+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli C-P lyase bound to a PhnK ABC dimer and ATP | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | protein complex / transferase / ABC / hydrolase / lyase / carbon phosphorus | |||||||||
| Function / homology |  Function and homology information Function and homology informationalpha-D-ribose 1-methylphosphonate 5-phosphate C-P-lyase / alpha-D-ribose 1-methylphosphonate 5-phosphate C-P-lyase activity / alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase / alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase activity / alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase complex / carbon phosphorus lyase complex / organic phosphonate metabolic process / organic phosphonate transport / organic phosphonate catabolic process / peptide transport ...alpha-D-ribose 1-methylphosphonate 5-phosphate C-P-lyase / alpha-D-ribose 1-methylphosphonate 5-phosphate C-P-lyase activity / alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase / alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase activity / alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase complex / carbon phosphorus lyase complex / organic phosphonate metabolic process / organic phosphonate transport / organic phosphonate catabolic process / peptide transport / 4 iron, 4 sulfur cluster binding / lyase activity / protein homodimerization activity / ATP hydrolysis activity / ATP binding / metal ion binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
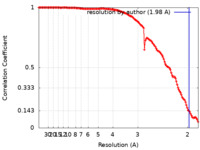
| Method | single particle reconstruction / cryo EM / Resolution: 1.98 Å | |||||||||
 Authors Authors | Amstrup SK / Sofos N | |||||||||
| Funding support |  Denmark, 1 items Denmark, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural remodelling of the carbon-phosphorus lyase machinery by a dual ABC ATPase. Authors: Søren K Amstrup / Sui Ching Ong / Nicholas Sofos / Jesper L Karlsen / Ragnhild B Skjerning / Thomas Boesen / Jan J Enghild / Bjarne Hove-Jensen / Ditlev E Brodersen /  Abstract: In Escherichia coli, the 14-cistron phn operon encoding carbon-phosphorus lyase allows for utilisation of phosphorus from a wide range of stable phosphonate compounds containing a C-P bond. As part ...In Escherichia coli, the 14-cistron phn operon encoding carbon-phosphorus lyase allows for utilisation of phosphorus from a wide range of stable phosphonate compounds containing a C-P bond. As part of a complex, multi-step pathway, the PhnJ subunit was shown to cleave the C-P bond via a radical mechanism, however, the details of the reaction could not immediately be reconciled with the crystal structure of a 220 kDa PhnGHIJ C-P lyase core complex, leaving a significant gap in our understanding of phosphonate breakdown in bacteria. Here, we show using single-particle cryogenic electron microscopy that PhnJ mediates binding of a double dimer of the ATP-binding cassette proteins, PhnK and PhnL, to the core complex. ATP hydrolysis induces drastic structural remodelling leading to opening of the core complex and reconfiguration of a metal-binding and putative active site located at the interface between the PhnI and PhnJ subunits. #1:  Journal: Biorxiv / Year: 2022 Journal: Biorxiv / Year: 2022Title: Structural remodelling of the carbon-phosphorus lyase machinery by a dual ABC ATPase Authors: Amstrup SK / Sofos N / Karlsen JL / Skjerning RB / Boesen T / Enghild JJ / Hove-Jensen B / Brodersen DE | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14444.map.gz emd_14444.map.gz | 110.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14444-v30.xml emd-14444-v30.xml emd-14444.xml emd-14444.xml | 29.5 KB 29.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14444_fsc.xml emd_14444_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_14444.png emd_14444.png | 207.1 KB | ||
| Filedesc metadata |  emd-14444.cif.gz emd-14444.cif.gz | 7.9 KB | ||
| Others |  emd_14444_additional_1.map.gz emd_14444_additional_1.map.gz emd_14444_half_map_1.map.gz emd_14444_half_map_1.map.gz emd_14444_half_map_2.map.gz emd_14444_half_map_2.map.gz | 4.4 MB 203.4 MB 203.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14444 http://ftp.pdbj.org/pub/emdb/structures/EMD-14444 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14444 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14444 | HTTPS FTP |
-Related structure data
| Related structure data |  7z18MC  7z15C  7z16C  7z17C  7z19C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14444.map.gz / Format: CCP4 / Size: 219.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14444.map.gz / Format: CCP4 / Size: 219.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.92189 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Map filtered to local resolution
| File | emd_14444_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map filtered to local resolution | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_14444_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_14444_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : E. coli C-P lyase bound to a PhnK ABC dimer and ATP
+Supramolecule #1: E. coli C-P lyase bound to a PhnK ABC dimer and ATP
+Macromolecule #1: Alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase subuni...
+Macromolecule #2: Alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase subuni...
+Macromolecule #3: Alpha-D-ribose 1-methylphosphonate 5-triphosphate synthase subuni...
+Macromolecule #4: Alpha-D-ribose 1-methylphosphonate 5-phosphate C-P lyase
+Macromolecule #5: Putative phosphonates utilization ATP-binding protein PhnK
+Macromolecule #6: ZINC ION
+Macromolecule #7: alpha-D-ribose-1,2-cyclic-phosphate-5-phosphate
+Macromolecule #8: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #9: MAGNESIUM ION
+Macromolecule #10: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: ATP up from a concentration from 0.25 to 1.5 and MgCl2 to a concentration of 6 mM was added 15 seconds before the sample was plunge frozen | |||||||||||||||||||||
| Grid | Model: UltrAuFoil R0.6/1 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. / Pretreatment - Atmosphere: AIR / Details: 10 mA for 90 seconds in a GloQube, Quorum | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 310 K / Instrument: LEICA EM GP / Details: Blotting for 6-9 seconds before plunging. | |||||||||||||||||||||
| Details | Sample was produced under ATP to ADP+Pi turnover conditions. Sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 6637 / Average electron dose: 62.0 e/Å2 Details: Images were collected in movie-mode with a total of 56 frames |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.4000000000000001 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 135000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)