[English] 日本語
 Yorodumi
Yorodumi- EMDB-14978: Structure of RQT (C1) bound to the stalled ribosome in a disome u... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of RQT (C1) bound to the stalled ribosome in a disome unit from S. cerevisiae | ||||||||||||
 Map data Map data | locally refined and filtered composite map ligand and ribosome | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | collision / splitting / RQC / RQT / RIBOSOME | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationRQC-trigger complex / ribosomal subunit / regulation of amino acid metabolic process / negative regulation of glucose mediated signaling pathway / translational readthrough / : / mTORC1-mediated signalling / Protein hydroxylation / positive regulation of protein kinase activity / ribosome disassembly ...RQC-trigger complex / ribosomal subunit / regulation of amino acid metabolic process / negative regulation of glucose mediated signaling pathway / translational readthrough / : / mTORC1-mediated signalling / Protein hydroxylation / positive regulation of protein kinase activity / ribosome disassembly / ribosome-associated ubiquitin-dependent protein catabolic process / pre-mRNA 5'-splice site binding / GDP-dissociation inhibitor activity / cytosolic large ribosomal subunit assembly / K63-linked polyubiquitin modification-dependent protein binding / nonfunctional rRNA decay / Formation of the ternary complex, and subsequently, the 43S complex / Translation initiation complex formation / response to cycloheximide / Ribosomal scanning and start codon recognition / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / Major pathway of rRNA processing in the nucleolus and cytosol / mRNA destabilization / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / negative regulation of mRNA splicing, via spliceosome / preribosome, large subunit precursor / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / L13a-mediated translational silencing of Ceruloplasmin expression / negative regulation of translational frameshifting / ribosomal large subunit export from nucleus / 90S preribosome / G-protein alpha-subunit binding / Ub-specific processing proteases / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosomal subunit export from nucleus / regulation of translational fidelity / protein-RNA complex assembly / maturation of LSU-rRNA / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / translation regulator activity / rescue of stalled cytosolic ribosome / cytosolic ribosome / cellular response to amino acid starvation / protein kinase C binding / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / helicase activity / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / positive regulation of apoptotic signaling pathway / macroautophagy / maturation of SSU-rRNA / ubiquitin binding / translational initiation / small-subunit processome / maintenance of translational fidelity / modification-dependent protein catabolic process / cytoplasmic stress granule / protein tag activity / rRNA processing / regulation of translation / ribosomal small subunit assembly / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / defense response to virus / cytosolic large ribosomal subunit / cytoplasmic translation / RNA helicase activity / negative regulation of translation / rRNA binding / structural constituent of ribosome / RNA helicase / protein ubiquitination / ribosome / translation / G protein-coupled receptor signaling pathway / ribonucleoprotein complex / negative regulation of gene expression / response to antibiotic / mRNA binding / ubiquitin protein ligase binding / positive regulation of DNA-templated transcription / nucleolus / ATP hydrolysis activity / mitochondrion / RNA binding / zinc ion binding / nucleoplasm / ATP binding / metal ion binding / nucleus Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
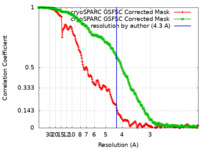
| Method | single particle reconstruction / cryo EM / Resolution: 4.3 Å | ||||||||||||
 Authors Authors | Best KM / Ikeuchi K / Kater L / Best DM / Musial J / Matsuo Y / Berninghausen O / Becker T / Inada T / Beckmann R | ||||||||||||
