxygenase / arachidonate 15-lipoxygenase activity / Packaging Of Telomere Ends / lipoxygenase pathway / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / telomere organization / arachidonate metabolic process / Chromatin modifying enzymes / lipid oxidation / Deposition of new CENPA-containing nucleosomes at the centromere / hepoxilin biosynthetic process / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / linoleic acid metabolic process / Meiotic synapsis / Inhibition of DNA recombination at telomere / nucleosomal DNA binding / RNA Polymerase I Promoter Opening / Assembly of the ORC complex at the origin of replication / DNA helicase activity / Interleukin-7 signaling / epigenetic regulation of gene expression / DNA methylation / Condensation of Prophase Chromosomes / HCMV Late Events / SIRT1 negatively regulates rRNA expression / Chromatin modifications during the maternal to zygotic transition (MZT) / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / Defective pyroptosis / Meiotic recombination / innate immune response in mucosa / DNA Damage/Telomere Stress Induced Senescence / HDACs deacetylate histones / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / Transcriptional regulation by small RNAs / Transcriptional regulation of granulopoiesis / HDMs demethylate histones / Formation of the beta-catenin:TCF transactivating complex / HCMV Early Events / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / G2/M DNA damage checkpoint / NoRC negatively regulates rRNA expression / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / PKMTs methylate histone lysines / B-WICH complex positively regulates rRNA expression / heterochromatin formation / RMTs methylate histone arginines / mitotic spindle / Pre-NOTCH Transcription and Translation / Metalloprotease DUBs / kinetochore / Activation of anterior HOX genes in hindbrain development during early embryogenesis / structural constituent of chromatin / UCH proteinases / nucleosome / antimicrobial humoral immune response mediated by antimicrobial peptide / Factors involved in megakaryocyte development and platelet production / Processing of DNA double-strand break ends / nucleosome assembly / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / chromatin organization / E3 ubiquitin ligases ubiquitinate target proteins / Senescence-Associated Secretory Phenotype (SASP) / RUNX1 regulates transcription of genes involved in differentiation of HSCs / HATs acetylate histones / gene expression / antibacterial humoral response / Oxidative Stress Induced Senescence / DNA helicase / Estrogen-dependent gene expression / chromatin remodeling / Ub-specific processing proteases / defense response to Gram-positive bacterium / cadherin binding / protein heterodimerization activity / Amyloid fiber formation / negative regulation of cell population proliferation / DNA repair / regulation of transcription by RNA polymerase II / regulation of DNA-templated transcription / ATP hydrolysis activity / protein-containing complex / DNA binding / extracellular space / extracellular exosome / extracellular region Similarity search - Function | |||||||||||||||
| Biological species |  Thermochaetoides thermophila (fungus) / Thermochaetoides thermophila (fungus) /  Homo sapiens (human) / Synthetic construct (others) / synthetic construct (others) Homo sapiens (human) / Synthetic construct (others) / synthetic construct (others) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | |||||||||||||||
 Authors Authors | Zhang M / Jungblut A / Hoffmann T / Eustermann S | |||||||||||||||
| Funding support |  Germany, European Union, 4 items Germany, European Union, 4 items
| |||||||||||||||
 Citation Citation | Journal: Acta Crystallogr D Biol Crystallogr / Year: 2010 Title: Features and development of Coot. Authors: P Emsley / B Lohkamp / W G Scott / K Cowtan /  Abstract: Coot is a molecular-graphics application for model building and validation of biological macromolecules. The program displays electron-density maps and atomic models and allows model manipulations ...Coot is a molecular-graphics application for model building and validation of biological macromolecules. The program displays electron-density maps and atomic models and allows model manipulations such as idealization, real-space refinement, manual rotation/translation, rigid-body fitting, ligand search, solvation, mutations, rotamers and Ramachandran idealization. Furthermore, tools are provided for model validation as well as interfaces to external programs for refinement, validation and graphics. The software is designed to be easy to learn for novice users, which is achieved by ensuring that tools for common tasks are 'discoverable' through familiar user-interface elements (menus and toolbars) or by intuitive behaviour (mouse controls). Recent developments have focused on providing tools for expert users, with customisable key bindings, extensions and an extensive scripting interface. The software is under rapid development, but has already achieved very widespread use within the crystallographic community. The current state of the software is presented, with a description of the facilities available and of some of the underlying methods employed. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17006.map.gz emd_17006.map.gz | 28.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17006-v30.xml emd-17006-v30.xml emd-17006.xml emd-17006.xml | 44.2 KB 44.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_17006.png emd_17006.png | 168 KB | ||
| Filedesc metadata |  emd-17006.cif.gz emd-17006.cif.gz | 11.6 KB | ||
| Others |  emd_17006_additional_1.map.gz emd_17006_additional_1.map.gz | 140.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17006 http://ftp.pdbj.org/pub/emdb/structures/EMD-17006 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17006 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17006 | HTTPS FTP |
-Related structure data
| Related structure data |  8oo7MC  8oo9C  8ooaC  8oocC  8oofC  8ookC  8oopC  8oorC  8oosC  8ootC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17006.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17006.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ctINO80 hexasome complex composite map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.822 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: ctINO80 hexasome complex overall refinement (used as a...
| File | emd_17006_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ctINO80 hexasome complex overall refinement (used as a reference map to generate composite map) | ||||||||||||
| Projections & Slices |
| ||||||||||||
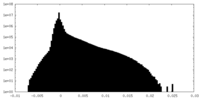
| Density Histograms |
- Sample components
Sample components
+Entire : INO80 core module in complex with hexasome
+Supramolecule #1: INO80 core module in complex with hexasome
+Supramolecule #2: Chromatin remodeler INO80
+Supramolecule #3: Histones
+Supramolecule #4: DNA
+Macromolecule #1: RuvB-like protein 1
+Macromolecule #2: RuvB-like protein 2
+Macromolecule #3: Chromatin-remodeling ATPase Ino80
+Macromolecule #4: Ino eighty subunit 2
+Macromolecule #5: Chromatin-remodeling complex subunit IES6
+Macromolecule #6: Actin-related protein 5
+Macromolecule #9: Histone H3.1
+Macromolecule #10: Histone H4
+Macromolecule #11: Histone H2A
+Macromolecule #12: Histone H2B
+Macromolecule #7: DNA Strand 1
+Macromolecule #8: DNA Strand 2
+Macromolecule #13: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #14: TETRAFLUOROALUMINATE ION
+Macromolecule #15: MAGNESIUM ION
+Macromolecule #16: ADENOSINE-5'-TRIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.88 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 30mM HEPES, pH7.5 50mM NaCl 0.25mM CaCl2 0.25mM DTT 2mM ADP 3.3mM MgCl2 10mM NaF 2mM AlCl3 0.05% octyl-beta-glucoside |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 15 sec. / Details: FISHIONE Instruments, Nanoclean 10% O2 90% Argon |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV Details: wait time of 5s, blot force at 3, and a blot time of 2s with Whatman blotting paper (Cytiva, CAT No. 10311807). |
| Details | 11-subunit ctINO80 reconstituted with hexasome |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 15384 / Average electron dose: 50.36 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Protocol: OTHER | ||||||||||
| Output model |  PDB-8oo7: |
 Movie
Movie Controller
Controller




























 Z
Z Y
Y X
X








 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)