| Funding support | European Union,  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural basis for clearing of ribosome collisions by the RQT complex. Authors: Katharina Best / Ken Ikeuchi / Lukas Kater / Daniel Best / Joanna Musial / Yoshitaka Matsuo / Otto Berninghausen / Thomas Becker / Toshifumi Inada / Roland Beckmann /    Abstract: Translation of aberrant messenger RNAs can cause stalling of ribosomes resulting in ribosomal collisions. Collided ribosomes are specifically recognized to initiate stress responses and quality ...Translation of aberrant messenger RNAs can cause stalling of ribosomes resulting in ribosomal collisions. Collided ribosomes are specifically recognized to initiate stress responses and quality control pathways. Ribosome-associated quality control facilitates the degradation of incomplete translation products and requires dissociation of the stalled ribosomes. A central event is therefore the splitting of collided ribosomes by the ribosome quality control trigger complex, RQT, by an unknown mechanism. Here we show that RQT requires accessible mRNA and the presence of a neighboring ribosome. Cryogenic electron microscopy of RQT-ribosome complexes reveals that RQT engages the 40S subunit of the lead ribosome and can switch between two conformations. We propose that the Ski2-like helicase 1 (Slh1) subunit of RQT applies a pulling force on the mRNA, causing destabilizing conformational changes of the small ribosomal subunit, ultimately resulting in subunit dissociation. Our findings provide conceptual framework for a helicase-driven ribosomal splitting mechanism. #1:  Journal: Acta Crystallogr D Struct Biol / Year: 2018 Journal: Acta Crystallogr D Struct Biol / Year: 2018Title: Real-space refinement in PHENIX for cryo-EM and crystallography. Authors: Afonine PV / Poon BK / Read RJ / Sobolev OV / Terwilliger TC / Urzhumtsev A / Adams PD | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14978.map.gz emd_14978.map.gz | 43.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14978-v30.xml emd-14978-v30.xml emd-14978.xml emd-14978.xml | 128.7 KB 128.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14978_fsc.xml emd_14978_fsc.xml emd_14978_fsc_2.xml emd_14978_fsc_2.xml | 18.6 KB 18.5 KB | Display Display |  FSC data file FSC data file |
| Images |  emd_14978.png emd_14978.png | 112.6 KB | ||
| Masks |  emd_14978_msk_1.map emd_14978_msk_1.map emd_14978_msk_2.map emd_14978_msk_2.map | 669.9 MB 669.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-14978.cif.gz emd-14978.cif.gz | 21.3 KB | ||
| Others |  emd_14978_additional_1.map.gz emd_14978_additional_1.map.gz emd_14978_additional_2.map.gz emd_14978_additional_2.map.gz emd_14978_additional_3.map.gz emd_14978_additional_3.map.gz emd_14978_additional_4.map.gz emd_14978_additional_4.map.gz emd_14978_additional_5.map.gz emd_14978_additional_5.map.gz emd_14978_half_map_1.map.gz emd_14978_half_map_1.map.gz emd_14978_half_map_2.map.gz emd_14978_half_map_2.map.gz | 53.5 MB 620.9 MB 5.1 MB 620.9 MB 457.5 MB 622.4 MB 618.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14978 http://ftp.pdbj.org/pub/emdb/structures/EMD-14978 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14978 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14978 | HTTPS FTP |
-Related structure data
| Related structure data |  7zuwMC  7zpqC  7zrsC  7zs5C  7zuxC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14978.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14978.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | locally refined and filtered composite map ligand and ribosome | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.045 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14978_msk_1.map emd_14978_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Mask #2
| File |  emd_14978_msk_2.map emd_14978_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: locally refined and filtered map of the ribosome
| File | emd_14978_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | locally refined and filtered map of the ribosome | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: halfmap 2 of the locally refined ligand
| File | emd_14978_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap 2 of the locally refined ligand | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: map of the locally refined and filtered ligand
| File | emd_14978_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | map of the locally refined and filtered ligand | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: halfmap 1 of the locally refined ligand
| File | emd_14978_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap 1 of the locally refined ligand | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: composite map of stalled and collided ribosome
| File | emd_14978_additional_5.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | composite map of stalled and collided ribosome | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap 2 of the ribosome
| File | emd_14978_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap 2 of the ribosome | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: halfmap 1 of the ribosome
| File | emd_14978_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | halfmap 1 of the ribosome | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : RQC trigger complex bound to the stalled ribosome of a disome unit
+Supramolecule #1: RQC trigger complex bound to the stalled ribosome of a disome unit
+Supramolecule #2: ribosome
+Supramolecule #3: RQC trigger complex with Slh1, Cue3, Rqt4
+Macromolecule #1: 18S ribosomal RNA
+Macromolecule #2: 5.8S ribosomal RNA
+Macromolecule #3: 5S ribosomal RNA
+Macromolecule #4: 25S ribosomal RNA
+Macromolecule #5: tRNA
+Macromolecule #6: 40S ribosomal protein S0-A
+Macromolecule #7: 40S ribosomal protein S1-A
+Macromolecule #8: RPS2 isoform 1
+Macromolecule #9: RPS3 isoform 1
+Macromolecule #10: 40S ribosomal protein S4-A
+Macromolecule #11: Rps5p
+Macromolecule #12: 40S ribosomal protein S6-A
+Macromolecule #13: 40S ribosomal protein S7-A
+Macromolecule #14: 40S ribosomal protein S8-B
+Macromolecule #15: 40S ribosomal protein S9-A
+Macromolecule #16: 40S ribosomal protein S10-A
+Macromolecule #17: 40S ribosomal protein S11-A
+Macromolecule #18: 40S ribosomal protein S12
+Macromolecule #19: 40S ribosomal protein S13
+Macromolecule #20: 40S ribosomal protein S14-B
+Macromolecule #21: RPS15 isoform 1
+Macromolecule #22: 40S ribosomal protein S16-A
+Macromolecule #23: 40S ribosomal protein S17-A
+Macromolecule #24: 40S ribosomal protein S18-A
+Macromolecule #25: 40S ribosomal protein S19-A
+Macromolecule #26: RPS20 isoform 1
+Macromolecule #27: 40S ribosomal protein S21-A
+Macromolecule #28: RPS22A isoform 1
+Macromolecule #29: 40S ribosomal protein S23-A
+Macromolecule #30: 40S ribosomal protein S24-A
+Macromolecule #31: 40S ribosomal protein S25
+Macromolecule #32: 40S ribosomal protein S26
+Macromolecule #33: 40S ribosomal protein S27-A
+Macromolecule #34: RPS28A isoform 1
+Macromolecule #35: RPS29A isoform 1
+Macromolecule #36: 40S ribosomal protein S30-A
+Macromolecule #37: RPS31 isoform 1
+Macromolecule #38: Guanine nucleotide-binding protein subunit beta-like protein
+Macromolecule #39: 60S ribosomal protein L2-A
+Macromolecule #40: 60S ribosomal protein L3
+Macromolecule #41: RPL4A isoform 1
+Macromolecule #42: RPL5 isoform 1
+Macromolecule #43: 60S ribosomal protein L6-B
+Macromolecule #44: 60S ribosomal protein L7-A
+Macromolecule #45: 60S ribosomal protein L8
+Macromolecule #46: RPL9A isoform 1
+Macromolecule #47: RPL10 isoform 1
+Macromolecule #48: RPL11B isoform 1
+Macromolecule #49: 60S ribosomal protein L13-A
+Macromolecule #50: 60S ribosomal protein L14-A
+Macromolecule #51: Ribosomal protein L15
+Macromolecule #52: 60S ribosomal protein L16-A
+Macromolecule #53: 60S ribosomal protein L17-A
+Macromolecule #54: 60S ribosomal protein L18-A
+Macromolecule #55: 60S ribosomal protein L19-A
+Macromolecule #56: 60S ribosomal protein L20-A
+Macromolecule #57: 60S ribosomal protein L21-A
+Macromolecule #58: 60S ribosomal protein L22-A
+Macromolecule #59: 60S ribosomal protein L23-A
+Macromolecule #60: RPL24A isoform 1
+Macromolecule #61: 60S ribosomal protein L25
+Macromolecule #62: 60S ribosomal protein L26-A
+Macromolecule #63: 60S ribosomal protein L27-A
+Macromolecule #64: 60S ribosomal protein L28
+Macromolecule #65: 60S ribosomal protein L29
+Macromolecule #66: 60S ribosomal protein L30
+Macromolecule #67: 60S ribosomal protein L31-A
+Macromolecule #68: RPL32 isoform 1
+Macromolecule #69: 60S ribosomal protein L33-A
+Macromolecule #70: 60S ribosomal protein L34-A
+Macromolecule #71: 60S ribosomal protein L35-A
+Macromolecule #72: 60S ribosomal protein L36-A
+Macromolecule #73: 60S ribosomal protein L37-A
+Macromolecule #74: RPL38 isoform 1
+Macromolecule #75: 60S ribosomal protein L39
+Macromolecule #76: 60S ribosomal protein L40-A
+Macromolecule #77: 60S ribosomal protein L41
+Macromolecule #78: 60S ribosomal protein L42-A
+Macromolecule #79: 60S ribosomal protein L43-A
+Macromolecule #80: RQC trigger complex helicase SLH1
+Macromolecule #81: CUE3 isoform 1
+Macromolecule #82: RQC trigger complex subunit RQT4
+Macromolecule #83: MAGNESIUM ION
+Macromolecule #84: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R3/3 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 44.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)